Abstract
Epulopiscium sp. type B (Lachnospiraceae) is an exceptionally large, highly polyploid, intestinal symbiont of the coral reef dwelling surgeonfish Naso tonganus. These obligate anaerobes do not form mature endospores and reproduce solely through the production of multiple intracellular offspring. This likely makes them dependent on immediate transfer to a receptive host for dispersal. During reproduction, only a small proportion of Epulopiscium mother-cell DNA is inherited. To explore the impact of this unusual viviparous lifestyle on symbiont population dynamics, we investigated Epulopiscium sp. type B and their fish hosts collected over the course of two decades, at island and reef habitats near Lizard Island, Australia. Using multi-locus sequence analysis, we found that recombination plays an important role in maintaining diversity of these symbionts and yet populations exhibit linkage disequilibrium (LD). Symbiont populations showed spatial but not temporal partitioning. Surgeonfish are long-lived and capable of traveling long distances, yet the population structures of Epulopiscium suggest that adult fish tend to not roam beyond a limited locale. Codiversification analyses and traits of this partnership suggest that while symbionts are obligately dependent on their host, the host has a facultative association with Epulopiscium. We suggest that congression of unlinked markers contributes to LD estimates in this and other recombinant populations of bacteria. The findings here inform our understanding of evolutionary processes within intestinal Lachnospiraceae populations.
Similar content being viewed by others
Introduction
Epulopiscium spp. and related bacteria, known as epulos, are a morphologically diverse group of intestinal symbionts renowned for their large size and disparate reproductive strategies [1,2,3,4]. Phylogenetically, this group is affiliated with the Lachnospiraceae, a family that comprises functionally crucial intestinal symbionts of vertebrates [5, 6]. Abundant populations of epulos are found in surgeonfish species that regularly consume algae or detritus [2]. Epulos likely contribute to the breakdown of refractory algal polysaccharides ingested by their hosts and serve as important mediators of carbon flow in coral reef systems [7,8,9]. The model for these unusual bacteria is referred to as Epulopiscium sp. type B, which displays many useful traits that have facilitated studies of Epulopiscium biology [10,11,12,13,14]. In addition to their distinct morphology, Epulopiscium sp. type B is often the only epulo located in adult Naso tonganus [15, 16]. Other hosts harbor multiple morphologically and phylogenetically distinct epulo lineages [2, 4, 17]. On the basis of draft metagenome sequences from Epulopiscium sp. type B populations [14] (NZ_ABEQ01000000), these heterotrophic symbionts are predicted to be obligate fermenters, as neither genes for cytochromes nor genes for enzymes to relieve oxidative stress (e.g. catalase or superoxide dismutase) have been found. Thus far, no free-living Epulopiscium sp. type B have been detected in environmental surveys. These observations along with its affilation with Clostridium cluster XIVb [18] suggest that the intestinal symbiont Epulopiscium sp. type B requires a host that regularly consumes algae for a stable anoxic environment rich in nutrients.
Many epulos produce endospores that might aid in dispersal and maintenance of populations in their surgeonfish host [4]. However, Epulopiscium sp. type B has lost the ability to produce mature endospores and no longer uses binary fission for reproduction [14]. Instead, these bacteria produce multiple intracellular offspring using a derived form of endosporulation (Fig. 1). The ecological and evolutionary processes leading to this unique form of bacterial viviparity are not well understood, but may have been determined in part by the symbiotic association of Epulopiscium sp. type B with its particular surgeonfish host, Naso tonganus. In a previous study, we suggested that the manner in which symbionts enter a host intestinal tract can influence the reproductive biology of endospore forming symbionts [19]. Metabacterium polyspora, a close relative of Epulopiscium spp., uses the production of multiple endospores as its primary means of reproduction in its guinea pig host. This pivot away from binary fission and toward multiple endospore formation likely reinforces the symbiotic association, as these bacteria regularly cycle out of and back into the host intestinal tract. Guinea pigs are coprophagous and reproduction by sporulation allows the offspring of M. polyspora to survive the oxic environment outside of the host, as well as transit through the mouth and stomach of the host. In epulos associated with fish, codiversification studies suggest specific relationships between distinct epulo clades and particular surgeonfish species that correlate with host feeding preference [17]. We hypothesize that these associations impact symbiont reproductive strategies as well. Most surgeonfish that host epulos contain multiple morphotypes and multiple phylotypes [2, 4, 17]. Adult N. tonganus are often observed to harbor only one large epulo, Epulopiscium sp. type B [15, 16]. This simple system provides a unique opportunity to explore how the symbiotic alliance impacts the population structure and reproductive strategy of epulos and other gut-associated Lachnospiraceae.
Epulopiscium sp. type B daily life cycle. These populations are maintained by an unusual reproductive strategy, which employs the formation of multiple intracellular offspring. Binary fission has never been observed in Epulopiscium sp. type B. a Accumulation of DNA (in blue) at the poles of intracellular daughter cells marks the beginning of formation of the next generation (granddaughter cells). b Bipolar division often occurs just prior to emergence of daughter cells from their mother cell. Note the DNA in the mother cell is less pronounced at this stage and mother cells do not survive daughter-cell release. c Polar cells are (d) fully engulfed by their mother cell. e Internal offspring cells continue to grow until (a) they nearly fill the mother-cell cytoplasm. Figure modified from [20]
Epulos, like most gut microbiota of vertebrates, are likely acquired from the environment. However, surveys of near-complete 16S rRNA gene sequences of Epulopiscium sp. type B cells from many host individuals recovered minimal genetic diversity (>99% sequence identity), suggesting that these symbiont populations may be clonal and experience dispersal limitation [15, 20]. Yet, given the dynamic life history of surgeonfish (e.g. aggregate spawning, dispersal of eggs and larvae, and pulses of juvenile settlement on a reef [21]), direct transmission of epulos from a parent to its offspring seems unlikely. A recent phylogeographic study of a broadly distributed surgeonfish species suggested that larval fish are dispersed widely [22]. Furthermore, surveys of gut symbionts in fish at different life stages revealed no epulos in newly settled juvenile surgeonfish and distinct epulo communities between juveniles and adults [23]. Surgeonfish most likely acquire epulos through conspecific coprophagy [23]. Unlike vertically transmitted insect endosymbionts [24], populations of horizontally transmitted obligate symbionts do not appear to be as genetically constrained by dispersal bottlenecks [25, 26]. The apparent clonality of Epulopiscium sp. type B suggests that other factors peculiar to the symbiont (e.g., physiology and cell cycle) or its host (e.g., N. tonganus genetics, anatomy, diet or behavior) may be influencing Epulopiscium sp. type B diversity.
