Abstract
Despite the significance of biogenic methane generation in coal beds, there has never been a systematic long-term evaluation of the ecological response to biostimulation for enhanced methanogenesis in situ. Biostimulation tests in a gas-free coal seam were analysed over 1.5 years encompassing methane production, cell abundance, planktonic and surface associated community composition and chemical parameters of the coal formation water. Evidence is presented that sulfate reducing bacteria are energy limited whilst methanogenic archaea are nutrient limited. Methane production was highest in a nutrient amended well after an oxic preincubation phase to enhance coal biofragmentation (calcium peroxide amendment). Compound-specific isotope analyses indicated the predominance of acetoclastic methanogenesis. Acetoclastic methanogenic archaea of the Methanosaeta and Methanosarcina genera increased with methane concentration. Acetate was the main precursor for methanogenesis, however more acetate was consumed than methane produced in an acetate amended well. DNA stable isotope probing showed incorporation of 13C-labelled acetate into methanogenic archaea, Geobacter species and sulfate reducing bacteria. Community characterisation of coal surfaces confirmed that methanogenic archaea make up a substantial proportion of coal associated biofilm communities. Ultimately, methane production from a gas-free subbituminous coal seam was stimulated despite high concentrations of sulfate and sulfate-reducing bacteria in the coal formation water. These findings provide a new conceptual framework for understanding the coal reservoir biosphere.
Similar content being viewed by others
Introduction
Gas produced by biological processes (biogas) has a large role to play in meeting rising energy needs worldwide [1]. Coal-bed methane (CBM) is a relatively untapped energy source representing up to 20% of the world’s biogas resources [2,3,4] and is contained in actively mined and abandoned coal reservoirs globally [5]. The increased utilization of this unconventional natural gas source can significantly reduce the environmental drawbacks of coal-fired power plants [6, 7] prompting commercial enterprises to focus on the enhancement of biogenic methane production. To date, only incremental improvements of in situ biogas yields from coal have been observed [5, 8].
Biogenic methane is the product of ongoing microbial coal degradation whereas thermogenic methane is formed during the coalification process [9]. Mixed biogenic (δ13C value less than −60‰) and thermogenic (δ13C value greater than −50‰) signatures [10, 11] have been observed in many coal reservoirs worldwide [5]. The yield of biogenic methane is dependent on the degree of coal biodegradation and the availability of metabolic products generated from bacterial activity such as methanol, H2 and acetate used by methanogenic archaea to produce methane [12,13,14,15,16,17,18,19].
Biogenic methanogenesis in coal reservoirs is slow in situ and needs to be stimulated to be economically viable [12]. Nitrogen (N) and phosphorus (P) are essential nutrients for microbial metabolism and are limiting in many methanogenic environments [20, 21], thus reducing the effectiveness of biogas production. The applications of multi-nutrient amendments (N, P, vitamins, trace elements) or methanogenic substrates (acetate and/or CO2/H2) has been trialled in field-scale pilot tests but are not cost-effective for commercial implementation, and the use of compounds that can be used by methanogens can result in the enhancement of coal-independent methanogenesis [22].
One successful strategy in the bioremediation of groundwater contaminants is the application of passive oxygen-releasing material, e.g. calcium peroxide that releases oxygen upon contact with water (2CaO2 (s) + 2H2O - > 2Ca(OH)2 + O2 (g) [23]. The oxidation of coal or aerobic hydrocarbon degradation processes can drive the release of soluble organics into the coal formation water which can then be further biodegraded by microorganisms into methanogenic substrates [13, 17]. Despite the practical significance of in situ stimulation of the microbial communities in coal reservoirs, there has been no systematic long-term study of how nutrient and oxygen treatments alter microbial communities over the production life time of a coal gas well. Earlier in situ amendments were applied on productive methane-generating coal sites or non-productive lignite reservoirs under low or depleted sulfate concentrations ( < 1 mM; [24, 25]). Waters rich in sulfate (2–5 mM) occur in many coalbed aquifers [24] but are generally not found in association with methane given sulfate reducing bacteria (SRB) can outcompete methanogenic archaea for substrates such as acetate [26].
Additional work is required to answer several important practical and fundamental questions which are significant in the context of treatments for enhanced methane production of coal reservoir microbiomes. Can an indigenous microbial community in sulfate-rich coal formation water be stimulated by the addition of basic nutrients ammonium (N) and phosphate (P)? Can an initial oxic incubation phase in addition to nutrient amendment reshape the coal-reservoir microbial community in favour of methane generation? Are there systematic changes in microbial community composition and ecological function driven by treatment strategies? What are the metabolic drivers of methane generation? The answers to these questions have important implications for the enhancement of methane generation from productive and non-productive coal reservoirs for gas enterprises worldwide. They also offer insights into fundamental aspects of the coal reservoir microbiology such as the effects of high sulfate concentration on the methane production regime, the ability of methanogenic communities to overcome intermittent oxygen intrusion and the effects of basic nutrient limitation on the coal associated subsurface microbiome.
A field trial to stimulate methane production was carried out in a gas-free sulfate-rich subbituminous coal-seam reservoir located in the western coalfields of New South Wales, Australia. Our aim was to assess the impact of basic nutrient addition and an oxic/anoxic treatment regime on the biogeochemistry and microbiology of the coal formation water. A system free of biogas enabled us to clearly ascribe biogenic methane production to treatment effects through exclusion of treatment effects on release of pre-existing methane. Four coal gas wells extending 80 m below ground into a 3 m thick subbituminous coal seam (Fig. 1a, E) were used to assess three treatment strategies. In the first treatment, the nutrients ammonium chloride (N) and potassium phosphate (P) were applied to approximately 450 L formation water in contact with the coal seam. In the second treatment, in addition to nutrients calcium peroxide was provided during the first three months to initiate oxic conditions in the formation water, and subsequently removed to initiate an anoxic phase. The third treatment was a positive control amended with the methanogenic substrate sodium acetate (20 mM) and nutrients. A fourth well was left untreated as a negative control for comparison.
Materials and methods
Gas well setting and amendments
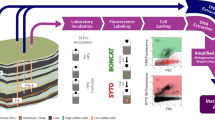
To stimulate the in situ biological conversion of coal to methane gas, four wells (bore holes) were drilled to 80 m depth, reaching 3 m into a subbituminous coal seam of the Sydney Basin at the Lithgow State Coal Mine, NSW, Australia (Fig. 1). All wells were lined with an iron casing and secured by backfilling with concrete. Wells were developed over 12 months by removing groundwater on a monthly basis and allowing them to refill (approx. 500 L influx per 24 h) to ensure the concrete mineralogy had no impact on the microbiology. The well diameter was 158 mm and the total well volume was 1588 L (1138 L well headspace and 450 L coal formation water – equilibrated with the aquifer in which the seam is submerged). The well head contained 3 gastight apertures for gas sampling. The field trial was conducted for 15–18 months starting in July 2011. The formation water submerged the coal and the lower 20 m of each well. The water temperature was 16 °C with a pH range of 7.8–8.7. Prior to the field trial, sodium bromide (Unilab Ajax, Australia) was added to the well water and monitored for decreases over time. Decreases were commensurate with the removal of water volume during sampling. This confirmed that the coal seam is poorly permeable (no hydraulic conductivity or water flow through the coal seam) and that there was no connectivity between the wells. To assess the impact of nutrient addition and aerobic/anaerobic treatment on the microbial community, water chemistry and methane production, the following treatments were applied: Well 1, nutrients + acetate (positive control); well 2, nutrients and calcium peroxide for oxygen release in the initial 3 months, well 3, nutrients only, and well 4, no amendment (negative control). Nutrients included 1.8 mM NH4Cl (Univar Ajax, Australia) and 1.9 mM Na2HPO4, (Unilab Ajax, Australia). Acetate (20 mM; Sigma Aldrich, Australia) was added to assess if the in situ conditions were generally suitable for methane production. The oxygenic treatment involved the addition of 100 mg calcium peroxide (Sigma Aldrich, Australia) contained in a fine mesh inside a permeable PVC canister, connected to the bottom of a larger permeable PVC canister containing coal pieces of the size of ~3 cm3 per coal piece taken from the original coal cores and placed in the formation water such that coal pieces could be retrieved for microbial community characterisation and the peroxide treatment could be terminated by removal of the connected canister (Fig. 1). The water sampling was accomplished using a bladder pump (PVC 3/8in discharge, ThermoFisher Scientific, Australia) deployed with a stainless steel drop tube (ThermoFisher, Australia) and operated at a low flow refill ratio of 40:20 to avoid the removal of dissolved gases in the formation water. The drop tube was lowered such that the bladder pump drew groundwater samples from the aqueous volume within the coal seam in each well. The pump and the connecting tubes were drained and flushed between wells to avoid the carriage of water between well sampling. Formation water used for chemical (anions, cations) and microbiological (cell counts) analyses were sampled monthly over a period of 15–18 months. Samples were immediately processed in the laboratory as outlined below.
