Abstract
Petroleum hydrocarbons reach the deep-sea following natural and anthropogenic factors. The process by which they enter deep-sea microbial food webs and impact the biogeochemical cycling of carbon and other elements is unclear. Hydrostatic pressure (HP) is a distinctive parameter of the deep sea, although rarely investigated. Whether HP alone affects the assembly and activity of oil-degrading communities remains to be resolved. Here we have demonstrated that hydrocarbon degradation in deep-sea microbial communities is lower at native HP (10 MPa, about 1000 m below sea surface level) than at ambient pressure. In long-term enrichments, increased HP selectively inhibited obligate hydrocarbon-degraders and downregulated the expression of beta-oxidation-related proteins (i.e., the main hydrocarbon-degradation pathway) resulting in low cell growth and CO2 production. Short-term experiments with HP-adapted synthetic communities confirmed this data, revealing a HP-dependent accumulation of citrate and dihydroxyacetone. Citrate accumulation suggests rates of aerobic oxidation of fatty acids in the TCA cycle were reduced. Dihydroxyacetone is connected to citrate through glycerol metabolism and glycolysis, both upregulated with increased HP. High degradation rates by obligate hydrocarbon-degraders may thus be unfavourable at increased HP, explaining their selective suppression. Through lab-scale cultivation, the present study is the first to highlight a link between impaired cell metabolism and microbial community assembly in hydrocarbon degradation at high HP. Overall, this data indicate that hydrocarbons fate differs substantially in surface waters as compared to deep-sea environments, with in situ low temperature and limited nutrients availability expected to further prolong hydrocarbons persistence at deep sea.
Similar content being viewed by others
Introduction
Every year more than 1000 million liters of petroleum hydrocarbons enter the sea via natural seeps or anthropogenic activities [1]. Many microorganisms use hydrocarbons as a carbon and energy source [2], with metabolism affected by the chemical nature of the hydrocarbon, electron acceptor availability and temperature [3]. Hydrostatic pressure (HP) has been a largely neglected factor so far, in spite of being a distinctive geophysical parameter of deep-sea environments [4]. At increasing depths, the greater amount of mass in the water column exerts a downward force from the sea surface which is equal to about 1 MPa every 100 m (thus, HP is about 10 MPa at 1000 m below sea surface level [bsl]). Microbial hydrocarbon degradation at natural seeps located up to 3500 m bsl generates sufficient biomass to feed invertebrate communities [5]. In hot, anaerobic and nutrient-limited (e.g., phosphate, sulphate) deep subsurface oil reservoirs, microbial hydrocarbon degradation can proceed on a geological timescale at the oil-water interface [6]. The consistent observation of microbial oil consumption in different deep-sea ecosystems suggests that this is a common process at increased HP.
Until recently, the main anthropogenic contribution to deep-sea oil contamination was linked to seepage from shipwrecks. There are about 9000 potentially polluting wrecks laying on seafloors worldwide up to 6000 m bsl, holding 3000–23,000 million liters of oil [7, 8]. A more accurate assessment of the pathways of spilled-oil in the deep sea was carried out after the Deepwater Horizon (DWH) oil well blowout (Gulf of Mexico, April 2010). The DWH spill was the largest marine oil spill in history and the first to originate underwater (at 1500 m bsl, ≈15 MPa) [9]. Research on microbial community composition and gene expression in deep-sea DWH samples indicated a response to petroleum hydrocarbons [10,11,12]. However, DWH deep-sea studies generally compared contaminated and uncontaminated samples from equivalent HPs. Environmental investigations comparing samples at different HPs along the water column cannot avoid temperature gradients, which complicates results interpretation. Microbial degradation represents the ultimate step for the clean-up of oil-contaminated environments, particularly for the poorly accessible deep-sea areas. The process by which petroleum hydrocarbons enter the deep-sea microbial food web and impact the carbon budget and other biogeochemical cycles remains unresolved. This knowledge gap hampers the development of bioremediation technologies to combat deep-sea spills. In particular, it is unclear whether HP alone affects the assembly of oil-degrading microbial communities and their metabolism.
In the present study, laboratory-scale cultivation was applied to selectively discriminate the role of the sole HP in shaping the physiology and ecology of microbial hydrocarbon degradation. Hydrocarbon-free, high HP-adapted surficial microbial communities in marine sediments collected from 1000 m bsl (≈10 MPa) were supplied with long-chain hydrocarbons as sole carbon source. Long-chain hydrocarbons were selected because they have a greater chance of reaching deep-sea environments (e.g., following offshore in situ burning, [13, 14]), where they persist longer than short-chain aliphatics [15]. Following enrichments in the pressure range 0.1–30 MPa, isolation was conducted to retrieve high HP-adapted microorganisms, which were tested further in synthetic communities.
Materials and methods
Sample collection
Sediment cores were collected at the West Iberian Margin (June 2–10, 2014, onboard the R/V Belgica) using a multicorer at 960 m bsl for C20 incubations (latitude 37°49’579; longitude 09°27’497), and at 955 m bsl for C30 incubations (latitude 37°58’849; longitude 09°23’353). The upper 2 cm of the sediment cores were used for experiments. Samples were kept at 4 °C and 10 MPa (in situ HP) using high HP reactors (HHPRs) until reaching the lab (17 days).
Microbial analyses
HHPRs
High HP incubations were conducted in stainless steel ISI 316 reactors (208 mL, maximum HP 60 MPa) (Nantong Feiyu Oil Science and Technology Exploitation, China). HP was delivered through a manual pump.
Microbial enrichments at different HP
The experimental set up is described in Fig. 1. Sediments were diluted 20% (w:v) with ONR7a medium [16], pH 7.5 ± 0.1. The n-alkane eicosane (C20) and triacontane (C30) (Sigma-Aldrich, Belgium) were supplied as sole carbon source 0.1% (w:v). The liquid phase was 200 mL, with 8 mL of gas phase. O2 was provided by injecting 2.5 MPa of air, subsequently increasing HP to 10 or 20 MPa by adding sterile medium. Two control reactors at ambient pressure (0.1 MPa) were set using different initial O2 supply. For high O2 levels, a Schott bottle was used (200 mL liquid phase, 950 mL of gas phase). For microaerophilic controls a HHPR was used (100 mL of liquid phase; 108 mL of gas phase). O2 supply in microaerophilic controls was thus 10- to 2-fold lower as compared to high O2 controls and HHPR, respectively. Two-negative controls were also prepared (marine sediment and no added carbon; and sterile ONR7a with either C20 or C30 but no sediment). These control reactors were tested at 0.1 MPa. Before any re-inoculation, reactors were washed with ethanol 70% (v:v, Sigma-Aldrich) and rinsed five times with autoclaved, milli-Q water. Media were autoclaved before use, but re-inoculation and incubation were not carried out aseptically. Reactors were incubated statically for 10 days at 20 °C. Afterwards, pressure was gently released to ambient levels, the culture diluted tenfold in fresh ONR7a medium and incubated again (nine consecutive incubations for a final enrichment of 3 months).