Epulopiscium sp. type B display several remarkable features. These large cigar-shaped cells range from ~100 – 300 µm in length. They are extremely polyploid and harbor tens of thousands of genome copies [11, 13]. Genomes within Epulopiscium sp. type B cells are hypothesized to be nearly identical [13], although some variation may exist. The daily reproductive cycle of Epulopiscium sp. type B (Fig. 1) is synchronized within an individual host fish [11, 12]. Normally, two offspring are produced each day per mother cell but as many as 12 intracellular offspring have been observed [27]. Importantly, it has been estimated that during reproduction, ~1% of Epulopiscium sp. type B mother-cell DNA is passed on to daughter cells [12]. Yet, even at late stages of offspring development, mother-cell chromosomes continue to replicate [10]. Consequently, genome copies take on either somatic or germline roles [10]; some intact copies are inherited by the next generation but most of the DNA appears to be required only to support mother-cell metabolism. Eventually, the somatic copies are dismantled or released into the environment by lysis of the dying mother cell (Fig. 1).
The limited inheritance of mother-cell DNA could restrict genetic diversity of the Epulopiscium sp. type B population by loss of novel genes or mutations accrued in somatic but not germline genome copies. Additionally, retention of daughter cells within a mother cell during much of the offspring growth cycle, which occurs separately from other developing cells, poses a physical obstacle to acquiring new genetic material through horizontal gene transfer (HGT). Furthermore, host-dependent transmission may limit the genetic material available for exchange in a confined population within an individual host. As a consequence, the overall fitness of the symbiont population is expected to decline due to clonal interference; a process where competition between individual clones with different beneficial mutations slows down the accumulation of beneficial mutations for the population as a whole [28].
We hypothesize that host-dependent interactions and the unusual reproductive biology of the symbionts are driving the evolution of Epulopiscium sp. type B by limiting the diversity of the symbiont populations. Epulos have not yet been cultivated in the lab and surgeonfish lose these symbionts when held in captivity [1]. Therefore, we used cultivation-independent techniques to explore the population structure of Epulopiscium sp. type B collected from wild-caught hosts. Taking advantage of the large size, polyploid genome and distinct morphology of the symbionts, we collected individual cells from archived N. tonganus gut contents and subjected each to single-cell, whole-genome amplification. We used a multi-locus sequence analysis (MLSA) approach for the first fine scale population survey of Epulopiscium sp. type B, which allowed us to identify genetic differences between individual cells. We also investigated the N. tonganus population using MLSA and analyzed codiversification between partners of this host-symbiont association. Our findings reveal the importance of recombination for maintaining population diversity in this gut symbiont. We predict that the patterns of inheritance and genotype mixing we observed are driven by mechanisms that likely impact the population structure of other intestinal Lachnospiraceae. Furthermore, we suggest that population analyses of intestinal symbionts like these can provide insight into the distribution, movement and life histories of long-lived reef fish.
Methods
Sample collection
Naso tonganus were collected by spear from island or outer barrier reef habitats around Lizard Island, Australia, at three different 10-year intervals: circa 1990, 2003, and 2013 (Fig. 2). Surgeonfish that regularly consume algae have a long, coiled intestinal tract [2]. Other than the coiling pattern, the intestine of N. tonganus has no distinct morphological features. To collect samples containing Epulopiscium sp. type B, the N. tonganus intestinal tract was removed and uncoiled. The unraveled intestine was laid out in 4 equal-length segments, as previously described [7]. In this scheme, the stomach is defined as segment I, and from anterior to posterior, the intestinal segments are referred to as segments II – V. Samples of Epulopiscium sp. type B were taken from segment IV, where large numbers of symbionts are located. Intestinal contents from each individual fish were fixed in 80% ethanol and stored at −20 °C. Sample information, a description of Epulopiscium cells present, and demographic data for each host were recorded (Table 1).
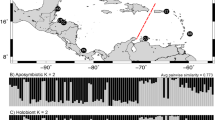
Collection sites for each Naso tonganus used in this study, located on a map of Lizard Island and nearby outer barrier reef. Arrow in the inset indicates location of the sampling area off the northeast coast of Australia. Islands (including Lizard Is., Bird Is. and South Is.) are shaded dark grey while dashed lines outline reefs. Approximate sampling locations for each N. tonganus individual are indicated by a unique color and sample ID. Details are provided in Table 1. Sampling time is indicated by either a circle, square, or triangle for year intervals circa 1990, 2003 and 2013, respectively
Single-cell, whole-genome amplification and MLSA of Epulopiscium sp. type B
Briefly, individual Epulopiscium cells were manually collected from fixed intestinal contents and subjected to whole-genome amplification (WGA) using the REPLI-g Kit (Qiagen) for 16 h at 30 °C, following the manufacturer’s protocol. Seven housekeeping genes (dnaC, ftsZ, mreB, radA, recA, rpoB, and secA) and the 16S rRNA gene were PCR amplified using primers and conditions described in Supplementary Table S1. See Supplementary Information for details of single-cell processing, WGA, MLSA scheme design, and sequence analysis.
DNA extraction and MLSA of Naso tonganus
Host DNA was extracted from fixed intestinal contents. Fish haplotypes were based on mitochondrial (cox1 and Cytb) and nuclear genes (ENC1, plagl2, and zic1). See Supplementary Information for details of DNA extraction and sequence analysis. Primer sets and amplification conditions are described in Supplementary Table S1. Haplotypes were reconstructed from the unphased sequence data using the phase option in DnaSP version 5.10 [29].
Phylogenetic analyses
Concatenated sequences were joined head-to-tail in-frame and trimmed for seven Epulopiscium loci and five N. tonganus loci (unphased). 16S rRNA gene sequences were not included because Epulopiscium has multiple rRNA operons [13]. Bayesian and maximum likelihood (ML) phylogenies were constructed in MrBayes version 3.2.4 [30] and PhyML version 3.0 [31], respectively, each using the generalized Time Reversible nucleotide substitution model [32] plus invariant sites (I) and Γ rate heterogeneity. Markov chain Monte Carlo was run for one million generations, sampling every 1000 generations. The first 25% of trees were discarded as burnin. A total of 1000 bootstrap replicates were performed for ML phylogenies.
Nucleotide diversity, recombination and gene flow analyses
For both Epulopiscium and host data sets, statistics for single genes and populations were calculated using DnaSP [29]. These tests included the number of polymorphic sites, nucleotide diversity (π) corrected using the Jukes Cantor method [33], haplotype diversity (Hd), and neutrality tests (Tajima’s D [34], Fu and Li’s D* [35], and Fu’s F [36]). Individual markers were analyzed for signatures of selection by calculating the ratios of nonsynonymous to synonymous substitutions (dN/dS) using the default NG86 nucleotide substitution model [37] in START2 (version 2) [38]. Markers were further analyzed with PAML 4.9 h [39], using multiple codon frequency models as described in the Supplementary Information.
