Abstract
Microbial communities are key engines that drive earth’s biogeochemical cycles. However, existing ecosystem models have only limited ability to predict microbial dynamics and require the calibration of multiple population-specific empirical equations. In contrast, we build on a new kinetic “Microbial Transition State” (MTS) theory of growth derived from first principles. We show how the theory coupled to simple mass and energy balance calculations provides a framework with intrinsically important qualitative properties to model microbial community dynamics. We first show how the theory can simultaneously account for the influence of all the resources needed for growth (electron donor, acceptor, and nutrients) while still producing consistent dynamics that fulfill the Liebig rule of a single limiting substrate. We also show consistent patterns of energy-dependent microbial successions in mixed culture without the need for calibration of population-specific parameters. We then show how this approach can be used to model a simplified activated sludge community. To this end, we compare MTS-derived dynamics with those of a widely used activated sludge model and show that similar growth yields and overall dynamics can be obtained using two parameters instead of twelve. This new kinetic theory of growth grounded by a set of generic physical principles parsimoniously gives rise to consistent microbial population and community dynamics, thereby paving the way for the development of a new class of more predictive microbial ecosystem models.
Similar content being viewed by others
Introduction
Microbes are the most abundant living things on earth [1] and are the key engines that drive earth’s biogeochemical cycles [2]. Developing models able to capture and predict their dynamics and community assembly patterns is therefore of the outmost importance for the study of global earth ecological equilibria and the development of innovative microbial biotechnology processes [3,4,5]. However, current microbial growth models are based on empirical equations, such as those from Monod [6], Contois [7] or Haldane-Andrews [8] and require extensive parameter calibration based on experimental data. Calibrated models are bound to specific experimental conditions, thus limiting their predictive abilities to a narrow domain. In order to build more generic models, there is a need to more thoroughly capture the fundamental drivers of microbial growth and to mathematically express how they contribute to the emergence of the many community assembly patterns observed in nature.
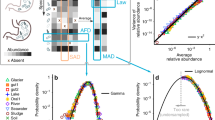
An increasing number of observations show that environmental physical–chemical factors shape the metabolic niches within a given biotope, resulting in stable functional microbial community structures despite random invasions [9]. This “functional convergence” phenomenon, i.e., the tendency of microbial functional groups to converge toward defined patterns in specific biotopes, has been observed in systems as diverse as sea water [10, 11], soil [12, 13], activated sludge [14], plant foliage [9], cheese [15], and numerous human body biomes [16]. These observations suggest that mechanistic processes may largely determine the functional patterns of microbial communities. Up to now, the search for mechanistic physics-based models of microbial community structure has been somewhat overlooked, as the very existence of general principles governing the structuring of ecosystems is still the subject of debate [17,18,19,20]. As complex as community dynamics can be, it is nevertheless recognized that an ecosystem complies with general laws such as those of thermodynamics, and the rule of stoichiometry [18], and that these laws play a role in ecosystem structure [21]. Whether achievable or not, trying to build on these laws to create a general framework that could result in ecologically consistent population and community patterns is a highly desirable research target [5].
The seeds for such a general framework were sown by the pioneering work of McCarty [22], followed by several other highly detailed studies [23,25,26,27] of the thermodynamic balance of microbial growth. This framework relies on thermodynamic and stoichiometric rules to describe the yield and output of microbial metabolism in terms of chemical species, heat, and entropy. These contributions led to the development of generic methods to predict the stoichiometry and the energetic balance of microbial growth (for a review see refs. [28] and [29]).
Several attempts have also been made to link thermodynamic balance calculations to the computation of microbial growth rates (reviewed in refs. [3] and [28]). For that purpose, some approaches combined balance calculations with heuristic Monod-like relationships between the concentration of the substrate and the absorption rate [24] and were used to simulate virtual microbial ecosystem dynamics [30]. Some authors made assumptions about the structure of the metabolic network, the electron transport chain or the organization of the pathway to establish a link between energy balances and rates [31,32,33,34]. It was also suggested to rely on the fact that the rate of microbial reactions is governed by enzyme kinetics in order to use the Michaelis–Menten theory and derive microbial rate equations encompassing thermodynamic constraints [35]. A collision frequency theory for microbial growth was also proposed [36] and coupled to thermodynamic considerations [37], which was the first attempt to conceptualize the growth phenomena on a physical basis. Recently, a more fundamental kinetic theory of microbial growth grounded by statistical physics principles was introduced [38]. For the sake of brevity, this theory is hereafter called “Microbial Transition State” (MTS) theory.