Coal and water chemical analyses
The coal composition was analysed at the Bureau Veritas Australia Pty Lt, Cardiff, NSW, Australia. Scanning electron microscopy (SEM) was performed according to Hazrin-Chong, Manefield [27]. Coal formation water was filtered through a 0.2 μm syringe filter (Millipore, Australia) and aliquots for cation analyses were subsequently acidified with formic acid (10% v/v; Sigma-Aldrich, Australia) to a pH < 2 and stored at −20 °C until further processing. Anion (F, Cl, Br, NO2, NO3, PO4, SO4) and cation (B, Ca, Fe, K, Mg, Mn, Na, P, S, Si) concentrations were analysed using ion chromatography (IC) with conductivity detection and inductive coupled plasma optical emission spectroscopy (ICP-OES) at the Mark Wainwright Centre (UNSW, Sydney, Australia). The instrument detection limits were 0.02 mg/L for B, Si and Mn, 0.2 mg/L for Fe and P, 0.5 mg/L for Ca, K, Mg, Na and S, 2 mg/L for F, NO2, NO3 and Br, and 5 mg/L for Cl and PO4.
Oxidation-reduction potential (ORP), pH and temperature analyses
ORP, pH and temperature of the formation water were monitored monthly over a period of 15–18 months using an EcoSense pH100A meter connected to a pH/Temperature and ORP (Ag/AgCl) probe (YSI, Xylem, Australia). The formation water was pumped out of the well using a bladder pump (Grundfos Pumps Pty. Ltd, Australia). The water was collected in a sampling cell and probes were submerged into the water. An average of 10 readings were taken after every refill of the pump and until temperature equilibration.
Total methane, acetate and sulfate analyses
Total methane (headspace and dissolved), acetate and sulfate concentrations were monitored monthly. Gas samples were taken from the well head apertures and were transferred directly into 10 mL gastight serum vials using a gastight glass syringe until further processed for CH4 and stable isotopes analyses (Intertek Geotech, Australia) as per Whiticar [28]. Dissolved methane was analysed according to Kampbell and Vandegrift [29]. Methane was analysed using a Shimadzu GC-2010 gas chromatograph with flame ionization detection (GC-FID) fitted with a GASPRO PLOT column (60 × 0.32 mm; Agilent Technologies, Australia). The carrier gas was helium (3 mL min−1), inlet temperature was 250 °C. The oven temperature program was isothermal 100 °C (1 min) and then 25 °C min−1 to 250 °C and held for 1 min. Gas samples of 100 μL were withdrawn directly from the sampling flasks using a pressure lockable gastight glass syringe (SGE Analytical Science, Australia) and injected into the GC. To describe the gas distribution throughout the well headspace columns samples were taken from the top of the well and 10 m and 30 m below ground level. Methane concentrations displayed a linear trend with increasing concentration. This linear relationship was used throughout the trial to calculate methane concentrations in the columns using the methane concentration in samples taken from the top of the wells. For the analyses of acetate and sulfate concentrations, water was filtered through a 0.2 μm syringe filter (Millipore, Australia) and subsequently 900 µL filtered water was acidified with 100 µL 10% v/v formic acid (Sigma-Aldrich, Australia) to a pH < 2. Acetate (1 µL) was analysed by GC-FID (Shimadzu, Australia) using a DB-FFAP column (30 m x 0.32 mm; Agilent Technologies, Australia). Carrier gas, helium (1 mL min−1). Injection port 250 °C, split mode (1:30). Oven temperature: 60 °C for 1 min and then 15 °C min−1 to 250 °C. Sulfate was measured by ion chromatography with conductivity detection (Mark Wainwright Centre, UNSW, Australia). Carbon dioxide was not monitored as the seam rests in a mineral carbonate subsurface matrix.
Cell counts
Coal formation water samples were immediately fixed by the addition of glutaric dialdehyde (0.2 μm filtered, 2% final concentration; Sigma Aldrich, Australia) and stored at 4 °C in the dark. Prior to analysis, an aliquot was diluted 1000 fold in particle-free phosphate-buffered saline (0.9 g of NaCl, sodium phosphate buffer 15 mM, pH 7.4, 0.2 μm filtered), thoroughly shaken and transferred to a microscopic slide that was treated with a mounting medium (9.6% polyvinylalcohol 4–88 moviol (Sigma-Aldrich, Australia) in 24% glycerol (Sigma-Aldrich, Australia) prior to applications. Cells were stained using SybrGreen I (Invitrogen, Australia), and counting was performed using a BX51 epifluorescence microscopy (Olympus, Australia) according to Lunau et al. [30].
DNA extraction
Coal formation water (3–20 L) was filtered through a 0.1 μm Supor®polyethersolfone (PES) membrane disc filter (Pall Corporation, Port Washington, NY) that was subsequently cut into quarters and stored in a mixture of 5 mL RNA later® Stabilization Solution (Life Technologies, Carlsbad, CA, USA) and 5 mL 1 × TE buffer (10 mM Tris-HCL, 0.1 mM EDTA; pH 8) at −20 °C until further processing. DNA was extracted from one quarter of the filter using phenol–chloroform extraction as described by Lueders et al. [31]. Subsequently, the DNA was precipitated using polyethylene glycol 6000 (Sigma Aldrich, Australia), and the DNA pellet was washed once with 70% (v/v) ethanol and resuspended in 50 μL nuclease free water (Qiagen, Australia). DNA concentration and purity were determined by agarose gel electrophoresis and fluorometrically using RiboGreen (Qubit Assay Kit, Invitrogen, Australia) according to the manufacturer’s instructions. The extracted DNA was used as target for quantitative PCR (qPCR), stable isotope probing (SIP) and 16 S rRNA gene sequencing.
Quantification of archaeal and bacterial 16S rRNA and functional genes
Quantitative PCR was used to determine the abundances of bacteria, archaea, methanogenic archaea and sulfate reducing bacteria in the coal formation water. Standards were prepared using the pGEM®-T Easy Vector System (Promega, Madison, WI, US) and 16 S rRNA PCR products of Desulfovibrio vulgaris (DSM 644) and Methanococcoides burtonii (DSM 6242, DSMZ Germany). Plasmid inserts were verified by Sanger sequencing (Ramaciotti Centre, UNSW, Australia) using the T7 promotor primer (5’-TAATACGACTCACTATAGGG-3’). DNA targets were measured in four different dilutions of DNA extracts (1:10, 1:50, 1:100, 1:500) and in triplicate. Primer sets specific for the different phylogenetic domains and functional genes were used according to [14, 16]. The qPCR reaction mixtures of 25 μL contained 12.5 μL of the premix solution of the iQ SybrGreen qPCR Kit (Biorad, Hercules, CA, US), 8 μL of PCR-grade water, 1.5 μL of each primer (final concentration 0.28 μM) and 2 μL of template DNA (10 ng). The qPCR was performed using a C1000 Thermal cycler with an CFX96 Real Time System (Biorad, Australia). The specificity of the reaction was confirmed by melting curve analysis and agarose gel electrophoresis to identify unspecific products. Amplification efficiencies for all reactions ranged from 96.3 to 110.5% with an r2 value of > 0.99 for standard curve regression. Cell abundances were calculated by accounting for the volume of formation water sampled, the volume eluted after DNA extractions, dilutions prior to qPCR and gene copy numbers.
QPCR cell abundance normalization
To compensate for the variation in DNA recovery from each sample, qPCR measurements of the 16 S rRNA gene, mcrA gene and dsrA gene in each sample were normalized with a DNA recovery ratio [32]. The DNA recovery ratio was calculated for each sample by comparing the cell number calculated from the DNA yield against the cell count by microscopy according to Eq. 1.
Where for each sample i, R is the DNA recovery ration, Y is the DNA yield measured by Qubit (g/mL), N is the microscopy cell count (cells/mL), and D is the mean mass of DNA per cell (g). In this study, D = 7.5 × 10−15 g, as calculated from previously published measurements of groundwater samples [33,34,35]. Each qPCR measurement was then normalized with the respective DNA recovery ratio and gene copy number as per Eq. 2.
Where for each sample i, Ĉ is the normalized abundance (cells/mL), C is the qPCR measured abundance (cells/mL), and R is the DNA recovery ratio. For each target gene j, P is the copy number per cell (copies/cell). Average 16 S rRNA copy numbers were used (4.02 for bacteria and 1.63 for archaea according to the rrnDB [36, 37]. The mcrA and dsrA copy numbers were assumed to be 1 copy per cell.
To calculate the abundance of the different microbial genera involved in methanogenesis and sulfate reduction, qPCR of the mcrA and dsrA genes were combined with the sequencing data of the archaeal and bacterial 16 S rRNA gene, respectively (Eq. 3).
Where for each genus k, A is the calculated absolute abundance (cells/mL), Ã is the relative abundance measured by 16 S rRNA gene sequencing. For each sample i, Ĉ is the functional gene (mcrA or dsrA) normalized abundance (cells/mL).
16S rRNA gene sequencing
The 16 S rRNA gene of the community DNA from the well sample was sequenced using a 454-FLX sequencer and universal primers 926 F (5’-AAACTYAAAKGAATTGACGG-3’) and 1392 R (5’-ACGGGCGGTGTGRC-3’) for the V6-V8 region as described by Matsuki et al. [38] at the Hawkesbury Institute for the Environment, Sydney, Australia. Sequences were filtered to exclude low-quality reads, primer/barcode regions were trimmed and flowgrams were denoised using MOTHUR (http://www.mothur.org/; [39, 40]). Reads shorter than 250 bp were removed. Chimeras were checked using the UCHIME algorithm [41]. Sequences were classified taxonomically with a confidence threshold of 80% using MOTHUR-formatted SILVA training sets (v.199; https://www.arb-silva.de; [42]).