For the Epulopiscium sp. type B data set, recombination breakpoints were identified using the Genetic Algorithm for Recombination Detection (GARD) [40] available through the Datamonkey webserver [41]. Results were visualized using R [42] and package “ggplot2” [43]. Population structure was modelled using STRUCTURE 2.3.4 [44], as described in Supplementary Information, to examine population subdivisions and presence of admixture. Admixture within and between subpopulations is indicative of recombination. Additionally, a pairwise homoplasy index test (ΦW) [45] was performed using SplitsTree 4.13 [46] to detect recombination by examining the genealogical history of pairs of sites. This approach is useful for data with complex population structure and demographic histories, differentiating between population growth and recombination. Estimates for linkage disequilibrium were performed using the standardized index of association (ISA) [47] from START2.
To determine the level of gene flow between symbiont groups at different subdivisions, an analysis of molecular variance (AMOVA) [48] was implemented and tested for significance against 1000 permutations with Arlequin 3.5 [49]. A Mantel test was performed to examine whether genetic distance correlated with geographic distance and those results were plotted using the R package “ade4” [50]. Two distinct hierarchical subdivisions were tested independently: time intervals and habitat location (reef vs. island). Variability was assessed among Epulopiscium individuals within time intervals (ΦGT), among Epulopiscium populations within each host (ΦST), and among populations within time intervals (ΦSG). Multiple linear regressions were performed in R to examine whether π or Hd are correlated with demographic parameters listed in Table 1.
Codiversification analysis of N. tonganus and Epulopiscium sp. type B
The global signal of codiversification and contribution of individual host-symbiont associations were analyzed using Parafit [51] and PACo [52], implemented in R packages “ape” [53] and “vegan” [54]. The null hypothesis differs between the two tests; Parafit tests independent host and symbiont evolution whereas PACo explicitly tests independence of symbiont phylogeny on host phylogeny. The input for both analyses were aligned concatenated sequences from individual hosts and symbiont sequence types (sSTs), converted to distance matrices using the K80 model [55]. The significance of both tests were established from 100,000 permutations. The P-value was determined for each host-symbiont pair by ParaFitLink1 tests. A tanglegram was generated with iTol version 4.2 [56].
Results
MLSA of Epulopiscium sp. type B cells revealed high variability
This study examined symbiont diversity and distribution among and within individual hosts collected over the course of ~20 years (1990–2014) in a sampling area that covers discontinuous island and reef habitats within a 17 km radius circle (Fig. 2). Comparisons of Epulopiscium populations collected over time were used to improve the detection of population structure. A total of 113 individual Epulopiscium sp. type B cells were collected and analyzed from 12 different N. tonganus (~10 cells/fish) (Supplementary Table S2). Cells were subjected to WGA and genes were PCR amplified from these DNA samples. The sequence of each PCR product was determined using the Sanger method. Amplified 16S rRNA gene sequences were 99-100% identical to published 16S rRNA gene sequences from Epulopiscium sp. type B (Supplementary Fig. S1). This supported previous observations of low population diversity. However, using a higher resolution approach with additional markers (dnaC, ftsZ, mreB, radA, recA, rpoB, and secA), we observed high sequence variability and identified 88 symbiont sequence types (sSTs) (Fig. 3, Supplementary Fig. S2). All housekeeping gene sequences shared 99-100% sequence identity with genes in the Epulopiscium sp. type B draft genome (Supplementary Table S3). Comparison of the concatenated sequences revealed 64 polymorphic sites across the 4005-base length alignment (Supplementary Table S4). For the entire data set, nucleotide diversity (π) was 0.00287 and was similar across time and space (ranging 0.00217-0.00287) (Table 2). Nucleotide diversity varied across individual markers (0.00543–0.00109), except secA had no polymorphisms (Supplementary Table S4).
Bayesian phylogenetic tree of Epulopiscium sp. type B cells. Tree was constructed using concatenated sequences of seven housekeeping markers (dnaC, ftsZ, mreB, radA, recA, rpoB, and secA). Nodes with Bayesian posterior probabilities ≥ 0.70 are indicated by solid circles and nodes with ML bootstrap values ≥ 50% from 1,000 replicates are indicated. Each symbiont subtype (sST) is color coded by host source as in Fig. 2, and sSTs found in multiple hosts are indicated by red asterisks. Colored branches represent habitat type (green = island; blue = reef) and sSTs found in both island and outer barrier reef habitats are indicated with a crossed branch. Scale represents nucleotide changes per position
Neutrality tests predict either a recent symbiont population expansion or recombination
Next, neutral processes were explored to determine whether stochastic processes of dispersal and genetic drift explained the level of diversity observed in the symbiont population. Subpopulations within different habitats as well as the entire population sampled across space and time were examined; Class I neutrality tests (Tajima’s D and Fu and Li’s D*) were not significant and the Class II neutrality test (Fu’s F) was significantly negative (Table 2). Genes that likely contribute to the significant Fu’s F were dnaC, radA, and recA (Supplementary Table S4). Although a significantly negative Fu’s F would suggest an excess number of alleles due to a recent population expansion or genetic hitchhiking, this test is strongly affected by recombination which may produce a false positive result [57]. Therefore, the contribution of recombination was analyzed to determine which demographic parameters were influencing the diversity of the symbiont populations.
Host and location influence symbiont population structure
Deviations from the neutrality model prompted the exploration of parameters that might be influencing symbiont allele frequencies. Most sSTs were unique, but sST65 was more frequently encountered and observed in hosts collected at the Lizard Island sites (Fig. 3, Supplementary Fig. S2). Epulopiscium populations showed high haplotype diversity, averaging 0.991 across all samples (Table 2). Generally, individual hosts harbored diverse symbiont populations (Supplementary Table S2). However, host Nt_050613 had the least diverse population (Hd 0.667). No host demographic information (fish size, and location collected) correlated with low symbiont Hd.
Time and location were discernible despite the notable location bias in the samples used here, in which fish collected prior to 2011 were predominantly from the outer barrier reef and after 2011 most were taken near Lizard Island (Fig. 2, Table 1). AMOVA indicated time intervals contributed ~8% of the variation (P = 0.0499), whereas habitat location (reef vs. island) contributed 16% (P < 0.0001) (Supplementary Table S5). In both subdivisions, the majority of the contribution was found within populations (P < 0.0001). Thus, factors within hosts had the greatest influence on the diversity of symbiont populations. These may include host genotype, social behavior or dietary differences.