Experimental set up. Microbial communities from hydrocarbon-free, marine sediments collected at 1000 m below sea surface level (≈10 MPa) were enriched in a HP range 0.1–20 MPa, using either C20 or C30 as sole carbon source. Biomasses from enriched consortia were characterized, including that from a third high HP reactor at 30 MPa inoculated from 20 MPa reactors. Isolation from 10 and 20 MPa reactors supplied with C20 was conducted, yielding multispecies colonies. These were pooled and tested further in a high HP-adapted synthetic community (HHP-SC) at 0.1 and 10 MPa, using either C20 or acetate as sole carbon source
Isolation procedure and biomass characterization
Besides HHPRs operated at 10 and 20 MPa, with either hydrocarbon consortia enriched at 20 MPa were used as inoculum for new HHPRs operated at 30 MPa. All reactors were used for biomass characterization (i.e., PLFAs and amino acids). Isolation was attempted with HHPRs supplied with C20 at 10 and 20 MPa (Fig. 1) first at high HP and subsequently by plating (details provided in Supplementary Information).
Synthetic community experiments
Multispecies colonies from −80 °C glycerol stocks (20%, v:v) were thawed and cultivated axenically on either acetate or C20 in ONR7a medium using 150 mL glass Schott bottles (50 mL liquid phase), under aerobic, static conditions at 20 °C for 14 days. Two high HP-adapted synthetic communities were thus prepared: one grown on acetate and one on C20. Synthetic communities had equal carbon content (i.e., 0.8495 gC L−1 with either acetic acid or C20) and initial cell number (2 × 106 cells mL−1) and were incubated in triplicate at 0.1 or 10 MPa in glass Schott bottles or HHRPs, under the same conditions as for enrichments.
Bacterial counts
Cell concentrations were assessed by flow cytometry with SYBR green I staining [17]. Cells were diluted 1 × 103 and 1 × 104 with autoclaved, filtered ONR7a medium (0.22 µm, Sartorius, Belgium), and fractioned according to their size using glass microfibers filters of 1.5 and 25 µm (Sartorius), with the latter used for total cell number.
Molecular analyses
DNA extraction
Samples (2 mL) were centrifuged in a FastPrep tube (5 min, 13,000 rpm). Then, pellets were supplied with 200 mg glass beads (0.11 mm, Sartorius) and 1 mL lysis buffer (100 mM Tris, 100 mM EDTA, 100 mM NaCl, 1% polyvinylpyrrolidone [PVP40], 2% sodium dodecyl sulphate [SDS]; pH 8). Tubes were placed in a FastPrep device (MP Biomedicals, USA) (16,000 rpm, 40 s, 2 runs), centrifuged (10 min, maximum speed, 4 °C), the DNA extracted with phenol-chloroform and precipitated with ice-cold isopropyl alcohol and 3M sodium acetate (1 h, −20 °C). Isopropyl alcohol was removed by centrifugation (30 min, max speed), DNA pellets dried and resuspended in TE buffer (10 mM Tris, 1 mM EDTA) and stored at −20 °C. DNA sample quality was assessed using 1% (w:v) agarose (Life technologiesTM, Spain) gel-electrophoresis, and quantified by a fluorescence assay (QuantiFluor® dsDNA kit; Promega, USA) using a Glomax®-Multi+system (Promega). Samples were normalized to 1 ng µL−1 DNA and sent to LGC Genomics (Germany) for library preparation and sequencing using the Illumina Miseq platform (details provided in Supplementary Information).
Metagenome sequencing
DNA extracted from multispecies colonies was used for metagenome sequencing on an Illumina MiSeq platform. 16S rRNA genes were extracted from assembled metagenomic contigs, with contig coverage calculated to estimate relative abundance of each strain in each enrichment (details provided in Supplementary Information).
Metaproteome analysis
Culture samples (100 mL) were centrifuged, pellets dissolved in 400 µL 50 mM Tris/HCl (pH 6.8) and protein extracted with liquid phenol [18]. After protein quantification with amido black assay, 7 µg of proteins from the enrichments and 25 µg from the synthetic communities were loaded into a 12% SDS-PAGE. For enrichments, the SDS-PAGE was conducted after proteins entered ~5 mm into the separation gel, while for synthetic communities each lane was cut afterwards in ten equal fractions for LC–MS/MS measurements. The complete protein fraction was digested with trypsin, and peptides were measured by LC–MS/MS using an Elite Hybrid Ion Trap Orbitrap MS with a 120 min gradient. For protein identification, a database search with Mascot [19] was performed, using a false discovery rate of 1% (details provided in Supplementary Information). All MS results were submitted to PRIDE [20], accession number PXD004328.
Statistical analysis
Bars in the graphs indicate a 95% confidence interval (95% CI) calculated using a Student's t-test with a two-sided distribution. Statistical significance was assessed using a non-parametric test (Mann–Whitney test), which considered a two-sided distribution with 95% CI. Differential abundance analysis of 16S rRNA amplicon OTUs was conducted using ALDEx2 (v.1.10.0) [21, 22] on OTUs combined into families. High HP samples (10 or 20 MPa) were compared to ambient pressure controls (aerobic and microaerophilic). Families where the Wilcoxon signed-rank test yielded p < 0.05 were considered significantly differentially abundant in between the two conditions.
Chemical analyses
Dissolved O2 was measured with a probe by Hach (Belgium). pH was determined with a probe by Metrohm (Belgium). Phosphate and sulphate were quantified with a Compact Ion Chromatograph (Metrohm, Switzerland) equipped with a conductivity detector. Dissolved inorganic carbon was determined by gas chromatography (SRI 310C, USA) after adding 10% H3PO4 (Sigma‐Aldrich).
Intracellular metabolites
Samples were prepared with minor modifications according to ref. [23]. NMR spectra were obtained on Bruker spectrometers operating at 500 and 700 MHz (1H), processed using MestReNova (v.11.0.4, Mestrelab Reserch), and analyzed by pcaMethods package [24] using R (v.3.4.4) (details provided in Supplementary Information).