DNA-stable isotope probing (SIP)
Coal formation water (90 mL) was transferred anoxically into 120 mL nitrogen degassed serum bottles supplemented with 10 mM 12C- or fully 13C-labelled acetate (99 atom% 13C, Sigma Aldrich, Australia) and incubated in the dark at 20 °C. All incubations were carried out in triplicates and sampled for DNA extraction and stable isotope analyses (DNA-SIP) after 0, 3, and 6 days of incubation. DNA was extracted from 5 mL of culture slurry according to Lueders et al. [31]. Three parallel extractions were carried out, and extracts were pooled for each incubation treatment. DNA was checked by agarose gel electrophoresis and quantified fluorometrically using RiboGreen (Qubit Assay Kit, Invitrogen, Australia) according to the manufacturer’s instructions. Gradient preparation, isopycnic centrifugation, and gradient fractionation were performed as described by Lueders et al. [43]. DNA from each gradient fraction was quantified fluorometrically and by qPCR using archaeal and bacterial primer systems as according to Beckmann et al. [16].
Community analysis of density fractions
Amplicon libraries were generated from the DNA of the density gradient fractions by following Illumina’s 16 S Metagenomic Sequencing Library Preparation Protocol, using 12.5 ng of template DNA per reaction. PCR cycles for the initial PCR were reduced to 21 to avoid PCR biases from over-amplification. The following universal primer pair was used for the initial amplification, consisting of an Illumina-specific overhang sequence and a locus specific sequence:
926F_Illum(5’TCGTCGGCAGCGTCAGATGTGTATAAGAGACAG[AAACTYAAAKGAATTGRCCG]-3’), 1392R_Illum(5’GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAG[ACGGGCGGTGTGTRC]-3’).
This universal primer pair targets the V6–V8 hypervariable regions of the 16 S Ribosomal RNA gene and has been shown to capture the microbial diversity of bacteria and archaea in a single reaction [44]. PCR products were purified using magnetic bead capture with the Agencourt AMpure XP kit (Beckman Coulter, Australia) and quantified using a fluorometric kit (RiboGreen, Qubit Assay Kit, Invitrogen, Australia) according to the manufacturer’s instructions. Purified amplicons were multiplex sequenced using the MiSeq platform (Ramaciotti Centre for Genomics, UNSW, Australia) according to the manufacturer’s instructions. Paired end sequences were joined using FastqJoin (http://code.google.com/p/ea-utils; Erik [45]). Illumina libraries were quality filtered to truncate reads at positions with Phred scores < Q20 retaining reads > 75 bp and with < 3 low quality bases and no N characters using MOTHUR (http://www.mothur.org/) according to Schloss et al. [39] and Kozich et al. [46]. Chimeras were detected and removed [41] and taxonomy was assigned against the SILVA Database (v.119; https://www.arb-silva.de; [42]).
Statistical analysis
Statistical analyses of the datasets were carried out according to Clarke [47] and Clarke and Warwick [48], using Primer V6 [49] and XLSTAT (AddinSoft, Paris, France).
Results
Biogeochemical characteristics of the western coalfield reservoir
Composition and geochemical characteristics of the coal formation water and the coal from all 4 wells used at the trial site in the Western Coalfields of NSW, Australia were analysed (Table 1, Table S1). The coal was of subbituminous rank [50] and core samples from within the solid seam yielded no detectable microorganisms or quantifiable or PCR amplifiable microbial DNA (Fig. 1d). The formation water had a low salinity (0.013 ± 0.007 ppt) and sulfate concentrations of 265 ± 105 mg/L. No significant sources of N (ammonium, nitrate, nitrite) or P (phosphate) were observed in the formation water (Table S1). The coal formation water was characterized by low biomass of 3.2 × 104 cells/mL (n = 5; SD = 1.4 × 104). The Oxidation-Reduction potential (ORP, Ag/AgCl) was suitable for methanogenic activity ranging from −134 to −305 mV with a pH range of 7.8 to 8.7 and a water temperature of 16 °C. Calcium peroxide treatment had a clear impact on the redox potential (Figure S1). The oxic phase of the formation water was characterized by an elevated ORP of −141 ± 82 mV for the first 3 months compared to the untreated and nutrient treated well with an ORP of −253 ± 32 mV and −245 ± 40 mV, respectively. After the removal of CaO2, the ORP decreased to −287 ± 66 mV initiating the anoxic phase for the following 15 months.
Methane production and cell concentrations
Methane production (sum total of methane in the well headspace and formation water) was detected in all nutrient amended wells (N + P, N + P + CaO2, N + P + Acetate), showing a distinct one or two phase exponential increase in methane concentration (Fig. 2a). The highest methane yields were observed in a positive control well amended with the methanogenic substrate acetate (Fig. 2a, b). Acetate (final concentration 10 mM) was added at the start of the field trial and after 3 months to a final concentration of 20 mM in the coal formation water. Two phases were apparent in early biomass proliferation up to 7 months with a 3.5 order of magnitude increase in cell numbers, then declining almost 1 order of magnitude for 2 months until a second proliferation from 10 to 18 months (Fig. 2c). Both phases of methane generation were linked to acetate consumption (Fig. 2b).
(a) Methane formation, (b) Acetate concentration, (c) Cell numbers, and (d) sulfate concentrations in all four in situ treatments over an incubation time of 18 months. Addition of nutrients and acetate (red squares), nutrients and calcium peroxide (blue squares), nutrients (green squares) and no amendment (black squares)
The second highest methane formation yields were observed in the nutrient plus CaO2 amended well (Fig. 2a). After the removal of CaO2, methane increased over 5-fold peaking at 12 months. Cell concentrations started to increase after the removal of the CaO2 from 105 to 107 cells/mL and decreased with the decline of methane (Fig. 2a, c). In the well amended with nutrients only, methane increased from 12–15 months (Fig. 2a) with an 10-fold increase in cell numbers (Fig. 2c). The stable carbon and hydrogen isotopic analyses of the methane in all wells exhibited light carbon isotopic signatures with ∂13C values ranging from −55 to −77 ‰ relative to VPDB (Vienna Peedee Belemnite) indicating a biogenic origin of methane and with ∂D-CH4 of −300 ‰ consistent with acetoclastic methanogenesis ([11, 28]; Table 1) indicating that acetate was the main precursor of methane formation in all treatments.
Microbial community composition in the coal formation water
DNA extraction from the coal formation water was performed every three months and DNA subjected to PCR amplification, 16S rRNA gene sequencing and quantitative PCR of 16S rRNA and functional genes (mcrA and dsrA) to determine the microbial community composition and abundance in all wells over an incubation time of 15–18 months. In addition, qPCR copy numbers were normalised to total cell counts as determined by microscopy. All nutrient amended well treatments altered the archaeal and bacterial community composition significantly in the coal formation water in favour of methane production, directly by methanogens and indirectly by potential hydrocarbon degrading bacteria, especially sulfate reducing bacteria (Figs. 2a, 3).
Cell numbers of archaea (white bars), methanogenic archaea (light grey bars), bacteria (dark grey bars) and sulfate reducing bacteria (grey bars) based on ribosomal RNA and the functional mcrA and dsrA genes determined by quantitative qPCR. Changes of absolute abundances of methanogens (coloured squares) and sulfate reducing bacteria (coloured circles) over an incubation time of 15–18 months
In response to the nutrient amendments, methanogenic archaea increased from < 1 to up to 20% of the microbial community composition based on 16S rRNA gene sequencing and mcrA gene qPCR data (Figs. 3, 4a). Methanogenic archaea detected in the coal formation water belonged to the orders Methanosarcinales (Methanosarcina sp., Methanosaeta sp., Methanolobus sp. Methanococcoides sp.), Methanomicrobiales (Methanoregula sp., Methanoculleus sp.) and Methanobacteriales (Methanobacterium sp.) and their abundance increased significantly up to 3-4 orders of magnitude in the nutrient amended wells with an initial oxic phase or the addition of acetate (106 and 107 cells/mL respectively; T-test, p = 0.023). The genera Methanosarcina, Methanosaeta, Methanoregula and Methanolobus were principally responsible for this increase with Methanobacterium, Methanococcoides and Methanoculleus species contributing to a later methane production phase (Fig. 3). Low methanogenic cell numbers ( < 103 cells/mL) were observed in the untreated control well congruent with infrequent detection of Methanosaeta and Methanosarcina species (Figs. 3, 4).