Here we investigate using the MTS approach for the modeling of microbial ecosystems. The theory underlying this approach analyzes an elementary microbial division event using first principles. The probability of a single cell to be surrounded by a sufficient amount of resources (electron donors, acceptors and nutrients) to allow division is expressed, considering their microscopic distribution in the culture medium. Thence, a growth rate formula is derived at the population level, as the statistical outcome of the probability of division computed at the level of each individual. This ab initio analysis is sufficiently fundamental to be independent of the microbial species and growth conditions. This implies that the growth of any chemotrophic microbial population catalyzing a defined metabolic reaction can be readily implemented with the MTS model. In cases where the model parameters are not calibrated, what is predicted is the dynamics of functional microbial populations as directly derived from the first principles grounding the MTS theory. The resulting equations of growth are seen here as the first fundamental layer determining microbial dynamics, as a result of physical laws, on top of which parametric equations can be added in the future to account for the complexity of biological or ecological phenomena that are beyond the scope of the current analysis. The approaches developed in this work represent an opposite take on modeling compared to many models in biology that are bound to a specific system, and that calibrate the parameters of an empirical expression in order to quantitatively reproduce the behaviors of the experimental system under study. In contrast, the objective of this contribution is to evaluate the qualitative properties of MTS dynamical models as derived solely from the set of fundamental hypotheses grounding the theory. To this end, we focus on generic and idealized situations starting from the simplest one, i.e., a pure culture grown on a minimal medium, and progressively add more complexity to the simulations, in order to analyze and to question the consistency of the population and community dynamics that arise directly (without parameter adjustment) from microbial transition state theory.
Methods
Modeling ecosystem reaction stoichiometry
The ecosystem includes a biotic and an abiotic component. The abiotic component comprises all chemical reagents, products, and spectator species. The biotic component consists of the whole microbial community. In our approach, the microbial community is subdivided into guilds. A guild is defined by the metabolism it catalyzes (aerobic acetotrophs, denitrifying acetotrophs etc., see list below). The population density of each guild is represented by the molar concentration of a generic, C-normalized biomass molecule. The biomass molecule used in the simulations is C1H1.613O0.557N0.158 [39]. Its enthalpy of formation is −126.83 kJ. mol−1 and its Gibbs energy of formation is −82.16 kJ. mol−1. The molecular weight of this molecule is 24.76 g.C-mol-Biomass−1. Assuming a cell volume of 1e−18 m3 per cell and a cell density of 1.09e6 g. m−3 [40], a factor of 22.7e12 cell. C-mol-Biomass−1 will be used for the purpose of illustration in the results. The choice of this molecule does not constitute the MTS model and theoretically, any other biomass formula could be used.
The system stoichiometry is formulated using a vectorial approach as proposed by Roels [26], thereby enabling a compact formulation. The generic formulation is described in this section. The reader should refer to the results section for specific examples of implementation. In all the following formulas, the * symbol denotes matrix product. Let r be the number of reagents involved in the system and C be a r × 1 vector storing the concentration of all reagents of the system, including biomasses, in mM at a given time. Let p be the number of processes affecting the concentrations of the reagents; the derivative of C over time is expressed from the balance equation of C as
where A is a r × p matrix storing the (unitless) stoichiometric coefficients of every process for every reagent, and R is the p × 1 vector of the rate (in time−1) of every process. By convention, the stoichiometric coefficients are either positive or negative depending on the production or consumption of the corresponding chemical species, respectively. The processes described by the MTS model are metabolic reactions. Let g be the number of guilds, Amet the matrix of dimension r × g storing the stoichiometric coefficients of the metabolism of every guild, and Rmet the g × 1 vector of the rate of each guild reaction. Eventual stoichiometric matrices describing other processes are horizontally concatenated to Amet, and their rates are vertically concatenated to Rmet. The Amet matrix is a linear combination of two matrices Aan and Acat, both of dimensions r × g, respectively, storing the coefficients of the anabolic and catabolic reactions, and adjusted to close the elemental balance in each reaction separately. The stoichiometric coefficients of a catabolic reaction are set so that exactly one electron donor molecule is consumed (unitless stoichiometric coefficients are actually mol/molDonor ratios). The stoichiometric coefficients of an anabolic reaction are set so that exactly one biomass molecule is produced (unitless stoichiometric coefficients are actually mol/C-mol-Biomass ratios) as the production of one unit of biomass is considered as the elementary event in the MTS model.