Despite the statistical noise contributed by variation within host populations, there was a significant correlation between geographic distance and pairwise FST (Mantel test: R2 = 0.136, P = 0.0014) (Supplementary Fig. S3). A Principle Coordinate Analysis (PCoA) of pairwise FST values further supported clustering based on location (Fig. 4). Island populations clustered tightly together with the exception of Nt_050913 from Research Station Beach, which grouped with Nt_031411 from North Day Reef. Other reef populations clustered together with the exception of the two populations from the southernmost outer reef collection location within our study area, Detached Reef. Both of these divergent symbiont samples were collected in the same location and year. These observations of population subdivisions and admixture were confirmed by simulations using STRUCTURE (Supplementary Information) (Supplementary Fig. S6). Altogether, these data suggest that symbiont diversity within an individual fish is dependent on host feeding and location.
PCoA of the genetic differentiation (FST) of Epulopiscium sp. type B populations. The 95% confidence interval for each habitat type (island or outer barrier reef) is indicated by ellipses. Sampling time is indicated by either a circle, square, or triangle for year intervals circa 1990, 2003 and 2013, respectively
Recombination contributes to diversity in Epulopiscium sp. type B populations
Symbiont population structure and a significantly negative Fu’s F suggested symbiont gene flow between hosts. Therefore we investigated whether recombination or spontaneous mutation was the mechanism contributing to the variation observed. Multiple tests found evidence of recombination among the Epulopiscium sp. type B populations. GARD identified at least one recombination breakpoint at position 615 of the concatenated sequences (Supplementary Fig. S4). Topological incongruence was supported by the KH test (P < 0.001) (Supplementary Fig. S5). The homoplasy index test (ΦW) had a significant level of convergence/recombination among the total population, within habitats, and even within certain hosts (Nt_101203, Nt_101803, Nt_031905, Nt_031411) (Table 2, Supplementary Table S2). These data suggest that HGT among Epulopiscium sp. type B cells helps maintain allelic diversity in the symbiont population and may diminish clonal interference.
Preliminary MLSA of Naso tonganus suggests deviation from neutrality
To examine the possible contribution of host genetics to symbiont population structure, host allelic frequencies were characterized. Previously reported acanthurid markers and four new loci that have not been reported for N. tonganus were used (Supplementary Table S1). Five loci were successfully amplified and analyzed from 11 out of 12 fish used in this study (Supplementary Table S6). Analysis of the concatenated sequences revealed 19 polymorphic sites across 4,125 bp, with π ranging from 0.00041 to 0.00211 per locus. All markers were under stabilizing or neutral selection. Neutrality tests for individual markers were not significant. However, Fu and Li’s D* were significantly positive (1.593, P < 0.05) across the length of the concatenated sequences, reflecting an excess of intermediate-frequency alleles which can result from population bottlenecks, structure and/or balancing selection.
Fourteen haplotypes were identified, in which 7 individuals were homozygous (hap1–3, hap6, hap13–14) and 4 were heterozygous (Supplementary Fig. S7). Since the heterozygous haplotypes clustered tightly within each individual, the multilocus phylogeny was constructed using unphased sequences and each host is referred herein per their sample ID (Supplementary Fig. S8). Although the genealogy is not well resolved, two main clades are distinguishable. One clade contains 3 out of the 4 Lizard Island samples and one host collected from the outer barrier reef. The other clade contains mostly hosts from the outer barrier reef and the other Lizard Island associated host.
Codiversification analysis reveals symbiont dependence on host phylogeny
Significant global codiversification links between sSTs and individual hosts were detected from both Parafit (3.268 × 10-9, P = 0.0002) and PACo (m2 = 8.644 × 10−5, P < 0.00001). However, only a few significant ParaFit1 links (P < 0.05) contributed to the codiversification signal (Fig. 5). Yet, the significant PACo results indicate that the symbiont phylogeny is dependent on the host. The Procrustean superimposition plot confirmed the influence of some host individuals (Nt_050913, Nt_120714, Nt_050613, Nt_102990, and Nt2_101203) on the symbiont phylogeny (Supplementary Fig. S9). Host Nt_050913 had the greatest number of significant links (9 links) with its symbionts. Including marginally significant links (P < 0.1), better resolved the symbionts/host relationship. These data suggest that Epulopiscium sp. type B and their surgeonfish host share a facultative relationship where symbionts are more dependent on their host than vice versa. This model is further supported by our observations of apparently healthy adult N. tonganus which were collected within our study area but appeared to harbor no Epulopiscium sp. type B. Notably, sSTs that occur in multiple hosts (asterisks, Fig. 5) often had significant links with hosts that were more phylogenetically related and collected in proximal locations, thus suggesting that host genetics or habitat sharing influences symbiont populations. For example, sST65 occurred in three hosts and was significantly linked to Nt_120714 and Nt_050913 but marginally linked to Nt_050613. Host Nt_050613 was more distantly related to the sister pair Nt_120714 and Nt_050913 but all three were collected from Island locations. sST51 occurred in hosts Nt_120714 and Nt_010212, but had a significant link with only Nt_120714. Likewise, sST8 was significantly linked with host Nt_050913 and not Nt_031411.
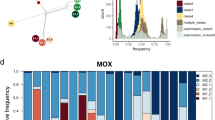
Tanglegram of host N. tonganus (left) and symbiont Epulopiscium sp. type B sSTs (right). Parafit Global test = 3.268 × 10-9, P = 0.0002 (10,000 permutations). 85 host-sST links were detected and are indicated by matching colors. sSTs found in multiple hosts are indicated with an asterisk; an outlined box specifies two hosts while a shaded box specifies three hosts (e.g., sST65). Significant ParaFitLink1 tests are shown by red dashed lines (P < 0.1) and red solid lines (P < 0.05). Host collection habitats are indicated with green (island) or blue (outer barrier reef) cladogram branches
Discussion
The Lachnospiraceae are recognized as influential members of the gastrointestinal tract microbiota of terrestrial vertebrates and some marine vertebrates, including surgeonfish. One member of the Lachnospiraceae family, Epulopiscium sp. type B, can form a specific relationship with the surgeonfish N. tonganus. Moreover, these intestinal symbionts display an unusual daily reproductive cycle. Both traits likely impact the evolutionary trajectory of the bacteria. Using a fine scale population survey of this intestinal symbiont and its host, we explored their codiversification and identified factors that contribute to the evolution of Epulopiscium sp. type B. Remarkably, we discovered high genetic diversity among Epulopiscium sp. type B populations within individual fish as well as evidence of extensive symbiont genotype mixing between fish. Despite this, widespread linkage disequilibrium (LD) in the symbiont population and the identification of one host-associated subpopulation with low haplotype diversity support the hypothesis that transmission bottlenecks are occurring. These data suggest that recombination contributes to Epulopiscium sp. type B genetic diversity and compensates for deleterious effects imposed by its lifestyle. Furthermore, our analyses suggest that Epulopiscium sp. type B is not only dependent on its host for transmission, but horizontal transmission of the symbiont is also necessary for the incursion of new genetic material.