a Changes in the bacterial and archaeal community composition in the coal formation water in response to different nutrient amendments over an incubation time of 15 months based on 16 S rRNA gene sequencing. From left to right: nutrients and acetate (positive control), nutrients and calcium peroxide, nutrients and no amendment (negative control). Colors indicate members of different phyla. Size of the bubble represents relative abundance of the genus or family in each sample. b Principal component analysis (PCA) comparing four different wells amended with nutrients and acetate (triangles), nutrients and calcium peroxide (circles), nutrients (squares) and no amendment (diamonds) over an incubation time of 15 months (white = 0 months to black = 15 months) and the showing the relationship to the different phyla (Bubbles are color coded as in a and methane production
The nutrient-stimulated microbial communities were significantly distinct from the untreated control (ANOSIM, p < 0.003) that harboured the indigenous community of the coal formation water. The indigenous community was dominated by bacterial members of the Beta- and Deltaproteobacteria, the Bacteroidetes/Chlorobi Group, the Firmicutes, the Actinobacteria and Fibrobacteres/Acidobacteria Group (Fig. 4a, b). In nutrient amended wells a decrease in relative abundance of members affiliated with the Bacteroidetes/Chlorobi group (mainly Unclassified Chitinophagaceae; explaining 19.16–28.08% of the dissimilarity compared to the untreated control well; SIMPER analysis) and the Actinobacteria (mainly Rhodococcus sp.; 9.46–10.02% dissimilarity; SIMPER analysis) was observed along with an increase in Deltaproteobacteria (mainly Desulfovibrio, Geobacter, Syntrophobacter, Pelobacter sp.; 16.18–18.93% of the dissimilarity; SIMPER analysis) and methanogenic archaea (mainly Methanosarcina, Methanosaeta, Methanoregula, Methanolobus, Methanobacterium sp.; 12.34–21.32% of the dissimilarity, SIMPER analysis). A higher dissimilarity of the microbial communities was observed between the untreated well and the nutrient plus acetate treated well (72.89% dissimilarity, SIMPER analysis) or the nutrient and calcium peroxide amended well (71.32% dissimilarity, SIMPER analysis). In comparison, the dissimilarity between the untreated well and the well only amended with nutrients (56.62% dissimilarity, SIMPER analysis) was lower (Fig. 4b). Nutrient amendments with acetate or calcium peroxide resulted in significantly different microbial community compositions compared to the nutrient amended well (ANOSIM, p = 0.029 and p = 0.034) whereas communities amended with acetate and calcium peroxide were not significantly different (ANOSIM, p = 0.07).
Sulfate reducing lineages, mainly Deltaproteobacteria and Archaeoglobales were detected in all wells and increased in abundance along with the dsrA gene (104 to 108 cells/mL) in the nutrient amended wells throughout the field trial (Fig. 3, 4a). Highest SRB abundance and diversity were observed in the acetate plus nutrient amended well compared to the well only amended with nutrients (Fig. 3). The sulfate reducing communities were dominated by Desulfovibrio spp. and members of the Desulfobacteraceae, Desulfovibrionaeceae and Desulfobulbaceae families (Fig. 3). The calcium peroxide treatment resulted in a decrease in SRB abundance and diversity (Desulfovibrio sp. being an exception) but recovered when calcium peroxide was removed and the anoxic incubation phase initiated (Figs. 3, 4a).
Microbial community composition on the coal surface
In response to the nutrient amendments, methanogenic archaea as well as bacteria were detected at the surface of the subbituminous coal after 3 months of the field trial (Fig. 5). The coal obtained from the untreated well and at the start of the field trial yielded no detectable microorganisms or microbial DNA. After 3 months of incubation, methanogenic archaea were detected at the coal surface of all amendments and belonged to the orders Methanosarcinales (Methanosarcina sp.), Methanobacteriales (Methanobacterium sp.) and Methanomicrobiales (Candidatus Methanoregula, Methanoculleus sp.). Their relative abundance increased from 0 to 37% after the addition of acetate and from 0 to 19% after the calcium peroxide induced oxic phase ceased compared to an increase from 0 to 12% when only amended with nutrients. Bacterial lineages within the phyla Alpha-Deltaproteobacteria, Actinobacteria and Firmicutes associated with sulfate- and sulfur reduction and hydrocarbon oxidation predominated.
Changes in the bacterial and archaeal community composition on the coal surface in response to different nutrient amendments over an incubation time of 15 months based on 16 S rRNA gene sequencing. From left to right: nutrients and acetate (positive control), nutrients and calcium peroxide, nutrients and no amendment (negative control). Colours indicate members of different phyla. Size of the bubble represents relative abundance of the genus or family in each sample
Sulfate reducing bacteria (SRB) within the orders Desulfovibrionales and Desulfobacterales increased in relative abundance after the addition of acetate (0 to 11%) and in the calcium peroxide treatment (0 to 19%) after the anoxic phase was initiated. Both treatments showed a high relative abundance of Geobacter uraniireducens (up to 14%) attached to the coal surface.
Microbial metabolic potential
DNA stable isotope probing (DNA-SIP) was used to track the incorporation of the methanogenic substrate acetate by bacteria and archaea in the coal formation water of the nutrient and acetate amended well (Fig. 6). Formation water sampled at the end of the field trial (18 months; Fig. 2a) was incubated with 13C2-acetate for 10 days. Labelled and unlabelled acetate pulses (10 mM) were consumed in 10 days (Fig. 6c). Methane increased after 4 days of incubation and continued to accumulate up to the conclusion of the incubation (Fig. 6c).
a, b Distribution and relative abundances of bacteria and archaea significantly associated with acetate utilisation in the ‘heavy’ and ‘light’ fractions of CsCl gradients derived from samples incubated in the presence of either 13C-labelled acetate or 12C-labelled acetate. c Acetate consumption and methane production in 13C- and 12C-labelled acetate amendments of coal formation water from the nutrient and acetate amended well (13C-acetate = black squares, 12C-acetate = black circles, 13C-methane = white squares, 12C-methane = white circles). DNA samples were taken from the 13C- and 12C-acetate enrichments during the time course of acetate consumption (red circles)
SSU 16S rRNA gene sequences from gradient fractions ranging in buoyant density from heavily labelled (1.764 g mL−1) to unlabelled (1.667 g mL−1) were compared within 13C-acetate-incubated samples and 12C-acetate-incubated controls (Fig. 6a). After 3 and 6 days, strong 13C dependent community shifts within the ‘light’ and ‘heavy’ gradient fractions were observed. Although many species were detected in both the ‘heavy’ and the ‘light’ gradient fractions, there was a clear distribution trend in the relative abundance of species (Fig. 6b). The early incorporation of 13C-label was predominantly detected in bacteria closely related to Geobacter uraniireducens, Geobacter pickeringii, Geobacter hydrogenophilus, Desulfovibrio vulgaris and Desulfobacca acetoxidans (Deltaproteobacteria) after day 3 in response to labelled acetate exposure (Fig. 6a, b). Organisms that incorporated carbon from acetate during the time course of 6 days belonged to the Azospira sp., Pseudomonas moraviensis (Betaproteobacteria) and Desulfovibrio dechloracetivorans (Deltaproteobacteria). Acetoclastic methanogenic archaea related to Methanosarcina acetivorans and Methanosaeta concilii incorporated 13C-acetate after 6 days when 65% acetate was consumed (Fig. 6b, c). Additionally, a non-methanogenic member within the phylum Euryarchaeota affiliated to Archaeoglobus lithotrophicus became highly enriched in the ‘heavy’ DNA fraction after 6 days (Fig. 6b). Archaeoglobus species are known as hyperthermophilic archaea found in high-temperature oil reservoirs potentially mediating sulfate reduction and hydrocarbon degradation [51,52,53]. Recently 16S rRNA sequences similar to those of Archaeaoglobus species were detected in two moderate-temperature coal reservoirs [54, 55].
Discussion
This study is the first to observe long-term (1.5 years) succession of microbial communities in a coal seam in response to biostimulation for biogas production. It was conducted in a gas-free sulfate-rich subbituminous coal seam resulting in confidence that all gas produced resulted from the stimulatory amendments applied and that biogas production is possible in the presence of significant quantities of sulfate. Basic nutrients (ammonium and phosphate) and the oxygen releasing reagent calcium peroxide (CaO2) were used for the first time to stimulate methane production.
Stimulation of multiple microbiomes in a single coal seam
We present a pragmatic longitudinal study in which four individual wells were amended with nutrients, nutrients with calcium peroxide, nutrients with acetate (positive control well) or not amended (negative control well) and responses monitored over a period of 15–18 months. Given the sampling and access constraints to gas production wells in general, this study represented a valuable opportunity to assess succession in the microbial communities over a long-term stimulation trial. Whilst there were some differences in starting conditions in each well (eg. variation in sulfate and chloride concentrations), pH and ORP, generally considered major drivers of community composition, were uniform. Regardless of initial differences, quantitative PCR of 16S rRNA and functional genes (mcrA, dsrA), 16S rRNA sequencing and geochemistry analysis of the coal formation water revealed that the microbiomes of the different wells changed in relation to treatments (Figs. 3, 4). Depending on the availability of nutrients and the presence of oxic or anoxic conditions, the microbial communities were dominated by members of the Beta- and Deltaproteobacteria, Firmicutes and methanogenic Euryarchaeota (Fig. 4). The metabolic capacities of these microbial communities were respectively centred either on oxidation of coal hydrocarbons, acetogenesis, sulfate reduction, iron reduction and methane formation. These findings may have important implications for gas enterprises to operate extraction wells and stimulate biogas formation.