For the anabolic reaction, we use the convention proposed by Heijnen: the C-source for anabolism is either the electron donor when it is organic or HCO3−. We posit here that the reduced and oxidized forms of the catabolism’s electron donor should be included in the anabolism. Our working hypothesis is that the nitrogen source used for anabolism is ammonium (no organic sources of nitrogen are present in the simulated culture media), because it is available in every culture medium simulated in this article. Another choice would have been nitrate, but its possible role as an electron acceptor would have made the interactions between guilds more complex and complicated the message of the simulations.
The Gibbs free energy of formation of every chemical species used in the simulations is taken from Kleerebezem and collaborators [28].
Linking growth stoichiometry to energy balance
The overall metabolism of the whole microbial community can thus be expressed as:
where λ is the number of times the catabolic reaction of a guild has to be performed for the total produced energy to equal the energy barrier of growth (therefore expressed as mol.DonormolBiomass−1). λ is then a diagonal matrix of guild-specific scalar factors (denoted λg) that ensures the coupling of energy and stoichiometric balances, as explained by Kleerebezem [28], a factor also sometimes denoted fcat:
where ΔGan is the Gibbs free energy change for the anabolic reaction, ΔGcat is the Gibbs free energy change for the catabolic reaction and ΔGdis is the dissipated free energy of growth. The dissipated free energy is the Gibbs free energy change in the overall growth reaction (anabolism and catabolism). This energy dissipation makes the overall Gibbs free energy change of growth negative, so the reaction is spontaneous. ΔGan and ΔGcat are computed from the Gibbs free energy change for anabolic and catabolic reactions, corrected for non-standard temperature and concentrations. We posit that only exergonic catabolic reactions can lead to growth. Therefore, if ΔGcat happens to be positive during the computations, it is set to zero, resulting in insignificant growth. The λg factors are computed at each time step of system integration.
Water and biomass activities are not included in the mass action ratio ΔGan, ΔGcat and ΔGdis values are expressed in kJ.C-molBiomass−1. The value of each λg is such that the Gibbs energy variation of the metabolic reaction is equal to the dissipated energy, which in this article, is assumed to be identifiable with the variable \(- Y_{GX}^{\mathrm {max}}\), empirically defined by Heijnen as
where the chain length of the carbon source is denoted \(\left( {NoC_{C_S}} \right)\) and (\(\gamma _{C_S}\)) is the degree of reduction of the carbon source required for heterotrophic growth. The degree of reduction of a carbon source is computed as in the original publication by Heijnen and collaborators [23]. That is, by summing 4 electron.carbon−1, 1 electron.hydrogen−1, −2 electron.oxygen−1, −3 electron.nitrogen−1 and ±1 electron.charge−1. For example, the reduction degree \(\gamma _{C_S}\) of the carbon source acetate (C2H3O2−) is 4×2+3−2×2+1=8 electron.acetate−1.
Coupling stoichiometry, energy balance, and microbial dynamics
The growth rate function used in the simulations is the multi-substrate growth rate function described in the supplementary materials of Desmond-Le Quéméner and Bouchez [38]. This formula arises from simple hypotheses concerning microbial growth at microscopic scale. These principles can be summarized as:
-
–a microbial cell needs to overcome a fixed energy barrier in order to divide
-
–this energy barrier can be broken down into anabolic energy ΔGan and dissipated energy ΔGdis
-
–the energy available to overcome the energy barrier is the catabolic energy ΔGcat obtained from the catabolism of substrate molecules
-
–substrate molecules are considered as particles randomly distributed around the cells
-
–if a fictional, fixed volume Vh (harvest volume) around the cell contains enough substrate to overcome the energy barrier, the cell is said to be in an “activated” state
-
–only an activated cell is able to divide
Considering these hypotheses, the proportion of activated cells in the culture medium at a given time can be expressed using a probabilistic reasoning (detailed in ref. [38]). Hence, for a given guild, the formula of the microbial growth rate is
where μ is the growth rate (time−1), Amet,i the negative stoichiometric coefficient of substrate i (mol.C-mol-Biomass−1) computed in Eq. 2, and [Si] the concentration of substrate i (mol.volume−1). Although visually different, this formula is consistent with the one presented in the introductory article of the MTS model [38] (see supplementary materials 1). This formula encompasses two parameters: μmax, which represents the maximum growth rate (time−1) and Vh, which represents the harvest volume (m3.C-mol-Biomass−1). These parameters both aggregate generic physical phenomena and particular biological characteristics that would be very difficult to assess accurately for each specific guild considered. As the purpose of this article is to document generic growth patterns as derived from MTS theory, we intentionally made simple generic choices for the value of these parameters. The value of μmax for every guild was set to \(\frac{{k_{\mathrm {B}}T}}{h}\) where kB is the Boltzmann constant (m2.kg.s−2.K−1), T the temperature of the system (K) and h Planck’s constant (m2.kg.s−1); the result is close to 2.23e16 h−1 at 298.15 K. This term comes from Eyring’s transition state theory [41], on which the MTS theory is based. The value of the Vh parameter was set to 1 m3.C-mol-Biomass−1 for all the guilds, except in the final simulation where its value was set to 10 m3.C-mol-Biomass−1 as a working hypothesis because, given the low yields of the autotrophs, a value of 1 m3.C-mol-Biomass−1 proved to be insufficient (see supplementary materials 2). Considering the previously estimated ratio of 22.7e12 cell.C-mol-Biomass−1, the individual cell harvest volume for a Vh of 1 m3.C-mol−1 would be 4.41e−14 m3.cell−1. A sphere of this volume would have a radius of approximately 14 µm (see supplementary materials 3). The Vh parameter modulates the probability for a cell to be surrounded by sufficient substrate to be activated. From a biological standpoint, Vh results from all adaptations implemented by the cells to increase their ability to collect chemical resources in the culture medium (such as specific membrane transporters or chemotaxis), but also may vary according to the physical characteristics of the biotope such as the diffusivity of the substrates or agitation.