Results
High HP selects for small-sized cell cultures with high nutrient uptake, but low biomass yield and hydrocarbon-degradation capacity
Hydrocarbon-free surficial marine sediment collected at 1000 m bsl (≈10 MPa) was incubated at increased HP (10 or 20 MPa) under aerobic conditions, using control cultures at atmospheric pressure (0.1 MPa) under aerobic and microaerophilic conditions. Cultures were supplied with either C20 or C30 as sole carbon source and grown for 3 months under repeated batch conditions (9 incubations of 10 days each, 10% dilution; experimental set up in Fig. 1).
To assess microbial oil degradation capacity, the pH of test reactors was compared with two-negative controls, one without added carbon and another without marine sediment (Fig. 2). Provided that either C20 or C30 were supplied as sole carbon source, decreased pH values indicated high hydrocarbon degradation activity, as CO2 ionization in water generates HCO3− + H+. All test reactors showed a lower pH with respect to negative controls throughout the whole enrichment (p < 0.05, Fig. 2). After three repeated inoculations (i.e., 30 days) the marine sediment was completely washed out from all reactors, thus the potential contribution of microbes attached to the sediment was removed. In controls with no C20 or C30, this resulted in negligible cell counts (as assessed by flow cytometry; these controls were stopped after 50 days). HP reactors showed a lower acidification capacity compared to both control cultures at 0.1 MPa (Fig. 2). This was particularly evident in enrichments with C30, as after 70 days cultures at increased HP showed a loss-of-acidification capacity and could not decrease the pH below about 7.15 at any new batch incubation (Fig. 2). This was not reflected in any other physiological measurement, as enriching cultures were comparable between 30 and 90 days (Fig. 3). Cultures at high HP generally had a lower cell number as compared to ambient controls with either carbon source (Fig. 3a, b), and were characterized by an increasingly smaller size (as assessed by flow cytometry on 1.5-µm-filtered samples, Fig. 3c, d). Small-sized cells were not merely due to a physical constraint (if any) imposed by high HP, as cell counts were conducted 1–2 h after decompression. Irrespective of the HP applied, cultures supplied with C20 had lower pH values than those at C30, whereas the latter had a higher cell number.
pH decrease in enriching consortia supplied with C20 a or C30 b as sole carbon source at different HPs (0.1, 10, and 20 MPa) and in two-negative controls at 0.1 MPa. Enriching consortia at 0.1 MPa were tested under aerobic (+O2) or microaerophilic (−O2) conditions. Each data point represent a 10-day incubation period, after which cultures were diluted 10% (v:v) and incubated again for another 10 days
Final cell numbers a–d, O2 respiration e, f, and nutrients consumption g–j in enriching consortia supplied with either C20 or C30 as sole carbon source at different HPs (0.1, 10, and 20 MPa). Enriching consortia at 0.1 MPa were tested under aerobic (+O2, green) or microaerophilic (−O2, yellow) conditions. In a–d, cells were sorted for their size using sterile filters of 25 a, b and 1.5 μm c, d prior to injection into the flow cytometer for counting. Bars indicate a 95% confidence interval, with values considered between the 3rd and 9th incubation (n = 6). Keys: * indicates significant difference (p < 0.05) with respect to 0.1 MPa +O2
At ambient conditions, C20 and C30 are solid and solubilize in water at less than 2 µg L−1 [25]. Quantitative hydrocarbon degradation measurements would thus require extraction with solvents of the entire culture, preventing further culturing. However, of the 72 reactors tested through the enrichments, hydrocarbons were found in the water phase only in one case (20 MPa, 9th incubation, C20), indicating that hydrocarbon solubilisation was likely followed by rapid bacterial consumption.
HP did not significantly alter O2 respiration per cell with either carbon source (p > 0.05, Fig. 3e, f). Nonetheless, high HP stimulated SO42− reduction per cell, which was higher than aerobic controls and generally comparable to microaerophilic controls (p < 0.05; Fig. 3g, h). This must take into account that the total amount of O2 respired by the cells in high HP reactors was higher than in microaerophilic controls (p < 0.05, Fig. S1A,D). Finally, with either carbon source high HP enhanced PO43− consumption per cell as compared to both controls at ambient pressure (p < 0.05; Fig. 3i, j).
High HP inhibits specialized, hydrocarbonoclastic bacteria, and selects for generic, non-specific oil-degraders
Long-chain-hydrocarbon supply to pristine sediments resulted in a remarkable microbial succession (Fig. 4a). A shared response to increased HP with either hydrocarbon was the significant abundance of Desulfuromonadaceae as opposed to Oceanospirillaceae in ambient controls (p < 0.05, Table S1A,B; Fig. 4a). At high HP, supply of C20 also significantly enriched Halomonadaceae, Pseudoalteromonadaceae and Shewanellaceae, with Vibrionaceae significantly more enriched only with C30 (p < 0.05, Table S1A,B). A time course of the most enriched genera is presented in Table S2, S3 and Fig. S2. Some of the so-called obligate hydrocarbonoclastic bacteria (OHCB), a group of specialized marine microorganisms growing almost exclusively on oil [26], were present in our enrichments (e.g., Thalassolituus and Alcanivorax). While predominating in both controls at ambient pressure, OHCB were almost totally suppressed by high HP (Fig. 4b).