Formation water chemistry dictates the indigenous microbial community composition
The formation water composition was a major factor shaping the composition and metabolic potential of the initial reservoir microbiome (Figs. 1, 3, 4 Table S1). Sequencing and qPCR of functional genes (mcrA and dsrA) indicated a strong potential for sulfate reduction and a very low potential for methanogenesis (methanogenic archaea; Figs. 3, 4a). The initial archaeal community composition was similar in all wells, with a predominance of Archaeoglobus sp. and uncultured Crenarchaeota (Fig. 4a), commonly observed in coal and oil reservoirs worldwide [51, 53, 55,56,57,58,59]. The considerable potential of Archaeoglobus species to ferment alkanes, a main constituent of coal and oil, to generate acetate has been shown recently [58]. The produced acetate could then provide a substrate source for methanogens as well as sulfate reducers. Archaeoglobus species also have the capacity to couple the degradation of hydrocarbons, like alkanes to sulfate reduction [53, 56, 60]. Metagenomic mining, genomic bin reconstruction and functional gene sequencing (dsrAB genes) indicated a strong potential for sulfate reduction and hydrogen sulphide production. Metagenomic bins contained the assA-gene, which encodes the catalytic subunit of an alkylsuccinate synthase, an enzyme for the activation of hydrocarbons by fumarate addition and genes for the utilization of activated hydrocarbons as well as genes involved in β-oxidation and the utilization of acetyl CoA [58]. Similar pathways have been identified in isolates [61] but the exact mechanism of hydrocarbon degradation is still unclear.
Crenarchaeota and closely related members of the Thaumarchaeota and Miscellaneous Crenarchaeotal Group (MCG) have been shown to carry out a heterotrophic metabolism rather than being obligate autotrophic. They can use organic carbon derived from the degradation of fossil organic matter and are likely to access substrates that are physically or chemically recalcitrant, utilizing organic compounds found in wastewater and petroleum reservoirs [62,63,64,65,66] . Crenarchaeota have been shown to dominate sulfate-reducing enrichments from petroleum reservoirs with a pronounced degradation of alkanes where the anaerobic activation of alkanes proceeded potentially via assA-genes, or alternative yet unknown mechanisms [67].
Members of Archaeoglobus and Crenarchaeota might potentially be involved in the breakdown of coal-associated hydrocarbon compounds like alkanes and fatty acids to acetate using sulfate as electron acceptor. One important characteristic of the coal formation water was the presence of moderate to high sulfate concentrations (0.5 to 3.7 mM sulfate; Fig. 2d) creating an environment where the methanogenic archaea might be outcompeted by SRB for acetate and H2 gas [26]. Elevated concentrations of sulfate have been observed in coalbed aquifers, but not in association with methane formation [24, 68]. Together this indicates that in elevated-sulfate, and low-nutrient formation waters, indigenous microbial communities were potentially linked to hydrocarbon oxidation coupled to sulfate reduction rather than to methanogenesis.
The rise of hydrocarbon degrading bacteria through the addition of nutrients
After nutrient treatments were applied, hydrocarbon degrading bacteria became abundant, affiliated with Azospira, Dechloromonas, Thauera, Azoarcus, Propionivibrio (Betaproteobacteria), Tolumonas, Pseudomonas (Gammaproteobacteria), Desulfovibrio, Geobacter, Pelobacter, Synthrophobacter (Deltaproteobacteria), Desulfotomaculum, Desulfosporosinus and Pelotomaculum species (Firmicutes; Fig. 4a). These lineages have a cosmopolitan distribution with their presence previously noted in coalbed seams worldwide, e.g. Powder River Basin, Ruhr Basin, Alberta Basin, Illinois Basin, Ishikari Basin, Waikato coalfields, deep sea coalbeds [14, 16, 21, 57, 59, 69,70,71]. Whereas the most dominant bacterial phyla were represented by the Firmicutes, Spirochaetes, Bacteroidetes and all five groups of the Proteobacteria [5]. Their predominance can be associated with the solubilisation of intermediates from coal using a range of genes (bssA, bamA, bamB, bzdN, bcrA, bcrC) involved in the degradation of a variety of hydrocarbons being capable of initially attacking and metabolizing aromatic hydrocarbons [72,73,74]. Specifically, Thauera, Azoarcus, Pseudomonas, and Geobacter species as well as SRB are recognized as key organisms for BTEX (Benzene, Toluene, Ethylbenzene, Xylene) and/or PAH (Polycyclic Aromatic Hydrocarbons) degradation under nitrate reducing, iron and sulfate reducing conditions [73, 75,76,77,78,79]. The potential utilization of different electron acceptors by hydrocarbon degrading bacteria suggests a high metabolic versatility of the microbial community which could respond more easily to changes in their availability and with the degradation of coal regardless of the electron acceptor available as observed in other coal habitats [57]. Another major group involved in anaerobic aromatic hydrocarbon degradation is the Peptococcaceae (Firmicutes) such as Desulfotomaculum, Desulfosporosinus and Pelotomaculum species (Fig. 4a; [72, 73]). These species were abundant in the calcium peroxide and nutrient amended well after the initial oxic conditions ceased and anoxic conditions were induced (Fig. 4a). Members of the phylum Firmicutes, potentially using methanol and methylamines as the sole carbon and energy source, dominate coal-bearing sediments in the deep sea [71, 80]. Congruently, Pelobacter and Syntrophobacter species increased in relative abundance (Fig. 4a) suggesting the initial oxic phase stimulated subsequent anaerobic hydrocarbon degrading processes through release of aliphatic and aromatic compounds. Additionally, acetogenic bacteria putatively involved in hydrocarbon degradation were observed in the formation water (Clostridium, Dechloromonas; Fig. 4a). Notably, sequences similar to Clostridium species were observed in earlier surveys in coal habitats [14, 16, 59, 81]. All the putative hydrocarbon-degrading bacteria may be involved in the initial breakdown of coal and its transformation into compounds further utilised by the remainder of the microbial community. During the long term incubations the accumulation of acetate was detected occasionally when methane production stalled (Fig. 2a, b) implying that the activity of acetogenic bacteria provided acetate to the methanogenic archaea.
Acetate is the central energy carrier responsible for methane production
Increases in abundance of methanogenic archaea and biogenic methane formation were stimulated in all nutrient amended wells (Figs. 2a, 3, 4a, b). Here we demonstrate that nutrient amendment results in the direct or indirect (via acetogenic bacteria) stimulation of methanogenic archaea, mainly Methanosaeta spp. and Methanosarcina spp., which are obligate and facultative acetoclastic methanogenic archaea respectively (Figs. 3, 4a). Compound specific isotope analyses of methane indicated acetoclastic methanogensis to be the dominant pathway for methane production (Table 1). Recent studies have shown that in organic-matter rich sediments, acetate was not directly converted to methane but rather oxidised to CO2 in a syntrophic relationship between acetotrophic bacteria and methanogens favouring hydrogenotrophic methanogenesis [82]. Hydrogenotrophic methanogenesis has been detected in many coal reservoirs predominantly under sulfate-limiting conditions [5, 71]. However, in the coal formation water, hydrogenotrophic methanogens appeared to be more abundant during the late methane production phase or the ultimate plateau when more biomass generated through biostimulation was likely hydrolysed and fermented (Figs. 2a, 3). H2 was not detected, most likely due to fast microbial utilization. Our results indicate that adding the methanogenic substrate acetate to prime the microbial system for enhanced methane production is not efficient since methane production accounted for only 28% of the acetate added.
A promising carbon independent amendment approach is the application of oxygen releasing agents such as calcium peroxide that lifts the ORP and potentially stimulates the initial oxidative biofragmentation of coal. This would result in the release of aliphatic and aromatic compounds that can be taken up by cells and fermented to yield electron donors such as acetate for methanogenic archaea. Microbial uptake of oxygen creates anaerobic microenvironments where the methanogenic archaea are protected, however oxidative stress strongly inhibits methanogenesis [83]. Methanosarcina and Methanolobus sp. were abundant during the first 3 months of incubation (Fig. 3), when the ORP was high (Figure S1), indicating their resilience to extended exposure to microaerophilic conditions and involvement in the initial methane production. Though methanogenic archaea are regarded as strictly anaerobic, some methanogenic archaea (eg. Methanosarcina) can tolerate prolonged periods of oxygen exposure [84, 85]. The abundance of obligate acetoclastic methanogenic genera, specifically Methanosaeta sp., increased after the removal of CaO2, potentially preferring a more reduced environment (Fig. 3).
The fate of acetate in the coal formation water
While 28% of acetate in the acetate amended positive-control well was converted to methane in situ, the fate of the remaining acetate was unclear (Fig. 2a, b). DNA stable isotope probing (DNA-SIP) was used to track the incorporation of acetate by bacteria and archaea in the coal formation water of the nutrient and acetate amended well.
Elevated sulfate concentrations and high relative abundances of specialised acetate oxidising bacteria, such as Desulfobacca acetoxidans [86], indicates a potential competition between sulfate reducing bacteria and acetoclastic methanogenic archaea, as described in various anoxic systems [26, 87, 88]. There was no increase in SRB abundance in the well amended with nutrients only (Fig. 3) indicating that acetate was the main stimulant for SRB. Intermittent acetate production with up to 4 mM acetate was observed in all nutrient amendments but otherwise the acetate concentration was below the detection limit of 50 μM (Fig. 2). Acetogenesis from H2 and CO2 would be possible below a concentration < 100 μM [89].
H2 was not detected in the coal formation water during the field trial assuming concentrations below the detection limit of 100 μM. Both, H2 and acetate, produced via bacterial fermentation of complex organic matter [90, 91], are key substrates to methanogens and SRB [87], and a rapid turnover results in acetate and H2 typically not accumulating to high concentrations, unlike the end products of sulfate reduction (H2S) and methanogenesis (CH4; [92, 93]).