While pH can have many different impacts on the regulation of microbial growth, here only its influence on reaction equilibrium is taken into account through the mass action ratio. Other types of pH effects are beyond the scope of the current MTS model.
The g × 1 Rmet vector storing the rate of each metabolic reaction is
where M is the g × 1 vector of the microbial growth rate of each guild and [X] the g × 1 vector of the biomass concentration of each guild.
This ordinary differential equation system is implemented and solved using Matlab (MATLAB Release 2014a, The MathWorks, Inc., Natick, Massachusetts, United States.).
Microbial guilds considered
Glucosotroph
-
○ Catabolism: C6H12O6 + 6 O2 → 6 HCO3− + 6 H+ (ΔG0′ = −2841.3 kJ.molDonor−1)
-
○ Anabolism: 0.167 C6H12O6 + 0.158 NH4+ → 0.430 H2O + 0.164 H+ + 0.00625 HCO3− + C1H1.613O0.557N0.158 (ΔG0′ = −28.3 kJ.C-mol-Biomass−1)
-
○ Dissipated energy: 236.05 kJ.C-mol-Biomass−1
-
Aerobic acetotroph (Ordinary Heterotrophic Organisms):
-
○ Catabolism: C2H3O2− + 2 O2 → + 2 HCO3− + 1 H+ (ΔG0′ = −844.4 kJ.molDonor−1)
-
○ Anabolism: 0.503 C2H3O2− + 0.158 NH4+ + 0.338 H+ → 0.4305 H2O + 0.0063 HCO3− + C1H1.613O0.557N0.158 (ΔG0′ = 23.9 kJ.C-mol-Biomass−1)
-
○ Dissipated energy: 432.12 kJ.C-mol-Biomass−1
-
-
Sulfate reducing acetotroph:
-
○ Catabolism: 1 C2H3O2− + 1 SO4−2 → 2 HCO3− + 1 HS−(ΔG0′ = -47.7 kJ.molDonor−1)
-
○ Anabolism: 0.503 C2H3O2− + 0.158 NH4+ + 0.338 H+ → 0.4305 H2O + 0.0063 HCO3− + C1H1.613O0.557N0.158 (ΔG0′ = 23.9 kJ.C-mol-Biomass−1)
-
○ Dissipated energy: 432.12 kJ.C-mol-Biomass−1
-
-
Denitrifying acetotroph:
-
○ Catabolism: C2H3O2− + 1.6 NO3− + 0.6 H+ → 0.8 N2 + 2 HCO3− + 0.8 H2O (ΔG0′ = −792.1 kJ.molDonor−1)
-
○ Anabolism: 0.503 C2H3O2− + 0.158 NH4+ + 0.338 H+ → 0.4305 H2O + 0.0063 HCO3− + C1H1.613O0.557N0.158 (ΔG0′ = 23.9 kJ.C-mol-Biomass−1)
-
○ Dissipated energy: 432.12 kJ.C-mol-Biomass−1
-
-
Iron reducing acetotroph:
-
○ Catabolism: C2H3O2− + 4 H2O + 8 Fe+3 → 9 H+ + 2 HCO3− + 8 Fe+2 (ΔG0′ = −809.6 kJ.molDonor−1)
-
○ Anabolism: 0.503 C2H3O2− + 0.158 NH4+ + 0.338 H+ → 0.4305 H2O + 0.0063 HCO3− + C1H1.613O0.557N0.158 (ΔG0′ = 23.9 kJ.C-mol-Biomass−1)
-
○ Dissipated energy: 432.12 kJ.C-mol-Biomass−1
-
-
Ammonium oxidizing bacteria (AOB):
-
○ Catabolism: NH4+ + 1.5 O2 → 1 NO2− + 1 H2O + 2 H+ (ΔG0′= -269.9 kJ.molDonor‒1)
-
○ Anabolism: HCO3− + 0.828 NH4+ → 1.101 H2O + 0.670 NO2− + 0.499 H+ + C1H1.613O0.557N0.158 (ΔG0′ = 267.7 kJ.C-mol-Biomass‒1)
-
○ Dissipated energy: 3500 kJ.C-mol-Biomass‒1
-
-
Nitrite oxidizing bacteria (NOB):
-
○ Catabolism: NO2− + 0.5 O2 → 1 NO3−(ΔG0′ = ‒79.1 kJ.molDonor‒1)
-
○ Anabolism: HCO3− + 2.64 NO2− + 1.16 H+ → 0.27 H2O + 2.49 NO3− + C1H1.613O0.557N0.158 (ΔG0′ = 241.3 kJ.C-mol-Biomass‒1)
-
○ Dissipated energy: 3500 kJ.C-mol-Biomass−1
-