Evolution of horizontally acquired symbionts depends on host
The limited number of significant codiversification links and structure of symbiont populations support the model that Epulopiscium sp. type B cells are horizontally transmitted. Samples collected over the course of more than two decades were used in the study to try to improve the likelihood of detecting changes in symbiont population structure over time. Notably, acanthurids collected from the Great Barrier Reef can live 30–45 years [58]. This suggests that sampling beyond the life span of N. tonganus may be needed to illuminate time as a more significant contributor to symbiont population variation. However, the presence of identical alleles and similar population structures between fish collected at different time points implies that the introduction of new symbionts to an established population may occur throughout the life span of an individual host. Since Epulopiscium populations are easily lost when surgeonfish are brought into captivity, we assert that in the wild, dietary changes or stress may alter symbiont populations leading to sweeps or facilitating introgression.
Codiversification analyses suggest that host genetics contributes to Epulopiscium sp. type B population structure. There is evidence that host genetics refines the composition of microbiota associated with animals, including some fish [59, 60]. Interactions between gut microbiota and the host immune system likely contribute to observed variation [61, 62]. A study of the threespine stickleback (Gasterosteus aculeatus) found that the composition of gut microbiota depended more on host genotype than on any other transient environmental factors [63], suggesting that the fish host filters through countless environmental microorganisms to establish its gut microbiome.
Selection of symbionts may also arise through host ecology (e.g., diet) and/or composition of the resident microbiota. In herbivorous acanthurids, Epulopiscium spp. can be the most dominant taxa in the hindgut [2, 60]. Compared to zooplanktivores, these fish have longer intestinal tracts, which contain higher levels of fermentation products (e.g., acetate) [7, 8]. Although N. tonganus has been referred to as an herbivore [64, 65], it is generally considered an omnivore [9]. Thus, individual feeding preferences or the availability of suitable food may be wide-ranging and may impose a strong selection for particular microbiota and compatible Epulopiscium strains. Evidence of codiversification of gut symbionts with their surgeonfish hosts suggests that there is pressure to retain specific phylotypes [4, 17]. Here we have increased the resolution of codiversification and extend it to the population level for Epulopiscium sp. type B.
The phylogenetic diversity of the Epulopiscium sp. type B parallels gut symbiont profiles described for social bees and mammals [66,67,68]. In these systems, communities tend to have low species richness, and specific lineages exhibit shallow fan-like branching patterns which suggests that hosts are inoculated with a few founder species that later diversify in situ. Diversification from founder species may enable niche partitioning among strains in a nutrient-rich gut environment. Recent studies in an experimental model using gut inoculations with an auxotrophic E. coli strain demonstrated that niche partitioning reduced clonal interference within a mouse host [69]. Epulopiscium sp. type B likely plays a major role in host nutrition and our data suggest niche partitioning may also be occurring here. The functional significance of these diversified populations is worth further study.
Gut microbiota footprint reveals limited patterns of movement of N. tonganus
Some coral reef fish, including some acanthurids, exhibit high genetic connectivity across large oceanic distances [70, 71]. This may involve dispersal of pelagic larvae over distances exceeding 10,000 km [72], whereas small ranging fish exhibit spatial structure influenced by suitable reef habitats and seascape discontinuity [73]. Recent studies of large, coral reef associated fish species have revealed high levels of larval retention to parental habitats [74], suggesting that larval dispersal is not as extensive as previously proposed. The small sample size and markers used in this study were insufficient to address genetic connectivity. However, the divergence of host-specific symbiont populations suggests host foraging and movement patterns are governed in part by seascape discontinuity. This observation is further supported by the host’s positive Fu and Li’s D* that suggest these fish may have experienced a population bottleneck or more likely have undergone population subdivision.
Spatial structuring at our study site was surprising given that the large N. tonganus would be expected to easily traverse this distance. However, traveling from Lizard Island to the outer reef would require a fish to cross 25 km of open water up to 50 m deep. Surgeonfish may be reluctant to venture far from a reef due to increased vulnerability to predators. With a few notable exceptions, symbionts from fish collected near Lizard Island were more closely related to one another than to symbionts of hosts from outer barrier reef, and vice versa. For example, host Nt_050913 is genetically more similar to Nt_120714 from Bird Island but its Epulopiscium population is more closely related to symbionts of Nt_031411 from North Day reef. Another form of symbiont population discontinuity was observed at the southern end of the outer barrier reef within our study area. Both symbiont populations from fish collected near Detached Reef are unique and suggest that these fish came from a more distant location, perhaps south of our sampling area. Clearly, more detailed surveys are needed to test the hypothesis, but data collected in this study indicate that Epulopiscium sp. type B populations provide a record of movement of individual fish among groups of fish associated with the Great Barrier Reef. These suggested patterns of movement are consistent with the ‘commuting’ and ‘foraying’ patterns observed in Naso unicornis studies using radio telemetry-based tracking [75, 76].
Genetic exchange within the Epulopiscium sp. type B populations
The results provided here support the hypothesis that Epulopiscium sp. type B populations depend on environmentally acquired alleles to conserve genetic diversity. Based on the draft genome, Epulopiscium sp. type B has the recombination and DNA maintenance genes needed to support this mechanism [11].
Adaptive rate studies highlight the advantage that recombination has over spontaneous mutations, especially in systems where population structures exist. Experimental studies of E. coli and Saccharomyces cerevisiae demonstrated that beneficial mutations became fixed sooner in strains with high recombination rates than in strains with high mutation rates [77, 78]. Model simulations have shown that higher rates of HGT in small, structured populations made these populations more resistant to Muller’s ratchet than larger, mixed populations [79]. This suggests that “cross-referencing” between subdivided populations, facilitated by HGT, enhances genetic diversity. The gut ecosystem provides a structural framework in which recombination could be highly impactful as we observed in Epulopiscium sp. type B populations.
Despite strong evidence for recombination, the Epulopiscium populations studied exhibited LD, suggesting that there is not enough recombination occurring to observe random assortment of markers. Previous reports estimated that at least a 20-fold relative contribution of recombination per point mutation (r/m) is needed for loci to assort independently [47, 80]. However, Spratt et al.[81] cautions against using LD as a proxy for relative recombination rates, recognizing that highly recombinant bacterial populations may still appear to be in LD. We suggest that LD may be common in naturally competent populations of bacteria. The ability to take up DNA from the environment and stably integrate that DNA into the genome is widespread in the bacteria and the archaea [82]. Congression, the phenomenon by which competent cells are co-transformed with unlinked DNA molecules at high frequencies [83, 84], has been used by geneticists for decades to introduce specific genetic changes without the need for selection of both markers. However, the frequency at which multiple unlinked pieces of DNA are incorporated in a single competent cell has only recently been analyzed systematically [85]. For both Vibrio cholerae and Streptococcus mutans, competent cells can take up two unlinked markers at surprisingly high co-transformation frequencies of 50–60%. This tendency for some members of a naturally competent population to be transformed by numerous unlinked, unselected genes would contribute to LD in natural populations.