Methanogenesis and sulfate reduction, using H2 as electron donor, were still thermodynamically favourable at the detection limit concentrations of 100 μM with energy yields exceeding the estimated biological energy quantum (BEQ) of ΔGr = −10 kJ mol−1 [89]. If H2 is the energy source, methanogens and SRB can support a H2 driven metabolism with energy yields as low as −10 and −20 kJ mol−1, respectively [89, 94]. Energy yields (ΔGr) for sulfate reduction and methanogenesis from H2 as electron donor, would be −130 and −90 kJ mol−1, for a detection limit of 100 μM H2 [89] with a higher free energy yield for SRB. Under H2 limiting conditions, methanogens can still gain energy down to a H2 concentration of 11 nM, whilst sulfate reducing bacteria can meet the BEQ down to a H2 concentration of 0.6 nM [89]. At an acetate concentration of 50 μM, the dG’ for acetate driven sulfate reduction would be −22.8 kJ mol−1 and for methanogenesis −6.5 kJ mol−1, suggesting sulfate reducers would outcompete methanogens for acetate.
SRB are usually obtained in higher cell densities and with higher turnover rates compared to methanogens, excluding methanogens from the consumption of acetate. SRB are capable of carrying out sulfate reduction using acetate concentrations as low as 1–20 μM. In the acetate amended well, the relative abundance of SRB was higher (106–108 cells/mL) than needed for the sulfate reduction observed in the later stage of the field trial where only 2 × 105 cells/mL would be sufficient for the reduction of 2.5–4 mM sulfate (Figs. 2, 3), indicating the relevance of potential electron acceptors other than sulfate, e.g. iron (III). Many SRB are capable of oxidizing acetate and reducing iron (III) [86, 95, 96]. SRB reduction rates were slow and the addition of nutrients did not lead to an exhaustion of the sulfate in the coal formation water during the 18 month field trial (Fig. 2), except in response to exogenous addition of acetate supporting the fact that SRB are energy (acetate) limited.
The early incorporation of 13C-label was predominantly into bacteria closely related to Geobacter uraniireducens, Geobacter pickeringii, Geobacter hydrogenophilus, Desulfovibrio vulgaris and Desulfobacca acetoxidans (Deltaproteobacteria) after day 3 in response to acetate exposure suggesting that these species are responsible for acetate degradation (Fig. 5a, b). The strong response of Geobacter species capable of acetate oxidation and Fe(III) reduction [97, 98] suggests that Fe(III) reduction might play an important role in acetate metabolism in coal formation water potentially influenced by iron well casing. Members of the genus Geobacter are often the predominant Fe(III)-reducing bacteria in subsurface anoxic environments [98]. It has been hypothesized that Desulfovibrio species may also play an important role in the reduction of Fe(III) [95, 99, 100]. Iron (III) was the only electron acceptor detected in the formation water at a concentration of 0.5 mM accounting for only 0.06 mM acetate oxidised to carbon dioxide regarding stoichiometric conversions suggesting that Fe(III) is potentially used from the sediment at the bottom of the well or from the iron well casing representing another potential source of Fe(III). Acetoclastic methanogenic archaea related to Methanosarcina acetivorans and Methanosaeta concilii incorporated 13C-acetate after 6 days when 65% of acetate was already consumed (Fig. 5b), possibly due to a slower response to acetate exposure alongside the abundant Geobacter species. The ex situ methane yield obtained supports our findings in situ that methane is not the major sink of acetate, with only 30–40% used by methanogenesis (Fig. 6c). Most notably, sulfate- and iron-reducing bacteria (Desulfovibrio and Geobacter) compete effectively for acetate.
The coal surface: a stage for interspecies interactions?
Many studies have elucidated the composition of microbial communities in coal formation water, but the presence of methanogenic archaea, sulfate reducing bacteria and hydrocarbon degrading bacteria on the coal surface in situ or over a long-term field trial has not previously been investigated. It has been hypothesised that hydrocarbon oxidising and fermentative bacteria occupy the coal surface and perform the first steps of coal degradation whilst methanogenic archaea use solublised reducing equivalents in the coal formation water to produce methane [5]. In conflict with this hypothesis we have shown that methanogenic archaea make up a crucial part of the microbial community on the coal surface (Fig. 5). These findings are supported by recent investigations by Trembath-Reichert et al. [80] and Inagaki et al. [71] showing that microbial growth is possible within a deep subseafloor coal bed, predominated by sulfate reducing and hydrocarbon degrading bacteria utilizing mainly methylated substrates and hydrogenotrophic methanogens being responsible for the vast majority of biogenic methane formation. All nutrient amendments in the long-term field trial resulted in the colonization of the coal surface by methanogenic archaea, sulfate reducing bacteria and hydrocarbon degrading bacteria (Fig. 5). No microorganisms were detected on the coal surface in the unamended control well. This is congruent with the idea that cell increase and methane production from coal is limited by the lack of available nutrients [101]. After the oxic phase ceased, the predominance of anaerobic hydrocarbon degraders and methanogenic archaea on the coal surface suggests enhanced degradation of available coal compounds to methanogenic substrates.
Conclusion
Long-term microbial community succession in coal formation water biostimulated for enhanced methanogenesis was described (Fig. 7). High sulfate concentrations and SRB did not prevent the stimulation of methanogenic archaea that are nutrient limited through the addition of a mineral nutrient amendment. Methanogenic archaea make up a major part of the coal associated biofilm communities carrying out acetoclastic methanogenesis in a gas-free coal seam. Acetate, and not H2, is the central energy carrier in this coal reservoir. SRB as well as iron reducing bacteria (IRB) are energy (acetate) limited and compete with the methanogenic archaea for acetate. Only 28% of the amended acetate accounted for methane production, the majority was consumed by SRB and IRB.
Long-term microbial community succession in coal formation water biostimulated for enhanced methanogenesis was described. High sulfate concentrations and SRB did not prevent the stimulation of methanogenic archaea that are nutrient limited through the addition of a mineral nutrient amendment. Methanogenic archaea make up a major part of the coal associated biofilm communities carrying out acetoclastic methanogenesis in a gas-free coal seam. Acetate, and not H2, is the central energy carrier in this coal reservoir. SRB as well as iron reducing bacteria (IRB) are energy (acetate) limited and compete with the methanogenic archaea for acetate. Only 28% of the amended acetate accounted for methane production, the majority was consumed by SRB and IRB
References
Mandil C. Gobal energy trends. France: IEA Publications; 2004. p. 57–81.
Dong Z, Holditch SA, McVay DA, Ayers WB. (2012). Global unconventional gas resource assessment. SPE Econ Manag. 4:222–34.
Pan Z, Wood DA. Virtual special issue: coalbed methane (CBM) exploration, reservoir characterisation, production, and modelling: a collection of published research (2009-2015). J Nat Gas Sci Eng. 2015;26:1491–4.
Rice DD, Claypool GE. Generation, accumulation, and resource potential of biogenic gas. AAPG Bull. 1981;65:5–25.
Strapoc D, Mastalerz M, Dawson K. Biogeochemistry of microbial coal-bed methane. Annu Rev Earth Planet Sci. 2011;38:617–56.
Dresselhaus MS, Thomas IL. Alternative energy technologies. Nature. 2001;414:332–7.
McJeon H, Edmonds J, Bauer N, Clarke L, Fisher B, Flannery BP, et al. Limited impact on decadal-scale climate change from increased use of natural gas. Nature. 2014;514:482–5.
Beckmann S, Welte C, Li X, Kroeninger L, Heo Y, Zhang M, et al. Novel phenazine crystals enable direct electron transfer to methanogens in anaerobic digestion by redox potential modulation. Energy Environ Sci. 2016;9:644–55.
Thielemann T, Cramer B, Schippers A. Coalbed methane in the Ruhr Basins, Germany: a renewable energy resource? Org Geochem. 2004;35:1537–49.
Schoell M. The hydrogen and carbon isotopic composition of methane from natural gases of various origins. Geochim Cosmochi Acta. 1980;44:649–61.
Whiticar MJ, Faber E, Schoell M. Biogenic methane formation in marine and freshwater environments: CO2 reduction vs. acetate fermentation – isotope evidence. Geochim Cosmochim Acta. 1986;50:693–709.
Fakoussa RM, Hofrichter M. Biotechnology and microbiology of coal degradation. Appl Microbiol Biotechnol. 1999;52:25–40.
Jones EJP, Voytek MA, Corum MD, Orem WH. Stimulation of methane generation from nonproductive coal by addition of nutrients or a microbial consortium. Appl Environ Microbiol. 2010;76:7013–22.
Beckmann S, Krueger M, Engelen B, Gorbushina AA, Cypionka H. Role of bacteria, archaea and fungi involved in methane release in abandoned coal mines. Geomicrbiol J. 2011a;28:347–58.
Krueger M, Beckmann S, Engelen B, Thielemann T, Cramer B, Schippers A, et al. Microbial methane formation from hard coal and mine timber in an abandoned coal mine. Geomicrobiol J. 2008;25:315–21.
Beckmann S, Lueders T, Krueger M, Von Netzer F, Engelen B, Cypionka H. Acetogens and acetoclastic Methanosarcinales govern methane formation in abandoned coal mines. Appl Environ Microbiol. 2011b;77:3749–56.