The ΔG0′ values indicated here correspond to standard Gibbs free energy changes corrected for a realistic H+ concentration of 1e−7 mol.L−1 at pH = 7. In the simulations, Gibbs free energy calculations are refined to account for concentrations of all chemical species. For each metabolism, the linear combination of the catabolic and the anabolic reaction gives what is assumed to be the overall growth reaction of the population considered.
Virtual culture conditions
The temperature is set to 298.15 K and the pH is assumed to be 7 in every system, unless specified otherwise.
Aerated batch cultures in minimal M9 medium
The culture medium used in the simulations is a minimal medium homologous to the M9 minimal medium. The concentrations of chemical species used for model initialization are glucose: 17.05 mM, proton: 3.98e−5 mM (pH = 7.40), ammonium: 18.69 mM, bicarbonate: 0 mM. When using an alternative carbon source, the same quantity of molC is used to enable yield comparison. Thus the concentration of acetate is 51.15 mM. Initial microbial inoculation was set to 1 mM (22.7e6 cell mL−1).
The only abiotic process implemented in aerated batch systems is aeration. Its rate is \(k_La \ast \left( {\left[ {O_2} \right]_{{\mathrm {sat}}} - \left[ {O_2} \right]} \right)\) where [O2] is the current oxygen concentration, [O2]sat the saturation concentration of oxygen in water (0.273 mM at the temperature of the system according to the Henry law) and kLa the oxygen transfer coefficient (time−1), set to 100 day−1 for the simulations. The stoichiometric coefficients of the aeration process consist in a r × 1 Aaeration matrix. The coefficient of Aaeration for O2 is 1, and 0 for every other reagent.
Chemostat culture system
The system is submitted to chemostat dynamics; the rate of matter transfer in a chemostat system is d * (Csat − C) [6] where Csat is the r × 1 matrix of the concentration of the concentration in the input of the chemostat and d is the dilution rate of the chemostat (input and output flow rate divided by the tank volume) in time−1. The stoichiometric coefficient of matter transfer on the reagent concerned is 1. The chemostat culture system is modeled as a 1 m3 perfectly mixed single compartment. It is subjected to chemostat dynamics with a flow rate of 1 m3 day−1. No aeration process is implemented in this system. Oxygen is fed to the culture medium through the chemostat inflow; its influent concentration is 2.73e−1 mM (8.73 g m−3, which is its saturation concentration). Input concentrations used for this system are meant to emulate a likely groundwater: ammonium 1 mM (18.04 g. m−3), bicarbonate 1 mM (61.02 g. m−3), nitrate 1.93e−1 mM (12 g.m−3), sulfate 6.24e−1 mM (60 g m−3), Fe+3 5.37e−1 mM (30 g. m−3). Multiple simulations were run using acetate concentrations ranging from 0 to 1.5 mM (88.56 g. m−3). The initial microbial inoculation was set to 1 mM (22.7e6 cell. mL−1) for each microbial guild. The simulation was run until biomass stabilized.