Relative 16S rRNA abundance of a bacterial families in the hydrocarbon-free, marine sediment used as inoculum (original HP ≈10 MPa) and in enriching consortia, and b OHCB genera detected in such enriching consortia, supplied with C20 (left) or C30 (right) as sole carbon source at different HPs (0.1, 10, and 20 MPa), as assessed by Illumina sequencing. Enriching consortia at 0.1 MPa were tested under aerobic (+O2) or microaerophilic (−O2) conditions
High HP downregulates β-oxidation and increases housekeeping proteins
High HP in enriched consortia (90 days) shaped cell metabolism as described by metaproteome analyses, particularly concerning housekeeping functions. Expression of proteins related to biological functions (UniProtKB keyword) such as ATP synthesis, ion and proton transport were upregulated with high HP as compared to ambient controls, while transcription was downregulated (p < 0.05, Fig. 5; Table S4). The increased importance of ions and protons transport may relate to the acidification following hydrocarbon oxidation, as cells at high HP might experience lower pH values due to increased CO2 solubility (16% equilibrium pressure increase every 10 MPa [27]) and facilitated ionization as compared to atmospheric pressure [28]. Among the low abundance proteins, high HP negatively impacted lipid degradation, fatty acid and lipid metabolism (p < 0.05, Fig. 5; Table S4). In particular, metaproteins related to fatty acids β-oxidation (EC: 4.2.1.17; 5.1.2.3; 5.3.3.8; 1.1.1.35; 2.3.1.9; 1.3.99.-) were remarkably downregulated at high HP with either carbon source (p < 0.025; log2 fold change [f.c.] −1.53 to −3.80; Table S5A,B). Mapped metaproteins comprised enzymes required for detoxification of radical O2 species, suggestive of an active hydrocarbon oxidation at high HP and aerobic controls at ambient pressure (Table S5). However, O2 stress at high HP was equal or lower than in aerobic controls at ambient pressure (Table S5A,B), indicating that the enhanced dissolved O2 levels imposed by a HP increase (14% equilibrium pressure increase every 10 MPa for O2 [27]) did not turn into a stress factor for the cultures. Nonetheless, several proteins for sulphite (SO32−) reduction in high HP enrichments confirmed that O2 respiration was followed by anaerobiosis and SO42− reduction with either carbon source (Fig. 3g, h). Finally, although proteins for alkane degradation were identified, the alkane 1-monooxygenase (EC: 1.14.15.3) responsible for terminal oxidation [29] was only detected with C30 in aerobic controls at ambient pressure (Table S6).
Heatmap of the high abundant metaproteins (top) and radar distribution of low abundant metaproteins (bottom) related to biological functions (UniProtKB keyword) in enriching consortia supplied with C20 (left) or C30 (right) as sole carbon source at different HPs (0.1, 10 and 20 MPa). Enriching consortia at 0.1 MPa were tested under aerobic (+O2) or microaerophilic (−O2) conditions. Numbers reported in the heatmap boxes (top) and in the radar graphs (bottom) represent the percentage of expressed metaprotein for a biological function relative to all detected metaproteins. Complete dataset reported in Table S4; all metaproteins in Table S6
High HP increases short and branched-chain PLFAs
With either hydrocarbon, cultures at 20 MPa were used to inoculate reactors at 30 MPa (Fig. 1), and HP impact on PLFA and amino acid profiles was investigated. High HP increased the relative abundance of shorter PLFAs, in particular C15 and C16 (p < 0.01, log2 f.c. 2.9–3.3; Fig. S3A,C). The relative content of iso-, anteiso- and in general total branched-chain PLFAs was remarkably higher at high HP as compared to both controls at ambient pressure, contrary to cyclopropane PLFAs (p < 0.015, log2 f.c. −0.9 to −3.1; Fig. S3B,D). High HP-enriched consortia accumulated i-C15:0, ai-C15:0, i-C16:0, C16:1ω7t, i-C17:1ω7c and two undetermined PLFAs (p < 0.05; Fig. S4A,B), whereas carrying less C18 monounsaturated PLFAs, especially C18:1ω9c (Fig. S4A,B). Amino acid profiles were not affected by HP except in consortia supplied with C30 ≥ 20 MPa (Table S7).
Isolation from HP-enriched consortia yields multispecies colonies of generic, non-specific oil-degrading bacteria
Isolation from enriched consortia at 10 and 20 MPa was attempted with C20 under aerobic conditions. Following dilution (up to 10−9), cultures were cultivated at their respective HP, then streaked on agar at ambient pressure. Colonies were generally no more than five per plate, possibly due to the reduced access to the solid C20 used as sole carbon source, and less than 1 mm in diameter. Each colony yielded metagenomes with two to five unique 16S rRNA gene sequences (Table S8), and thus represented a reduced complexity of the source community rather than a pure isolate. The assembly of such multispecies colonies did not differ when derived from either 10 or 20 MPa (Table S8). However, when considered together the 11 multispecies colonies retrieved from high HP-enriched consortia were formed by a core community of four frequently recurrent genera (Thalassospira, Vibrio, Halomonas and Pseudoalteromonas) which were among the most abundant at high HP (Table S2C,D) or whose family was significantly enriched at high HP (Halomonadaceae and Pseudoalteromonadaceae with C20, Table S1). None of these recurring genera was a specialized OHCB. On the contrary, the less frequent genera (e.g., Thalassolituus, Pseudomonas, Microbacterium) were neither abundant at high HP (Table S4C,D) nor belonged to families associated to high HP (Table S1). The reason for yielding multispecies colonies in place of individual species is unclear. The absence of Deltaproteobacteria is considered a consequence of adopting aerobic conditions for isolation.
High HP-adapted synthetic communities confirm a shift from OHCB to generic, non-specific oil-degraders with reduced hydrocarbon-degradation capacity
Multispecies colonies originated from the C20 consortia enriched at 10 and 20 MPa were used as inoculum to assemble a high HP-adapted synthetic community (HHP-SC), which was tested at 0.1 and 10 MPa using as sole carbon source either C20 or acetate as control (Fig. 1).
HHP-SCs performance in terms of acidification capacity and growth was comparable with that of enriched consortia selected under equivalent increased HPs (i.e., 10 MPa; p > 0.05, Fig. 6a, b). Thus, HHP-SCs reliably reproduced the hydrocarbon degradation capacity of HP enrichments even in the absence of isolated Deltaproteobacteria. Moreover, when such HHP-SCs were tested at ambient pressure their performance was lower as compared to consortia enriched in long-term experiments at 0.1 MPa (p < 0.05, Fig. 6a, b), which were largely dominated by the OHCB Thalassolituus (Fig. S2).
Physiological response of HHP-SCs as compared to long-term enrichments from which they derived when supplying C20 as sole carbon source at 0.1 and 10 MPa (n = 3) (a, pH value; b, cell number) and physiological response of HHP-SCs when supplied with C20 or acetate as sole carbon source at 0.1 and 10 MPa (n = 3) (c, final cell number; d, O2 respiration per cell; e, pH value; f, CO2 production per cell). Data for enrichments are that of the last three incubations (Figs. 2 and 3a, b). For convenience of comparison, HHP-SCs data when supplying C20 is reported twice (pH, a and e; cell number, b and c). Bars indicate the standard deviation from the mean. Keys reported in the graph
Increased HP reduced cell growth of HHP-SCs irrespective of the supplied carbon source (Fig. 6c), resulting in low total CO2 productions (Fig. S5) although HHP-SCs microbial community composition was not dramatically altered by the different conditions applied. Despite being tested under non-axenic conditions, HHP-SCs were represented by only eight main OTUs (99.0–99.4% of the total 16S rRNA sequences, Fig. 7), all originally present in the multispecies colonies used as inoculum (Fig. S6). In particular, HHP-SCs were constituted by a core community of four OTUs which were the most represented, especially in the test condition (i.e., 10 MPa, C20; 96.4%, Fig. 7). This core community was constituted by the four generic, non-specific oil-degrading genera Vibrio, Thalassospira, Halomonas and Pseudoalteromonas found to be frequently recurrent in multispecies colonies (Table S8). The OHCB T. oleiovorans (OTU00005, SSU_type8, Fig. S6) grew at ambient pressure in the presence of C20, but was inhibited at 10 MPa (16S rRNA abundance from 10.7 to 1.5%, log2 f.c. −2.8; Fig. 7), as occurred in enrichments (Fig. 4b).