Orem WH, Yoytek MA, Jones EJ, Lerch HE, Bates AL, Corum MD, et al. Organic intermediates in the anaerobic biodegradation of coal to methane under laboratory conditions. Org Geochem. 2010;41:997–1000.
Orem WH, Tatu C, Varonka M, Lerch H, Bates A, Engle M, et al. Organic substances in produced and formation water from unconventional natural gas extraction in coal and shale. Int J Coal Geol. 2014;126:20–31.
Schink B. Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev. 1997;61:262–80.
Bates BL, McIntosh JC, Lohse KA, Brooks PD. Influence of groundwater flowpaths, residence times and nutrients on the extent of microbial methanogenesis in coal beds: Powder River Basin, USA. Chem Geol. 2011;284:45–61.
Penner TJ, Foght JM, Budwill K. Microbial diversity of western Canadian subsurface coal beds and methanogenic coal enrichment cultures. Int J Coal Geol. 2010;82:81–93.
Ritter D, Vinson D, Barnhart E, Akob DM, Fields MW, Cunningham AIB, et al. Enhanced microbial coalbed methane generation: a review of research, commercial activity, and remaining challenges. Int J Coal Geol. 2015;146:28–41.
Cassidy DP, Irvine RL. Use of calcium peroxide to provide oxygen for contaminant biodegradation in a saturated oil. J Hazard Matter. 1999;69:25–39.
Van Voast WA. Geochemical signature of formation waters associated with coalbed methane. AAPG Bull. 2003;87:667–76.
Fallgreen PH, Zeng C, Ren Z, Lu A, Ren S, Jin S. Feasibility of microbial production of new natural gas from non-gas-producing lignite. Int J Coal Geol. 2013;115:79–84.
Lovley DR, Klug MJ. Sulfate reducers can outcompete methanogens at freshwater sulfate concentrations. Appl Environ Microbiol. 1983;45:187–92.
Hazrin-Chong H, Manefield M. An alternative SEM drying method using hexamethyldisilazane (HMDS) for microbial cell attachment studies on sub-bituminous coal. J Microbiol Methods. 2012;90:96–99.
Whiticar MJ. Carbon and hydrogen isotope systematics of bacterial formation of methane. Chem Geol. 1999;161:291–314.
Kampbell DH, Vandegrift SA. Analysis of dissolved methane, ethane, and ethylene in ground water by a standard gas chromatography technique. J Chromatogr Sci. 1998;36:253–6.
Lunau M, Lemke A, Walther K, Martens-Habbena W, Simon M. An improved method for counting bacteria from sediments and turbid environments by epifluorescence microscopy. Environ Microbiol. 2005;7:961–8.
Lueders T, Manefield M, Friedrich MW. Enhanced sensitivity of DNA- and rRNA –based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ Microbiol. 2004a;6:73–78.
Luk A, Beckmann S, Manefield M. Dependency of DNA extraction efficiency on cell concentration invalidates molecular quantification of microorganisms in groundwater. FEMS Microbiol. Ecol. 2018;94. FEMSEC-18-03-0167.R1.
McCoy WH, Olsen BH. Fluorometric determination of the DNA concentration in municipal drinking water. Appl Environ Microbiol. 1985;49:811–7.
Paul JH, Carlson DJ. Genetic material in the marine environment: Implication for bacterial DNA. Limnol Oceanogr. 1984;29:1091–6.
Paul JH, Jeffrey WH, DeFlaun M. Particulate DNA in subtropical oceanic and estuarine planktonic environments. Mar Biol. 1985;90:95–101.
Klappenbach JA, Saxman PR, Cole JR, Schmidt TM. Rmdb: The ribosomal RNA operon copy number database. Nucleic Acids Res. 2001;29:181–4.
Stoddard SF, Smith BJ, Hein R, Roller BRK, Schmidt TM. rrnDB: Improved tools for interpreting rRNA gene abundance in bacteria and archaea and a new foundation for future development. Nucleic Acids Res. 2015;43:D598.
Matsuki T, Watanabe K, Fujimoto J, Miyamoto Y, Takada T, Matsumoto K, et al. Development of 16S rRNA-gene-targeted group-specific primers for the human feces. Appl Environ Microbiol. 2002;68:5445–51.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing MOTHUR: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Schloss PD, Gevers D, Westcott SL. Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies. PloS One. 2011;6:e27310.
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics. 2011;27:2194–200.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucl Acids Res. 2013;41:D590–6.
Lueders T, Wagner B, Claus P, Friedrich MW. Stable isotope probing of rRNA and DNA reveals a dynamic methylotroph community and trophic interactiona with fungi and protozoa in oxic rice field soil. Environ Microbiol. 2004b;6:60–72.
Wilkins D, Van Sebille E, Rintoul SR, Lauro FM, Cavicchioli R. Advection shapes southern ocean microbial assemblages independent of distance and environmental effects. Nat Commun. 2013;4:2457.
Aronesty E. (2011). ea-utils: “Command-line tools for processing biological sequencing data”.
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD. Development of a dual-index sequencing strategy and curation pipeline for analysing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol. 2013;79:5112–20.
Clarke KR. Non-parametric multivariate analyses of changes in community structure. Aust J Ecol. 1993;18:117–43.
Clarke KR, Warwick RM. Change in marine communities: an approach to statistical analysis and interpretation. Plymouth: PRIMER-E; 2001. p. 172.
Clarke KR, Gorley RN. PRIMERv7: User manual/tutorial. Plymouth: PRIMER-E; 2015. p. 296.
Ward CR. Coal geology and coal technology. Palo Alto, CA: Blackwell Scientific Publications, Inc; 1984.
Beeder J, Nilsen RK, Rosnes JT, Torsvik T, Lien T. Archaeoglobus fulgidus isolated from hot North Sea oil field waters. Appl Environ Microbiol. 1994;60:1227–31.
Stetter KO, Huber R, Bloechl E, Kurr M, Eden RD, Fielder M, et al. Hyperthermophilic archaea are thriving in deep North Sea and Alaskan oil reservoirs. Nature. 1993;365:743–5.
Vigneron A, Alsop EB, Lomans BP, Kyrpides NC, Head IM, Tsesmetzis N. Succession in the petroleum reservoir microbiome through an oil field production lifecycle. ISME J. 2017;11:2141–54.
He H, Han Y, Jin D, Leng W, Sun Q, Shen L, et al. Microbial consortium in a non-production biogas coal mine of eastern China and its methane generation from lignite. Energ Source. 2016;38:1377–84.
Li D, Hendry P, Faiz M. A survey of the microbial populations in some Australian coalbed methane reservoirs. Int J Coal Geol. 2008;76:14–24.
Birkeland NK, Schoenheit P, Poghosyan L, Fiebig A, Klenk HP. (2017) Complete genome sequence analysis of Archaeoglobus fulgidus strain 7324 (DSM 8774), a hyperthermophilic archaeal sulfate reducer from a North Sea oil field. Stand Genomic Sci. 12: https://doi.org/10.1186/s40793-017-0296-5.
Gruendger F, Jimenez N, Thielemann T, Straaten N, Lueders T, Richnow HH et al. (2015). Microbial methane formation in deep aquifers of a coal-bearing sedimentary basin, Germany. Front Microbiol. 6: https://doi.org/10.3389/fmicb.2015.00200.
Liu YF. (2018). Metabolic capability and in situ activity of microorganisms in an oil reservoir. Microbiome. 6: https://doi.org/10.1186/s40168-017-0392-1.
Shimizu S, Akiyama M, Naganuma T, Fujioka M, Nako M, Ishijima Y. Molecular characterization of microbial communities in deep coal seam groundwater of northern Japan. Geobiol. 2007;5:423–33.
Khelifi N, Amin Ali O, Roche P, Grossi V, Brochier-Armanet C, Valette O, et al. Anaerobic oxidation of long-chain n-alkanes by the hyperthermophilic sulfate-reducing archaeon. Archaeoglobus fulgidus ISME J. 2014;8:2153–66.
Klenk HP, White O, Ketchum KA. The complete genome sequence of the hyperthermophilic, sulphate-reducing archaeon Archaeoglobus fulgidus. Nature. 1997;390:364–70.
Kubo K, Lloyd KG, Biddle JF, Amann R, Teske A, Knittel K. Archaea of the Miscellaneous Crenarchaeotal Group are abundant, diverse and widespread in marine sediments. ISME J. 2012;6:1949–65.
Mussmann M, Brito I, Pitcher A, Damste JSS, Hatzenpichler R, Richter A, et al. Thaumarchaeotes abundant in refinery nitrifying sludges express amoA but are not obligate autotrophic ammonia oxidizers. PNAS. 2011;108:16771–76.
Ren HY. (2011). Comparison of microbial community compositions of injection and production well samples in a long-term water-flooded petroleum reservoir. PLOS ONE 6 : https://doi.org/10.1371/journal.pone.0023258.
Tang YQ, Li Y, Zhao JY, Chi CQ, Huang LX, Dong HP et al. (2012). Microbial communities in long-term, water-flooded petroleum reservoirs with different in situ temperatures in the huabei oilfield, china. PLOS ONE 7: https://doi.org/10.1371/journal.pone.0033535.
Biddle JF, Lipp JS, Lever MA, Lloyd KG, Sorensen KB, Anderson R, et al. Heterotrophic archaea dominate sedimentary subsurface ecosystems off Peru. PNAS. 2006;103:3846–51.