Activated sludge system
The activated sludge system is modeled as an aerated batch. Aeration conditions are identical to those used for aerated batch cultures in M9 medium, except that the oxygen saturation concentration is set to 0.2556 mol. m−3, according to an empirical relationship used for wastewater modeling [42]. The composition of the medium is set to simulate a simplified filtered urban wastewater containing 1.76 mM acetate (103.9 g m−3) and 3.78 mM ammonium (68.19 g m−3). The initial microbial inoculation was set to 1 mM (22.7e6 cell mL−1) for each microbial guild. Values of Vh were common to all guilds and set to 10 m3C-mol-Biomass−1. For the sake of simplicity, the influent was considered to be free of particles and only aerobic growth of microbial populations was modeled.
Results
Modeling a pure culture growing in a minimal medium containing multiple elemental resources
To show how energy balance, stoichiometry and microbial dynamics are inherently coupled in our modeling framework, here we present model implementation in the simplest case of a single population growing in a minimal medium. The results enable analysis of the model’s dynamic properties.
We consider the growth of an axenic culture in a minimal M9 medium, in an aerated batch inoculated with 1 mM biomass (22.7e6 cell. mL−1). A single glucosotroph guild (see Methods) is simulated.
The metabolism of the population consists in the combination of an anabolic and a catabolic reaction and can be expressed as a function of λ;
−0.167 C6H12O6 −0.158 NH4+ + 0.430 H2O + 0.164 H+ + 0.00625 HCO3− + 1 C1H1.613O0.557N0.158 + λ (−1 C6H12O6 − 6 O2 + 6HCO3− + 6 H+) = 0
where λ can be calculated at each time step according to anabolic, catabolic, and dissipated Gibbs free energy variations is
(see Eq. 3 in Methods for details)
The growth rate μ of the population is then expressed as a function of its substrates (namely chemical species having a negative stoichiometric coefficient): \(\mu = \mu _{{max}}\,{\prod}_{i} e^{\frac{{A_{{met},i}}}{{V_{{h}}[ {S_i} ]}}}\)
where μmax is a constant that is independent of the microbial population considered (see Methods), Amet,i is the negative stoichiometric coefficient of the substrate Si, Vh is the harvest volume (independent of substrate and population, see Methods) and [Si] is the concentration of the substrate. In the following description of the MTS model predictions, each term \(e^{\frac{{A_{\mathrm {met},i}}}{{V_{\mathrm {h}}\left[ {S_i} \right]}}}\) of Eq. 5 for each substrate will be called “tuning factor of substrate i”. For each substrate, this tuning factor corresponds to the inhibition it exerts on the growth rate depending on the ratio of its demand to its supply in the culture medium. During the course of the simulation, the Gibbs free energy changes in the anabolic and catabolic reactions vary according to the changes in temperature and in the concentration of the reagents over time. The λ factor preserves the energy balance of the metabolism as a whole and is consequently dynamically adjusted. This process ensures the dynamic coupling of microbial growth rates, stoichiometry, and energy balance at each time step of the simulation, according to changes in the concentration of the reagents over time.
Figure 1a tracks the concentrations in the system during the dynamic simulation. The simulated population grows linearly until it reaches a plateau at 74.47 mM (1.69e9 cell mL−1) in 30 h (1.25 days). Figure 1a also shows the tuning factors of the population, which help understand population dynamics in the MTS model; in the current simulation, they indeed reveal a dynamic, two-step growth limitation phenomenon. In the beginning, as glucose is abundant in the culture medium, the growth rate is mostly limited by oxygen, the electron acceptor. As the aeration rate is constant, the population grows according to this linear aeration rate and depletes glucose and ammonium. At the 30th hour of the simulation, the concentration of glucose reaches a level at which it becomes significantly limiting; its tuning factor plummets from 0.98 to 8e−23 in <5 h (0.2 day). As glucose becomes scarce, the growth rate of the population decreases exponentially as a result of the limitation of the electron donor, to reach 3.3e−6 day−1 at the end of the simulation. With such a low metabolic rate, aeration replenishes the oxygen concentration in the batch culture almost to saturation. Consequently, the tuning factor of oxygen (electron acceptor) increases during the glucose-limited phase of growth, and glucose (electron donor) becomes the main limiting substrate. Conversely, simulations were performed using exactly the same model structure and only changing the initial nitrogen concentration to represent growth in a nitrogen-poor M9 medium. In that case, the growth rate appears to be limited by oxygen in the first phase, after which growth virtually ceases due to ammonia exhaustion (Fig. 1c). These simulations reveal that the MTS model can jointly capture the influence of all resources on growth dynamics, whether these resources contribute to the energy supply of the cell (electron donors or acceptors) directly or not (nutrients such as ammonia). It should be outlined that dynamic patterns exhibiting more progressive start-up phases can be obtained using different parameter values (see supplementary materials 4).