The detection of at least one recombination site, statistical support of topology incongruence, and admixture within and between subpopulations provide additional support for recombination in Epulopiscium sp. type B. Evidence of a transmission bottleneck (low Epulopiscium Hd in host Nt_050613) suggests that the detected recombination events were relatively recent in Epulopiscium evolutionary history. Even in the obligate symbiosis between the bivalve Solemya velum and its bacterial gill endosymbionts, mixed infections and recombination occurs at a high enough frequency to maintain symbiont diversity [25]. This further suggests that symbiont allele frequencies reflect a dynamic state in which populations may not reach equilibrium. Therefore, we speculate that microbial populations in gut systems might be predominantly in a state of flux.
The extreme polyploidy of Epulopiscium sp. type B may be confounding the typically observed relationship between LD and high recombination rate as well. Intracellular genetic diversity appears to be low, suggesting a strong pressure for gene conversion. Genome redundancy may mask the effects of deleterious mutations by purifying genes through gene conversion as observed in asexual amoeba [86]. High conversion rates may also limit genetic diversity and thus contribute to LD through clonal interference.
Model for the genetic inheritance of externally acquired DNA in Epulopiscium sp. type B
Multiple Firmicutes species coordinate DNA release with competence, either by regulating cell lysis [87, 88] or secretion of DNA [89]. The circadian cell cycle of Epulopiscium sp. type B provides an opportunity to coordinate competence induction with the daily release of genomic DNA from the population. This scenario would allow a newly independent offspring cell to take up and incorporate DNA from both its own mother cell and others in the population (Fig. 6). We hypothesize that some somatic genomic DNA is released when mother cells lyse (as in Fig. 1 stage B). If competence complexes are located near the poles of emerging daughter cells, where the next generation (granddaughters) have been initiated but are not yet fully engulfed (see Fig. 1 stages B & C), extracellular DNA could be incorporated into pole-associated chromosomes, thus increasing the likelihood of vertical transmission to future generations. Recombination would likely take place once replication begins in the offspring (Fig. 1 stage D).
Model for coordinated mother-cell lysis and offspring competence induction in Epulopiscium sp. type B. Within each host, Epulopiscium sp. populations are synchronized with respect to development. The nearly simultaneous lysis of mother cells could provide a diverse pool of genomic DNA available for uptake by competent cells. We hypothesize that daughter cells, emerging at the time of mother-cell lysis, are competent. The newly released cells would be in the earliest stages of offspring development. Some may have divided asymmetrically but not yet engulfed the polar cells, others may be more advanced in their development. Those cells at later stages of development, after polar-cell engulfment is complete, would have physical barriers to the uptake of DNA that could be inherited. We suggest that the uptake of DNA at the poles of a newly emerged Epulopiscium cells would increase the chances of inheritance of DNA acquired by HGT whereas uptake away from polar cells would not be inherited. Transformation of cells by the uptake of DNA from a different sequence type is indicated by color changes
Conclusions
The size and extreme polyploidy of Epulopiscium sp. type B make it an ideal model for using the single-cell genome-amplification approaches which facilitated this population study. We found that population bottlenecks imposed by unusual life history strategies, which are closely tied to maintaining a symbiotic association, can be overcome by simple changes to widely available mechanisms: increasing ploidy, and allele exchange using HGT and homologous recombination. There is a growing appreciation for the impact of polyploidy and recombination on the evolution of bacterial populations [90]. Given the broad range of bacterial life histories of many gut microbes (including non-endospore-forming Lachnospiraceae) [91], we suggest that these populations may be under similar pressures as we have observed for Epulopiscium sp. type B. Furthermore, the close association of microbes in a densely populated gut ecosystem is ideal for HGT-based mechanisms to develop diverse populations.
References
Fishelson L, Montgomery WL, Myrberg A Jr. A unique symbiosis in the gut of tropical herbivorous surgeonfish (Acanthuridae: Teleostei) from the Red Sea. Science. 1985;229:49–51.
Clements KD, Sutton DC, Choat JH. Occurrence and characteristics of unusual protistan symbionts from surgeonfishes (Acanthuridae) of the Great Barrier Reef, Australia. Mar Biol. 1989;102:403–12.
Angert ER. Alternatives to binary fission in bacteria. Nat Rev Microbiol. 2005;3:214–24.
Flint JF, Drzymalski D, Montgomery WL, Southam G, Angert ER. Nocturnal production of endospores in natural populations of Epulopiscium-like surgeonfish symbionts. J Bacteriol. 2005;187:7460–70.
Meehan CJ, Beiko RG. A phylogenomic view of ecological specialization in the Lachnospiraceae, a family of digestive tract-associated bacteria. Genome Biol Evol. 2014;6:703–13.
Thompson CL, Vier R, Mikaelyan A, Wienemann T, Brune A. ‘Candidatus Arthromitus’ revised: segmented filamentous bacteria in arthropod guts are members of Lachnospiraceae. Environ Microbiol. 2012;14:1454–65.
Clements KD, Choat JH. Fermentation in tropical marine herbivorous fishes. Physiol Zool. 1995;68:355–78.
Choat JH, Clements KD, Robbins W. The trophic status of herbivorous fishes on coral reefs I. Dietary analysis. Mar Biol. 2002;140:613–23.
Choat JH, Robbins WD, Clements KD. The trophic status of herbivorous fishes on coral reefs: II. Food processing modes and trophodynamics. Mar Biol. 2004;145:445–54.
Ward RJ, Clements KD, Choat JH, Angert ER. Cytology of terminally differentiated Epulopiscium mother cells. DNA Cell Biol. 2009;28:57–64.
Hutchison E, Yager NA, Taw MN, Taylor M, Arroyo F, Sannino DR, et al. Developmental stage influences chromosome segregation patterns and arrangement in the extremely polyploid, giant bacterium Epulopiscium sp. type B. Mol Microbiol. 2018;107:68–80.
Angert ER, Clements KD. Initiation of intracellular offspring in Epulopiscium. Mol Microbiol. 2004;51:827–35.
Mendell JE, Clements KD, Choat JH, Angert ER. Extreme polyploidy in a large bacterium. Proc Natl Acad Sci USA. 2008;105:6730–4.
Miller DA, Suen G, Clements KD, Angert ER. The genomic basis for the evolution of a novel form of cellular reproduction in the bacterium Epulopiscium. BMC Genom. 2012;13:265.
Angert ER The enigmatic cytoarchitecture of Epulopiscium spp. In: Shively JM, editor. Complex intracellular structures in prokaryotes. Berlin Heidelberg: Springer-Verlag; 2006. p. 285–301.
Angert ER, Clements KD, Pace NR. The largest bacterium. Nature. 1993;362:239–41.