High HP leads to intracellular citrate and dihydroxyacetone accumulation
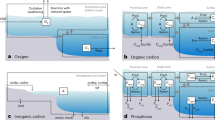
Cell metabolism in HHP-SCs was analysed. The full metaproteomes related to alkane activation and β-oxidation could be reconstructed, with the exception of the alkane 1-monooxygenase (EC: 1.14.15.3) responsible for terminal oxidation [29] (Fig. S7), as for enrichments (Table S6). A deeper analysis of the alkane-activation mechanism is proposed in the Supplementary Information. Incubation of HHP-SCs under ambient pressure did not restore high expression levels of β-oxidation-related proteins as compared to high HP (Table S9), nor did it with lipid and fatty acid metabolism, or lipid degradation-related metaproteins (p > 0.05; Table S10). However, high HP upregulated glycerol metabolism (log2 f.c. +0.76 [C20] and +0.67 [acetate], Table S10). The possibility that glycerol may have been used to produce biosurfactants [30], a group of glycolipids, phospholipids and lipoproteins enhancing the apparent solubility of oil in water, was not supported by surface tension and emulsification property analysis (Table S11). However, the entire metaproteome connecting glycerol metabolism to the tricarboxylic acid cycle (TCA) could be reconstructed (Fig. 8a). Two key intermediates of these pathways interconnected by few enzymatic reactions, namely citrate and dihydroxyacetone, were significantly accumulated in cells at high HP (Fig. 8b), in the frame of a general HP-dependent rearrangement of water and lipid-soluble intracellular metabolites (p < 0.05; Fig. 8c, Fig. S8–10).
Mapped metaproteins involved in glycerol metabolism linked to the TCA cycle in HHP-SCs supplied with C20 a. Selected intracellular metabolites from the aqueous phase b and intracellular metabolite profiles c in cells derived from HHP-SCs supplied with C20 or acetate as sole carbon source at 0.1 (shaded area) and 10 MPa. Analyses are normalized per cell content
Discussion
The use of increased HP in laboratory-scale experiments is an emerging approach to investigate deep-sea oil degradation [31,32,33,34,35,36,37,38]. Whereas the use of reactors may reduce microbial biodiversity to cultivable species, in the case of HP it allows to simulate a poorly accessible environment, such as the deep sea. Existing literature indicates that enhanced HP affects bacterial hydrocarbon consumption, however it does not explain the relationship between HP, microbial community assembly, and hydrocarbon degradation capacity. Whether oil-degradation pathways differ between surface and deep sea remains unclear. In the present study, we used pristine marine sediment microbial communities natively adapted to 10 MPa (i.e., 1000 m bsl) to decipher the sole effect of HP on microbial hydrocarbon metabolism independent of other parameters that may differ along the water column. HPs up to 30 MPa (3000 m bsl) were tested on either (1) enriched consortia or (2) synthetic communities adapted to high HP (HHP-SCs). The first approach tested the impact of HP on the long-term selection of different microbial community members, whereas the second focused on the short-term impact of HP on the metabolism of comparable communities already adapted to high HP.
Long-term selection of high HP in enriching oil-degrading consortia
The supply of long-chain hydrocarbons (either C20 or C30) to pristine sediments resulted in a HP-dependent restructuring of microbial communities (Fig. 4a). The OHCB Thalassolituus and Alcanivorax that largely predominated in ambient pressure controls (irrespective of O2 availability) were suppressed by a HP increase to only 10 MPa (Fig. 4b). As HP application enhances gas solubilisation [27], cells ≥10 MPa might have experienced higher dissolved O2 levels, which potentially influenced hydrocarbon metabolism and microbial community assembly. However, O2 respiration per cell was not impacted by high HP (Fig. 3e, f) and proteins related to O2 stress were equally expressed in aerobic controls and high HP-enriched consortia (Table S5).
High HP increasingly selected for small-sized (Fig. 3c, d), slow-growing (Fig. 3a, b) cells. Although not impacting cell respiration, at high HP anaerobiosis was established during each 10-day incubation, prompted SO42− reduction (Fig. 3g, h; Table S5) and stimulated the enrichment of unique Deltaproteobacteria (i.e., Desulfuromonadaceae; Table S1, S2, S3; Fig. S2) as compared to microaerophilic controls at ambient pressure. The observed shift in PLFA profiles reflected these changes in microbial community assembly. In fact, the PLFA composition at ambient pressure resembled that of obligate oil-degraders [39,40,41], whereas the increase in uneven branched-chain PLFAs under high HP (particularly i17:1ω7c) probably mirrored the increase in Deltaproteobacteria [42,43,44]. In particular, high HP selected for consortia remarkably enriched in branched-chain PLFAs (Fig. S3), a typical response of HP-tolerant microbes [45].
High HP-enriched consortia were characterized by low expression levels of β-oxidation-related proteins (Fig. 5; Table S5), consistent with previous findings by members of the present group on transcript levels of two axenic Alcanivorax species inhibited by 10 MPa while supplied with n-dodecane [36, 37]. β-oxidation represents the main metabolic pathway for hydrocarbon degradation following their activation [2]. A reduced protein expression level does not imply per se that a pathway is not operating, rather that it plays a less relevant role under the tested conditions. When compared to ambient controls, high HP-enriched consortia were featured by increased expression levels of housekeeping proteins, particularly basic cellular functions, such as ATP synthesis and ion transport including hydrogen (Fig. 5, Table S4). pH homeostasis is based on a H+-ATPase and is influenced by CO2 production [28], and hydration and ionization of CO2 is facilitated at increased HP as it entails-negative volume changes [28]. Thus, maintenance of pH homeostasis in acidifying cultures oxidizing hydrocarbons may represent a critical function at increased HP. In bacteria, high HP increases membrane permeability and inactivates pH-maintaining enzymes [46,47,48]. Permeability to ions impairs proton-driven forces used by several pumps (e.g., Na/K ATPase, [49]), a correlation being observed between ion pump activity and HP [50]. A similar molecular response (i.e., impacted ATP synthesis and expression of Na+-translocating reductases) was reported in the transcriptomic studies on the two axenic Alcanivorax species supplied with n-dodecane and inhibited at 10 MPa [36, 37]. Sustained hydrocarbons oxidation at increased HP may thus entail increased cell maintenance, rendering high β-oxidation levels less favourable.