Aerobic microbial cultures growing in different conditions. Graphs on left column are concentrations over time; acetate concentration in dark green, glucose concentration in light green, ammonium concentration in dark blue, oxygen concentration in light blue, acetotroph biomass concentration in plain black, glucosotroph biomass concentration in dashed black. Graphs on right column are natural logarithm of the tuning factors over time; tuning factors follow the same color scheme as in the left column; tuning factors for acetotroph guild are represented by plain lines, tuning factors for glucosotroph guild are represented by dashed lines. The lower a tuning factor is, the more significant is the limitation exerted by the substrate on the microbial population’s growth rate. Time is in hour, concentration is in mM, secondary concentration axis (right side) on the left column is for oxygen. Biomass is quantified as mM of carbon-normalized biomass. a Glucosotroph guild monoculture. b Acetotroph guild monoculture. c ammonium-limited glucosotroph guild monoculture. d Acetotroph and glucosotroph co-culture
Modeling two competing populations: emergence of an energy-driven ecological succession
One of the factors that influence community structure is the outcome of competition between populations. The MTS model’s ability to predict the outcome of competition between multiple populations in the sense of functional guilds has been questioned.
To answer this question, we implemented the minimalist case of two heterotrophic populations growing together in an aerated batch, modeled as a glucose oxidizing guild and an aerobic acetate oxidizing guild. Both guilds have the same absolute μmax value, and the same Vh value. The difference between the dynamics of the two guilds only resulting from thermodynamic constraints was then studied.
The two guilds differ in the energy they dissipate during their respective metabolic reaction according to Heijnen’s formula; −432.12 kJ C-mol-Biomass−1 for acetate oxidizers vs. −236.05 kJ.C-mol-Biomass−1 for glucose oxidizers. However, the number of electrons donated per carbon atom of electron donor is the same (4 electrons per carbon atom of electron donor). The initial quantity of each electron donor is 17.05 mM of glucose and 51.15 mM of acetate. These quantities were adjusted in this simulation so that they represented the same quantity of carbon (and consequently the same quantity of electrons) in the raw element count.
Figure 1d tracks the system’s variables over the course of the simulation. The model predicts that the glucose oxidizer guild grows first. Their population is predicted to stabilize at 74.47 mM (1.69e9 cell mL−1) after 31 h (1.29 days). The population of acetate oxidizers is predicted to grow from the 31st to the 84th hour (1.3–3.5 days) and to stabilize at 50.19 mM (1.14e9 cell mL−1). The growth of the glucose oxidizer guild follows the same kinetics and the same sequential inhibition as in the previous, mono-guild simulation (Fig. 1b). The growth of the acetate oxidizer guild is also sequentially inhibited by its electron acceptor (oxygen) then by its electron donor (acetate).
As the two guilds have a different electron donor (glucose and acetate) but the same electron acceptor (oxygen), they compete for the electron acceptor. The results shows that the guild of glucose oxidizers grows first and also develops a higher stabilized population density than the acetate oxidizers guild (Fig. 1d). In presence of the glucose oxidizer guild, the growth of the acetate oxidizer guild is delayed compared to its growth in the absence of the glucose oxidizer guild (Fig. 1b). This outcome arises from the thermodynamic properties of the metabolisms involved. The same amount of oxygen per carbon is needed to oxidize glucose or acetate, in order to preserve the elemental balance, and the batch simulation is initialized with the same quantity of carbon for both substrates. However, in the simulation conditions, glucose oxidation is more exergonic than acetate oxidation when normalized by carbon atom. The guild growing on glucose does not need to dissipate as much energy. These two properties are aggregated and related to the growth yields by the λ factor, more acetate than glucose must be oxidized in order to produce the energy required for growth (approximately 0.503 + 1 × 0.6 = 1.1 mol-Acetate.mol-Biomass−1 vs. approximately 0.167 + 1 × 0.09 = 0.25 mol-Glucose.mol-Biomass−1). Consequently, the tuning factor associated with oxygen is lower for acetate oxidizers than for glucose oxidizers (Fig. 1d). Both guilds reduce the oxygen concentration while growing, but the thermodynamic considerations implemented by the MTS model give a better yield to the glucose oxidizers. Hence they can draw down O2 to lower levels (compared with acetate oxidizers). Consequently the growth of glucose oxidizers brings the oxygen concentration to a level at which the growth rate of acetate oxidizers is insignificant and in practice prevents their growth during the first 30 h (1.25 days) of the simulation. This effect might be less pronounced when using different parameter values, as illustrated in supplementary materials 5. Interestingly, the possibility to obtain such a pattern could be experimentally tested in a system where the oxygen supply rate would be less than the oxygen uptake rate of the glucosotrophic guild.