Miyake S, Ngugi DK, Stingl U. Phylogenetic diversity, distribution, and cophylogeny of giant bacteria (Epulopiscium) with their surgeonfish hosts in the Red Sea. Front Microbiol. 2016;7:285.
Collins MD, Lawson PA, Willems A, Cordoba JJ, Fernandez-Garayzabal J, Garcia’ P, et al. The phylogeny of the genus Clostridium: proposal of five new genera and eleven new species combinations. Int J Syst Bacteriol. 1994;44:812–26.
Angert ER, Losick RM. Propagation by sporulation in the guinea pig symbiont Metabacterium polyspora. Proc Natl Acad Sci USA. 1998;95:10218–23.
Angert ER. DNA replication and genomic architecture of very large bacteria. Annu Rev Microbiol. 2012;66:197–212.
Randall JE. Surgeonfishes of the world. Honolulu: Mutual Publishing; 2001.
Eble Ja, Rocha La, Craig MT, Bowen BW. Not all larvae stay close to home: insights into marine population connectivity with a focus on the brown surgeonfish (Acanthurus nigrofuscus). J Mar Biol. 2011;2011:1–12.
Clements KD. Fermentation and gastrointestinal microorganisms in fishes. In: Mackie RI, White BA, editors. Gastrointestinal microbiology. USA, Boston, MA: Springer; 1997. p 156–98. .
Wernegreen JJ. Genome evolution in bacterial endosymbionts of insects. Nat Rev Genet. 2002;3:850–61.
Russell SL, Corbett-Detig RB, Cavanaugh CM. Mixed transmission modes and dynamic genome evolution in an obligate animal–bacterial symbiosis. ISME J. 2017;11:1359–71.
Bright M, Bulgheresi S. A complex journey: transmission of microbial symbionts. Nat Rev Microbiol. 2010;8:218–30.
Hutchison EA, Miller DA, Angert ER. Sporulation in bacteria: beyond the standard model. Microbiol Spectr. 2014;2:87–102.
Vos M. Why do bacteria engage in homologous recombination? Trends Microbiol. 2009;17:226–32.
Librado P, Rozas J. DnaSPv5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics. 2009;25:1451–2.
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol. 2012;61:539–42.
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol. 2010;59:307–21.
Tavaré S. Some probabilistic and statistical problems in the analysis of DNA Sequences. Am Math Soc. Lect Math Life Sci. 1986;17:57–86.
Jukes T, Cantor C. Evolution of protein molecules. In: Munro H, editor. Mammalian protein metabolism. New York: Academic Press; 1969. p. 21–132. .
Tajima F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics. 1989;123:585–95.
Fu YX, Li WH. Statistical tests of neutrality of mutations. Genetics. 1993;133:693–709.
Fu YX. Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics. 1997;147:915–25.
Nei M, Gojobori T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol. 1986;3:418–26.
Jolley KA, Feil EJ, Chan MS, Maiden MC. Sequence type analysis and recombinational tests (START). Bioinformatics. 2001;17:1230–1.
Yang Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol. 2007;24:1586–91.
Kosakovsky Pond SL, Posada D, Gravenor MB, Woelk CH, Frost SDW. Automated phylogenetic detection of recombination using a genetic algorithm. Mol Biol Evol. 2006;23:1891–901.
Delport W, Poon AFY, Frost SDW, Kosakovsky Pond SL. Datamonkey 2010: a suite of phylogenetic analysis tools for evolutionary biology. Bioinformatics. 2010;26:2455–7.
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing. 2014. http://www.r-project.org.
Wickham H. Ggplot2: elegant graphics for data analysis. New York: Springer-Verlag; 2009.
Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics. 2000;155:945–59.
Bruen TC, Philippe H, Bryant D. A simple and robust statistical test for detecting the presence of recombination. Genetics. 2006;172:2665–81.
Huson DH, Bryant D. Application of phylogenetic networks in evolutionary studies. Mol Biol Evol. 2006;23:254–67.
Smith JM, Smith NH, O’Rourke M, Spratt BG. How clonal are bacteria? Proc Natl Acad Sci. 1993;90:4384–8.
Excoffier L, Smouse PE, Quattro JM. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics. 1992;131:479–91.
Excoffier L, Lischer HEL. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010;10:564–7.
Dray S, Dufour A-B. The ade4 package: implementing the duality diagram for ecologists. J Stat Softw. 2007;22:1–20.
Legendre P, Desdevises Y, Bazin E. A statistical test for host-parasite coevolution. Syst Biol. 2002;51:217–34.
Balbuena JA, Míguez-Lozano R, Blasco-Costa I. PACo: a novel procrustes application to cophylogenetic analysis. PLoS ONE. 2013;8:e61048.
Paradis E, Claude J, Strimmer K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics. 2004;20:289–90.
Oksanen J, Guillaume F, Kindt BRLP, Minchin PR, O’Hara RB, Simpson GL, et al. vegan: community ecology package. R package version 2.3-5. 2016. http://cran.r-project.org/package=vegan.
Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16:111–20.
Letunic I, Bork P. Interactive tree of life (iTOL)v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016;44:W242–5.
Ramirez-Soriano A, Ramos-Onsins SE, Rozas J, Calafell F, Navarro A. Statistical power analysis of neutrality tests under demographic expansions, contractions and bottlenecks with recombination. Genetics. 2008;179:555–67.
Choat JH, Axe LM. Growth and longevity in acanthurid fishes: an analysis of otolith increments. Mar Ecol Prog Ser. 1996;134:15–26.
McKnite AM, Perez-Munoz ME, Lu L, Williams EG, Brewer S, Andreux PA, et al. Murine gut microbiota is defined by host genetics and modulates variation of metabolic traits. PLoS ONE. 2012;7:e39191.
Miyake S, Ngugi DK, Stingl U. Diet strongly influences the gut microbiota of surgeonfishes. Mol Ecol. 2015;24:656–72.
Hooper LV, Littman DR, Macpherson AJ. Interactions between the microbiota and the immune system. Science. 2012;336:1268–73.
Gómez GD, Balcázar JL. A review on the interactions between gut microbiota and innate immunity of fish. FEMS Immunol Med Microbiol. 2008;52:145–54.
Smith CCR, Snowberg LK, Gregory Caporaso J, Knight R, Bolnick DI. Dietary input of microbes and host genetic variation shape among-population differences in stickleback gut microbiota. ISME J. 2015;9:2515–26.
Sullam KE, Essinger SD, Lozupone CA, O’Connor MP, Rosen GL, Knight R, et al. Environmental and ecological factors that shape the gut bacterial communities of fish: a meta-analysis. Mol Ecol. 2012;21:3363–78.
Wong S, Rawls JF. Intestinal microbiota composition in fishes is influenced by host ecology and environment. Mol Ecol. 2012;21:3100–2.