Li W, Wang LY, Duan RY, Liu JF, Gu JD, Mu BZ. Microbial community characteristics of petroleum reservoir production water amended with n-alkanes and incubated under nitrate-, sulfate-reducing and methanogenic conditions. Int Biodeterior Biodegrad. 2012;69:87–96.
Taulis M, Milke M. Chemical variability of groundwater samples collected from a coal seam gas exploration well, Maramarua, New Zealnad. Water Res. 2013;47:1021–34.
Barnhart EP, Weeks EP, Jones EJP, Ritter DJ, McIntosh JC, Clark AC, et al. Hydrogeochemistry and coal-associated bacterial populations from a methanogenic coal bed. Int J Coal Geol. 2016;162:14–26.
Fry JC, Horsfield B, Sykes R, Cragg BA, Heywood C, Kim GT, et al. Prokaryotic populations and activities in an interbedded coal deposit, including a previously deeply buried section (1.6-2.3 km) above ~150 Ma basement rock. Geomicrobiol J. 2009;26:163–78.
Inagaki F, Hinrichs KU, Kubo Y, Bowles MW, Heuer VB, Hong WL, et al. Exploring deep microbial life in coal-bearing sediment down to ∼2.5 km below the ocean floor. Science. 2015;349:420–4.
Von Netzer F, Kuntze K, Vogt C, Richnow HH, Boll M, Lueders T. Functional gene markers for fumarate-adding and dearomatizing key enzymes in anaerobic aromatic hydrocarbon degradation in terrestrial environments. J Mol Microbiol Biotechnol. 2016;26:180–94.
Weelink SAB, Eekert MHA, Stams AJM. Degradation of BTEX by anaerobic bacteria: physiology and application. Rev Environ Sci Biotechnol. 2010;9:359–85.
Widdel F, Knittel K, Galushko A. Anaerobic hydrocarbon-degrading microorganisms: an overview. In: Timmis KN, (ed.). Handbook of hydrocarbon and lipid microbiology. Berlin/Heidelberg: Springer; 2010. p. 1997–2021.
Chakraborty R, Coates JD. Anaerobic degradation of monoaromatic hydrocarbons. Appl Microbiol Biotechnol. 2004;64:437–46.
Chakraborty R, O’Connor SM, Chan E, Coates JD. Anaerobic degradation of benzene, toluene, ethylbenzene, and xylene compounds by Dechloromonas strain RCB. Appl Environ Microbiol. 2005;71:8649–55.
Meckenstock RU, Mouttaki H. Anaerobic degradation of non-substituted aromatic hydrocarbons. Curr Opin Biotechnol. 2011;22:406–14.
Parales RE. Hydrocarbon degradation by betaproteobacteria. In: Timmis K, ed.. Handbook of Hydrocarbon and Lipid Microbiology. Berlin: Springer; 2010. p. 1715–24. Doi: 10.1007/978-3-540-77587-4_121
Rockne KJ, Chee-Sanford JC, Sanford RA, Hedlund BP, Staley JT, Strand SE. Anaerobic naphthalene degradation by microbial pure cultures under nitrate-reducing conditions. Appl Environ Microbiol. 2000;66:1595–601.
Trembath-Reichert E, Morono Y, Ijiri A, Hoshino T, Dawson KS, Inagaki F et al. (2017) Methyl-compound use and slow growth characterize microbial life in 2-km-deep subseafloor coal and shale beds. PNAS: https://doi.org/10.1073/pnas1707525114.
Shin SG, Lee CS, Hwang K, Ahn JH, Hwang S. Use of order-specific primers to investigate the methanogenic diversity in acetate enrichment system. J Indust Microbiol Biotechnol. 2008;35:1345–52.
Beulig F, Roy H, Glombitza C, Jorgensen BB. Control on rate and pathway of anaerobic organic carbon degradation in the seabed. PNAS. 2018;115:367–72.
Yuan Y, Conrad R, Lu Y. Responses of methanogenic archaeal community to oxygen exposure in rice field soil. Environ Microbiol Rep. 2009;1:347–54.
Kato MT, Field JA, Lettinga G. High tolerance of methanogens in granular sludge to oxygen. Biotechnol Bioeng. 1993;42:1360–6.
Kiener A, Leisinger T. Oxygen sensitivity of methanogenic bacteria. Syst Appl Microbiol. 1983;4:305–12.
Oude Elferink SJWH, Akkermans-van Vliet WM, Bogte JJ, Stams AJM. Desulfobacca acetoxidans gen. nov., sp. nov., a novel acetate-degrading sulfate reducer isolated from sulfidogenic granular sludge. Int J Syst Evol Microbiol. 1999;49:345–50.
Conrad R, Lupton FS, Zeikus JG. Hydrogen metabolism and sulfate-dependent inhibition of methanogenesis in a eutrophic lake sediment (Lake Mendota). FEMS Microbiol Ecol. 1987;45:107–15.
Muyzer G, Stams AJM. The ecology and biotechnology of sulfate-reducing bacteria. Nat Rev. 2008;6:441–54.
Lever MA. (2012). Acetogenesis in the energy-starved deep biosphere – a paradox? Front Microbiol. 2: https://doi.org/10.3389/fmicb.2011.00284.
Dolfing J. Acetogenesis. In: Zehnder AJB, Ed. Biology of anaerobic microorganisms. New York, NY: Wiley-Interscience; 1988. p. 417–68.
McInerney MJ. Anaerobic hydrolysis and fermentation of fats and proteins. In: Zehnder AJB, Ed. Biology of anaerobic microorganisms. New York, NY: Wiley-Interscience; 1988. p. 373–415.
Zinder SH. Physiological ecology of methnaogens. In: Ferry JG, Ed. Methanogenesis. New York, NY: Chapman and Hall; 1993. p. 128–206.
Hoehler TM, Alperin MJ, Albert DB, Martens CS. Thermodynamic control on H2 concentrations in an anoxic marine sediment. Geochim Cosmochim Acta. 1998;62:1745–56.
Hoehler TM, Alperin MJ, Albert DB, Martens CS. Apparent minimum free energy requirements for methanogenic archaea and sulfate-reducing bacteria in an anoxic marine sediment. FEMS Microbiol Ecol. 2001;38:33–41.
Coleman ML, Hedrick DB, Lovley DR, White DC, Pye K. Reduction of Fe(III) in sediments by sulfate-reducing bacteria. Nature. 1993;361:436–8.
Widdel F, Bak F. Gram-negative mesophilic sulfate-reducing bacteria. The prokaryotes: a handbook on the biology of bacteria: ecophysiology, isolation, identification, application.. New York, NY: Springer; 1992. p. 3353–78.
Coates JD, Bhupathiraju VK, Achenbach LA, McInerney MJ, Lovley DR. Geobacter hydrogenophilus, Geobacter chapellei and Geobacter grbiciae, three new, strictly anaerobic, dissimilatory Fe(III)-reducers. Int J Syst Evol Microbiol. 2001;51:581–8.
Shelobolina ES, Nevin KP, Blakeney-Hayward JD, Johnsen CV, Plaia TW, Krader P, et al. Geobacter pickeringii sp. nov., Geobacter argillaceus sp. nov. and Pelosinus fermentans gen. nov., sp. nov., isolated from subsurface kaolin lenses. Int J Sys Evol Microbiol. 2007;57:126–35.
Lovley DR. Dissimilatory metal reduction. Annu Rev Microbiol. 1993;47:263–90.
Sun B, Cole JR, Sanford RA, Tiedje JM. Isolation and characterization of Desulfovibrio dechloracetivorans sp. nov., a marine dechlorinating bacterium growing by coupling the oxidation of acetate to the reductive dechlorination of 2-chlorophenol. Appl Environ Microbiol. 2000;66:2408–13.
Lyles CN, Parisi VA, Beasley WH, Van Nostrand JD, Zhou J, Suflita JM. (2017) Elucidation of the methanogenic potential from coalbed microbial communities amended with volatile fatty acids. FEMS Microbiol Ecol 93 https://doi.org/10.1093/femsec/fix040.
Acknowledgements
This work was supported by the Australian Research Council Linkage Project LP100100128, Discovery Project DP140100621 and industry partner Biogas Energy Pty Ltd.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Beckmann, S., Luk, A.W.S., Gutierrez-Zamora, ML. et al. Long-term succession in a coal seam microbiome during in situ biostimulation of coalbed-methane generation. ISME J 13, 632–650 (2019). https://doi.org/10.1038/s41396-018-0296-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0296-5
This article is cited by
-
A review on microbial metabolism to increase coalbed methane generation and coal pretreatment to improve its bioavailability
Frontiers of Earth Science (2023)
-
Coexistence of Psychrophilic, Mesophilic, and Thermophilic Sulfate-Reducing Bacteria in a Deep Subsurface Aquifer Associated with Coal-Bed Methane Production
Microbial Ecology (2023)
-
Characteristics of microbial communities in water from CBM wells and biogas production potential in eastern Yunnan and western Guizhou, China
Frontiers of Earth Science (2023)
-
Subsurface hydrocarbon degradation strategies in low- and high-sulfate coal seam communities identified with activity-based metagenomics
npj Biofilms and Microbiomes (2022)
-
Activity-based, genome-resolved metagenomics uncovers key populations and pathways involved in subsurface conversions of coal to methane
The ISME Journal (2022)