Short-term effect of high HP on hydrocarbon metabolism in synthetic communities
Isolation from high HP-enriched consortia yielded multispecies colonies where a core community of high HP-adapted genera was associated with satellite microorganisms not related to HP. Among the latter, the OHCB T. oleiovorans was detected (Table S8). When tested in HHP-SCs with C20, T. oleiovorans could grow at ambient pressure but was inhibited by a HP increase to only 10 MPa (Fig. 7). This was consistent with the enrichment findings (Fig. 4b) and extends to Thalassolituus the so-called “Alcanivorax paradox” hypothesis proposed by some of the present authors [51]. This notes that in both field and lab-scale experiments so far OHCB appear to be affected by increased HPs [4]. Variation in temperature, salinity, electron acceptor availability, hydrocarbons and possibly pH in concomitance with increased HP is expected to refine this observation. For instance, the increased relative abundance of the OHCB genus Cycloclasticus [26] was correlated with the enrichment of aromatic hydrocarbons in underwater oil plumes during the DWH [52], however experimental evidence with lab-scale HP tests is missing. Recent findings report that a close relative of the OHCB, psychrophilic Oleispira antarctica RB-8 could grow in DWH deep-seawater samples in short-term (32 days), lab-scale tests when high HP was applied in combination with low temperature (0.1–30 MPa, 4 °C) using Macondo oil [53]. O. antartica predominated at all HPs, its selective advantage likely being dependent on the low temperature concomitantly applied (it was isolated from Antarctic coastal water [54]). Interestingly, its relative abundance slightly decreased with increasing HP (81–65%, 0.1–30 MPa). No other OHCB was detected, contrary to several generic, non-specific oil-degraders, many of which consistent with the present study (e.g., Vibrio, Photobacterium and Marinifilum; Table S2, S3; [53]). Another prominent effect of high HP in combination with low temperature was to reduce growth [53], as reported earlier [31]. In the present study, this occurred only by increasing HP (Fig. 3a, b; Fig. 6c). In particular, comparable HHP-SCs (Fig. 7) underwent a fivefold decrease in growth yields when increasing HP to only 10 MPa. The latter was consistent with the intracellular accumulation of citrate and dihydroxyacetone (Fig. 8b), two metabolic intermediates linked by few reactions whose enzymes were detected by metaproteomics (Fig. 8a). Citrate is the key intermediate of the TCA cycle, an aerobic process involved in the final steps of fatty acids (and carbohydrates) oxidation generating NADH for use e.g. in ATP synthesis, a biological function upregulated at increased HP in long-term enrichments (Fig. 5). Citrate HP-dependent accumulation suggests a reduction of TCA cycle rates, apparently related with the accumulation of dihydroxyacetone. The significant upregulation of the biological functions glycerol metabolism and glycolysis interconnecting these two metabolic intermediates supports this hypothesis (Tab. S10). The possibility that dihydroxyacetone represents a novel piezolyte, i.e., a solute whose biosynthesis is triggered by HP increases [55], cannot be discarded. The reason for reduced TCA cycle rates under increased HP is unclear. Aconitase and isocitrate dehydrogenase, the enzymes using citrate and its product in the TCA cycle, can be completely inhibited after only 15 min at HPs 5–15 times greater than what applied in the present study [56]. Whether their catalytic activity is partially inhibited at 10 MPa needs further investigation.
In conclusion, the present dataset reports the first comprehensive overview describing how HP shapes the physiology and ecology of microbial hydrocarbons degradation. A reduced capacity to conduct the final steps of fatty acids oxidation (i.e., TCA cycle) would decrease β-oxidation levels, resulting in low cell growth and hydrocarbon mineralization. The selective advantage of OHCB to sustain high hydrocarbon degradation rates would thus be prevented at high HP, allowing generic, non-specific oil-degraders to thrive. In fact, reduced hydrocarbon oxidation in high HP reactors occurred notwithstanding the availability of electron acceptors, with enhanced HP requiring a higher expenditure for maintenance of cell homeostasis (e.g., ATP synthesis and ion transport), which involved cell membrane composition (enriched in branched-chain PLFAs). The interplay between TCA cycle, ATP synthesis, pH homeostasis and hydrocarbon oxidation at deep-sea HP should be investigated further. This is particularly relevant to assess the fate of hydrocarbons entering the deep sea following anthropogenic spills, where degradation of the overabundant hydrocarbon input is further reduced by low temperature and lack of nutrients.
Data accessibility analysis
Raw sequencing reads for 16S rRNA genes in cultures enriched at ambient and increased HP up to 20 MPa were submitted to the NCBI, accession number PRJNA506445. Raw sequencing reads for the 11 multispecies colony metagenomes were submitted to the NCBI Sequence Read Archive under sequential accession numbers from SRR7825327 to SRR7825337. The 16 full-length 16S rRNA gene sequences assembled from this data were submitted to GenBank under sequential accession numbers from MH894278 to MH894293.
References
Maribus. World Ocean Review 3. 2014.
Rojo F. Degradation of alkanes by bacteria: minireview. Environ Microbiol. 2009;11:2477–90.
Head IM, Jones DM, Röling WFM. Marine microorganisms make a meal of oil. Nat Rev Microbiol. 2006;4:173–82.
Scoma A, Yakimov MM, Boon N. Challenging oil bioremediation at deep-sea hydrostatic pressure. Front Microbiol. 2016;7:1203.
Jørgensen BB, Boetius A. Feast and famine—microbial life in the deep-sea bed. Nat Rev Microbiol. 2007;5:770–81.
Head IM, Jones DM, Larter SR. Biological activity in the deep subsurface and the origin of heavy oil. Nature. 2003;426:344–52.
NOAA. Oil spill case histories 1967-91. Seattle, Washington; NOAA, Hazardous Materials Response and Assessment Division 1992.
Michel J, Gilbert T, Etkin DS. Potentially polluting wrecks in marine waters. An Issue Paper Prepared for the 2005 International Oil Spill Conference. 2005.