Kwong WK, Moran NA. Gut microbial communities of social bees. Nat Rev Microbiol. 2016;14:374–84.
Engel P, Stepanauskas R, Moran NA. Hidden diversity in honey bee gut symbionts detected by single-cell genomics. PLoS Genet. 2014;10:e1004596.
Ley RE, Peterson DA, Gordon JI. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell . 2006;124:837–48.
Sousa A, Ramiro RS, Barroso-Batista J, Güleresi D, Lourenço M, Gordo I. Recurrent reverse evolution maintains polymorphism after strong bottlenecks in commensal gut bacteria. Mol Biol Evol. 2017;34:2879–92.
Horne JB, van Herwerden L, Choat JH, Robertson DR. High population connectivity across the Indo-Pacific: congruent lack of phylogeographic structure in three reef fish congeners. Mol Phylogenet Evol. 2008;49:629–38.
Klanten OS, Choat JH, van Herwerden L. Extreme genetic diversity and temporal rather than spatial partitioning in a widely distributed coral reef fish. Mar Biol. 2007;150:659–70.
Lester SE, Ruttenberg BI. The relationship between pelagic larval duration and range size in tropical reef fishes: a synthetic analysis. Proc Biol Sci. 2005;272:585–91.
D’Aloia CC, Bogdanowicz SM, Harrison RG, Buston PM. Seascape continuity plays an important role in determining patterns of spatial genetic structure in a coral reef fish. Mol Ecol. 2014;23:2902–13.
Harrison HB, Williamson DH, Evans RD, Almany GR, Thorrold SR, Russ GR, et al. Larval export from marine reserves and the recruitment benefit for fish and fisheries. Curr Biol. 2012;22:1023–8.
Meyer CG, Holland KN. Movement patterns, home range size and habitat utilization of the bluespine unicornfish, Naso unicornis (Acanthuridae) in a Hawaiian marine reserve. Environ Biol Fishes. 2005;73:201–10.
Hardman E, Green JM, Sabrina Desiré M, Perrine S. Movement of sonically tagged bluespine unicornfish, Naso unicornis, in relation to marine reserve boundaries in Rodrigues, western Indian Ocean. Aquat Conserv Mar Freshw Ecosyst. 2010;20:357–61.
Chaguza C, Cornick JE, Everett DB. Mechanisms and impact of genetic recombination in the evolution of Streptococcus pneumoniae. Comput Struct Biotechnol J. 2015;13:241–7.
Cooper TF. Recombination speeds adaptation by reducing competition between beneficial mutations in populations of Escherichia coli. PLoS Biol. 2007;5:e225.
Takeuchi N, Kaneko K, Koonin EV. Horizontal gene transfer can rescue prokaryotes from Muller’s ratchet: benefit of DNA from dead cells and population subdivision. G3. 2014;4:325–39.
Hudson RR. Analytical results concerning linkage disequilibrium in models with genetic transformation and conjugation. J Evol Biol. 1994;7:535–48.
Spratt BG, Hanage WP, Feil EJ. The relative contributions of recombination and point mutation to the diversification of bacterial clones. Curr Opin Microbiol. 2001;4:602–6.
Lorenz MG, Wackernagel W. Bacterial gene transfer by natural genetic transformation in the environment. Microbiol Rev. 1994;58:563–602.
Erickson RJ, Copeland JC. Congression of unlinked markers and genetic mapping in the transformation of Bacillus subtilis 168. Genetics. 1973;73:13–21.
Mell JC, Lee JY, Firme M, Sinha S, Redfield RJ. Extensive cotransformation of natural variation into chromosomes of naturally competent Haemophilus influenzae. G3. 2014;4:717–31.
Dalia AB, McDonough E, Camilli A. Multiplex genome editing by natural transformation. Proc Natl Acad Sci. 2014;111:8937–42.
Maciver SK. Asexual amoebae escape Muller’s ratchet through polyploidy. Trends Parasitol. 2016;32:855–62.
Steinmoen H, Knutsen E, Håvarstein LS. Induction of natural competence in Streptococcus pneumoniae triggers lysis and DNA release from a subfraction of the cell population. Proc Natl Acad Sci Usa. 2002;99:7681–6.
Steinmoen H, Teigen A, Håvarstein LS. Competence-induced cells of Streptococcus pneumoniae lyse competence-deficient cells of the same strain during cocultivation. J Bacteriol. 2003;185:7176–83.
Zafra O, Lamprecht-Grandío M, de Figueras CG, González-Pastor JE. Extracellular DNA release by undomesticated Bacillus subtilis is regulated by early competence. PLoS ONE. 2012;7:e48716.
Soppa J. Polyploidy and community structure. Nat Microbiol. 2017;2:16261.
Browne HP, Forster SC, Anonye BO, Kumar N, Neville BA, Stares MD, et al. Culturing of ‘unculturable’ human microbiota reveals novel taxa and extensive sporulation. Nature. 2016;533:543–6.
Acknowledgements
We thank Rick Harrison and Lynn Johnson for their advice about study design and statistical analysis. We also thank Amy Dapper, Xavier Didelot, and Melanie Smee for advice on our population analyses. We thank Zanah Francis for help screening samples and David Sannino for insightful discussions. We are grateful to Daniel Buckley for the use of his server to run analyses. Fish were collected under the James Cook University Ethics approvals A503 and A1641 and with approval of the Great Barrier Reef Marine Park Authority permits G01-356 and G10-33239.1. This study was supported by National Science Foundation grants MCB1244378 and IOS1354911.
Data availability
The sequences obtained in this study have been deposited in GenBank under Accession No. MH259595-MH259695 (16S rRNA genes), MH268406-MH269196 (Epulopiscium sp. type B MLSA genes), and MH282951-MH283005 (N. tonganus MLSA genes).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Arroyo, F.A., Pawlowska, T.E., Choat, J.H. et al. Recombination contributes to population diversification in the polyploid intestinal symbiont Epulopiscium sp. type B. ISME J 13, 1084–1097 (2019). https://doi.org/10.1038/s41396-018-0339-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0339-y
This article is cited by
-
Host individual and gut location are more important in gut microbiota community composition than temporal variation in the marine herbivorous fish Kyphosus sydneyanus
BMC Microbiology (2023)
-
Host species and habitat shape fish-associated bacterial communities: phylosymbiosis between fish and their microbiome
Microbiome (2023)
-
Sources of variation in community composition of the hindgut microbiota in two tropical Kyphosus species
Coral Reefs (2022)
-
The gut microbiome and metabolome of Himalayan Griffons (Gyps himalayensis): insights into the adaptation to carrion-feeding habits in avian scavengers
Avian Research (2021)
-
Non-equivalent genomes in polyploid prokaryotes
Nature Microbiology (2021)