Federal Interagency Solutions Group. Oil Budget Calculator Science and Engineering team 2010. Oil budget Calculator Technical Documentation. 2010:1–49.
Kimes NE, Callaghan AV, Suflita JM, Morris PJ. Microbial transformation of the deepwater horizon oil spill-past, present, and future perspectives. Front Microbiol. 2014;5:603.
Joye SB, Teske AP, Kostka JE. Microbial dynamics following the macondo oil well blowout across gulf of Mexico environments. Bioscience. 2014;64:766–77.
King GM, Kostka JE, Hazen TC, Sobecky PA. Microbial responses to the deepwater horizon oil spill: from coastal wetlands to the deep sea. Ann Rev Mar Sci. 2015;7:15.1–15.25.
Buist I, Trudel K, Morrison J, Aurand D. Laboratory studies of the properties of in-situ burn residues. 1997 Int Oil Spill Conf. 1997;1997:149–56. https://doi.org/10.7901/2169-3358-1997-1-149
Jézéquel R, Simon R, Pirot V. Development of a burning bench dedicated to In Situ Burning Study: assessment of oil nature and weathering effect. In: Proceedings of the Thirty-seventh AMOP Technical Seminar on Environmental Contamination and Response, Environment Canada, Ottawa, ON, 2014. 555–566.
Bagby SC, Reddy CM, Aeppli C, Fisher GB, Valentine DL. Persistence and biodegradation of oil at the ocean floor following Deepwater Horizon. Proc Natl Acad Sci USA. 2017;114:E9–E18. https://doi.org/10.1073/pnas.1610110114
Dyksterhouse SE, Gray JP, Herwig RP, Lara JC, Staley JT. Cycloclasticus pugetii gen. nov., sp. nov., an aromatic hydrocarbon-degrading bacterium from marine sediments. Int J Syst Bacteriol. 1995;45:116–23.
De Roy K, Clement L, Thas O, Wang Y, Boon N. Flow cytometry for fast microbial community fingerprinting. Water Res. 2012;46:907–19.
Heyer R, Kohrs F, Benndorf D, Rapp E, Kausmann R, Heiermann M, et al. Metaproteome analysis of the microbial communities in agricultural biogas plants. N Biotechnol. 2013;30:614–22.
Perkins DN, Pappin DJC, Creasy DM, Cottrell JS. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis. 1999;20:3551–67.
Vizcaíno JA, Côté RG, Csordas A, Dianes JA, Fabregat A, Foster JM, et al. The Proteomics Identifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res. 2013;41:D1063–9.
Fernandes AD, Macklaim JM, Linn TG, Reid G, Gloor GB. ANOVA-Like Differential Expression (ALDEx) analysis for mixed population RNA-Seq. PLoS ONE. 2013;8:e67019.
Fernandes AD, Reid JN, Macklaim JM, McMurrough TA, Edgell DR, Gloor GB. Unifying the analysis of high-throughput sequencing datasets: characterizing RNA-seq, 16S rRNA gene sequencing and selective growth experiments by compositional data analysis. Microbiome. 2014;2:15.
Tremaroli V, Workentine ML, Weljie AM, Vogel HJ, Ceri H, Viti C, et al. Metabolomic investigation of the bacterial response to a metal challenge. Appl Environ Microbiol. 2009;75:719–28.
Stacklies W, Redestig H, Scholz M, Walther D, Selbig J. pcaMethods - A bioconductor package providing PCA methods for incomplete data. Bioinformatics. 2007;23:1164–7.
MacKay D, Shiu WY. A critical review of Henry’s law constants for chemicals of environmental interest. J Phys Chem. 1981;10:1175–99.
Yakimov MM, Timmis KN, Golyshin PN. Obligate oil-degrading marine bacteria. Curr Opin Biotechnol. 2007;18:257–66.
Enns T, Scholander PF, Bradstreet ED. Effect of hydrostatic pressure on gases dissolved in water. J Phys Chem. 1965;69:389–91.
Abe F, Horikoshi K. Analysis of intracellular pH in the yeast Saccharomyces cerevisiae under elevated hydrostatic pressure: A study in baro- (piezo-) physiology. Extremophiles. 1998;2:223–8.
Ji Y, Mao G, Wang Y, Bartlam M. Structural insights into diversity and n-alkane biodegradation mechanisms of alkane hydroxylases. Front Microbiol. 2013;4:58.
Dobler L, Vilela LF, Almeida RV, Neves BC. Rhamnolipids in perspective: gene regulatory pathways, metabolic engineering, production and technological forecasting. N Biotechnol. 2016;33:123–35.
Schwarz JR, Walder JD, Colwell RR. Deep-sea bacteria: growth and utilization of n-hexadecane at in situ temperature and pressure. Can J Microbiol. 1975;21:682–7.
Schwarz JR, Walder JD, Colwell RR. Deep-sea bacteria: growth and utilization of hydrocarbons at ambient and in situ pressure. Appl Microbiol. 1974;28:982–6.
Grossi V, Yakimov MM, Ali BAl, Tapilatu Y, Cuny P, Goutx M, et al. Hydrostatic pressure affects membrane and storage lipid compositions of the piezotolerant hydrocarbon-degrading Marinobacter hydrocarbonoclasticus strain #5. Environ Microbiol. 2010;12:2020–33.
Schedler M, Hiessl R, Valladares Juárez AG, Gust G, Müller R. Effect of high pressure on hydrocarbon-degrading bacteria. AMB Express. 2014;4:77.
Fasca H, de Castilho LVA, de Castilho JFM, Pasqualino IP, Alvarez VM, de Azevedo Jurelevicius D, et al. Response of marine bacteria to oil contamination and to high pressure and low temperature deep sea conditions. Microbiologyopen. 2018;7:e00550.
Scoma A, Barbato M, Borin S, Daffonchio D, Boon N. An impaired metabolic response to hydrostatic pressure explains Alcanivorax borkumensis recorded distribution in the deep marine water column. Sci Rep. 2016;6:31316.
Scoma A, Barbato M, Hernandez-Sanabria E, Mapelli F, Daffonchio D, Borin S, et al. Microbial oil-degradation under mild hydrostatic pressure (10 MPa): which pathways are impacted in piezosensitive hydrocarbonoclastic bacteria? Sci Rep. 2016;6:23526.
Scoma A, Boon N. Osmotic stress confers enhanced cell integrity to hydrostatic pressure but impairs growth in Alcanivorax borkumensis SK2. Front Microbiol. 2016;7:729.
Yakimov MM, Giuliano L, Denaro R, Crisafi E, Chernikova TN, Abraham WR, et al. Thalassolituus oleivorans gen. nov., sp. nov., a novel marine bacterium that obligately utilizes hydrocarbons. Int J Syst Evol Microbiol. 2004;54:141–8.
Yakimov MM, Golyshin PN, Lang S, Moore ERB, Abraham W, Lunsdorf H, et al. Alcanivorax borkurnensis gen. now, sp. nov., a new, hydrocarbon-degrading and surfactant-producing marine bacterium. Int J Syst Bacteriol. 1998;48:339–48.
Liu C, Shao Z. Alcanivorax dieselolei sp. nov., a novel alkane-degrading bacterium isolated from sea water and deep-sea sediment. Int J Syst Evol Microbiol. 2005;55:1181–6.
Taylor J, Parkes RJ. The cellular fatty acids of the sulphate-reducing bacteria, Desulfobacter sp., Desulfobulbus sp. and Desulfovibrio desulfuricans. Microbiology. 1983;129:3303–9.
Vainshtein M, Hippe H, Kroppenstedt RM. Cellular fatty acid composition of desulfovibrio species and its use in classification of sulfate-reducing bacteria. Syst Appl Microbiol. 1992;5:554–66.
Kohring LL, Ringelberg DB, Devereux R, Stahl DA, Mittelman MW, White DC. Comparison of phylogenetic relationships based on phospholipid fatty acid profiles and ribosomal RNA sequence similarities among dissimilatory sulfate-reducing bacteria. FEMS Microbiol Lett. 1994;119:303–8.
Wang F, Xiao X, Ou H-Y, Gai Y, Wang F. Role and regulation of fatty acid biosynthesis in the response of shewanella piezotolerans WP3 to different temperatures and pressures. J Bacteriol. 2009;191:2574–84.
Tholozan JL, Ritz M, Jugiau F, Federighi M, Tissier JP. Physiological effects of high hydrostatic pressure treatments on Listeria monocytogenes and Salmonella typhimurium. J Appl Microbiol. 2000;88:202–12.
Molina-Gutierrez A, Stippl, VolkerDelgado A, Gänzle MG, Vogel RF, Ga MG. In situ determination of the intracellular pH of Lactococcus lactis and Lactobacillus plantarum during pressure treatment. Appl Environ Microbiol. 2002;68:4399–406.
Molina-Höppner A, Doster W, Vogel RF, Gänzle MG. Protective effect of sucrose and sodium chloride for Lactococcus lactis during sublethal and lethal high-pressure treatments. Appl Environ Microbiol. 2004;70:2013–20.
Chong PLG, Fortes PAG, Jameson DM. Mechanisms of inhibition of (Na,K)-ATPase by hydrostatic pressure studied with fluorescent probes. J Biol Chem. 1985;260:14484–90.
Smelt JPPM, Rijke AGF, Hayhurst A. Possible mechanism of high pressure inactivation of microorganisms. High Press Res. 1994;12:199–203.
Mapelli F, Scoma A, Michoud G, Aulenta F, Boon N, Borin S, et al. Biotechnologies for marine oil spill cleanup: indissoluble ties with microorganisms. Trends Biotechnol. 2017;35:860–70.
Dubinsky EA, Conrad ME, Chakraborty R, Bill M, Borglin SE, Hollibaugh JT, et al. Succession of hydrocarbon-degrading bacteria in the aftermath of the deepwater horizon oil spill in the gulf of Mexico. Environ Sci Technol. 2013;47:10860–7.
Marietou A, Chastain R, Beulig F, Scoma A, Hazen TC, Bartlett DH. The effect of hydrostatic pressure on enrichments of hydrocarbon degrading microbes from the Gulf of Mexico following the deepwater Horizon oil spill. Front Microbiol. 2018;9:808.
Yakimov MM, Giuliano L, Gentile G, Crisafi E, Chernikova TN, Abraham WR, et al. Oleispira antarctica gen. nov., sp. nov., a novel hydrocarbonoclastic marine bacterium isolated from Antarctic coastal sea water. Int J Syst Evol Microbiol. 2003;53:779–85.
Martin DD, Bartlett DH, Roberts MF. Solute accumulation in the deep-sea bacterium Photobacterium profundum. Extremophiles. 2002;6:507–14.
Simpson RK, Gilmour A. The effect of high hydrostatic pressure on the activity of intracellular enzymes of Listeria monocytogenes. Lett Appl Microbiol. 1997;25:48–53.
Acknowledgements
These findings were financially supported by the FP7-EU project Kill Spill (312139), the Geconcentreerde Onderzoeksactie, Ghent University (BOF15/GOA/006), the Danish Ministry of Higher Education and Science (AU-2010-612-181) and The Novonordisk Foundation (NNF16OC0021110). F.-M.K. was supported by the Inter-University Attraction Pole ‘μ-manager’ (BELSPO, P7/25). A.S. thanks Dr. Ann Vanreusel (Ghent University, Belgium) for her supervision during deep-sea sampling. Dr. Xiao Xiang and Yu Zhang (Shanghai Jiao Tong University, China) are acknowledged for their assistance with high-pressure reactors. A. Bastian (Otto von Guericke University of Magdeburg) and Katrine Bay Jensen (Aarhus University) are acknowledged for their technical assistance.
Author contributions
A.S. conceived, designed and performed the experiments, and wrote the manuscript. R.H. performed the metaproteome analyses and co-wrote the manuscript. C.D. performed the metaproteome analyses. R.R. performed the experiments at high pressure and isolation of the micro-colonies. F.-M.K. performed the 16S rRNA analyses. I.M.B. performed the statistical analysis and the sequencing of the isolates. A.M. co-wrote the manuscript. P.V. analysed the amino acid data. H.B. and F.M. analysed the PLFA data. I.M.B. performed surfactants analysis and general editing. K.M. and T.V. analysed intracellular compounds. D.B. performed the metaproteome data analysis. U.R. supervised the metaproteome analysis. N.B. funded and supervised the project. All authors reviewed the manuscripts.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Scoma, A., Heyer, R., Rifai, R. et al. Reduced TCA cycle rates at high hydrostatic pressure hinder hydrocarbon degradation and obligate oil degraders in natural, deep-sea microbial communities. ISME J 13, 1004–1018 (2019). https://doi.org/10.1038/s41396-018-0324-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0324-5
This article is cited by
-
Response mechanism of Vibrio parahaemolyticus at high pressure revealed by transcriptomic analysis
Applied Microbiology and Biotechnology (2022)