It should be noted that the two populations have exactly the same growth parameter values implemented in the MTS model. Despite similar kinetic parameters for the two populations, a microbial succession emerges from the model. Therefore, simple mass and energy balance calculations coupled to the flux-force relationship between energy and rate determine the outcome of competition and an ecological succession emerges. Our model thus appears to inherently exhibit an original property: the ability to account for microbial successions as a result of competition for available resources and energy. In view of this result, the possibility to directly account for community assembly patterns observed in nature was further questioned.
Modeling competition between multiple populations: emergence of a community structured according to a “redox tower” of microbial metabolism
Natural systems in which electron donors are available in excess and electron acceptors are limited exhibit typical compartmentalization of microbial activities. This occurs particularly during hypolymnia of eutrophic freshwater lakes, when oxygen depletion results in anoxic zones [43]. This phenomenon has been studied for decades [44, 45]. It leads to sequential vertically stratified consumption of available electron acceptors according to the hierarchy of Gibbs free energy changes of half reduction reactions, leading to “redox towers of microbial metabolisms”. To assess the ability of our approach to account for such compartmentalization patterns, we simulated competition between multiple populations for a single electron donor (acetate) and in the presence of various electron acceptors. The competition was analyzed in a chemostat. In these systems, microbial populations are subjected to a continuous flow of substrate and an output rate for biomass. Each population then has two possible stationary states: either its growth rate negates the dilution rate or the population is washed out. We implemented a microbial community channeling electrons from acetate to oxygen, nitrate, sulfate and iron, and we simulated its dynamics according to the MTS model. The implemented guilds are aerobic acetotroph, denitrifying acetotroph, sulfate reducing and iron reducing guilds (see Methods).
The culture medium is modeled as a perfectly mixed 1 m3 single compartment subjected to chemostat dynamics with a flow rate of 1 m3.day−1, corresponding to a dilution rate of 1 day−1.
Figure 2 shows the concentration of each guild during the stationary phase, as a function of the influent acetate concentration. The concentration of all populations as a function of time for an input acetate concentration of 1.76 mM is shown as supplementary materials 6. As previously shown, the growth of the most thermodynamically advantaged guild delays the growth of competing guilds. The population able to adjust its growth rate to the dilution rate with the minimal amount of acetate is the aerobes guild. This is explained by the superior exergonicity of their metabolism as discussed in connection with the competition between two species above. With increasing acetate concentrations, aerobes coexist with nitrate reducers, then iron reducers, then sulfate reducers. Indeed, when the quantity of electrons provided in the form of acetate exceeds the quantity of electrons acceptable by a given acceptor (i.e. oxygen, nitrate, iron, or sulfate), some unoxidized acetate molecules remain in the culture medium. These acetate molecules are available for a guild catalyzing a less exergonic reaction. Therefore the complexity of the community increases with an increase in the incoming electron donor molecules in the system, as the possible electron acceptors are saturated one by one. A community structure pattern compliant with the typical redox tower pattern is thus parsimoniously predicted by the MTS model.
Competition outcome in a chemostat fed with acetate in the presence of various electron acceptors as predicted by the MTS model. The proportion of each guild at stationary state is depicted as a function of the acetate input concentration (mM). Simulation results predict patterns ranging from competitive exclusion at low acetate concentration to full coexistence at non-limiting acetate concentrations
Towards more predictive models for environmental biotechnology applications: microbial dynamics in a simplified activated sludge community
To illustrate the use of the MTS model in the practical case of environmental bioprocess modeling, we implemented a simplified model of the community found in the aerated tanks of wastewater treatment plants (WWTPs). A set of engineering models called “Activated Sludge Model” (ASM) is commonly used for the operation, design and optimization of WWTPs [46]. These models focus on the accuracy of the system state variables. This accuracy depends on how closely the simulated system resembles the reference system from which a priori knowledge is inputted through calibration. The calibrated parameters are thus bound to a range of experimental conditions. On the contrary, the MTS-based approach we present here does not rely on calibration based on experimental data: we derived kinetics and yields directly, i.e., without calibration, from fundamental generic assumptions, mass, and energy balance calculations.
An aerated batch incubation containing wastewater and a simplified activated sludge inoculum was then simulated using the MTS and ASMN models. The ASMN model is a modified version of the ASM no. 1 model [47], with nitrification split into two steps between two guilds [42]. This model thus splits the microbial community into three functional guilds:
-
○ Ordinary heterotrophic organisms (OHO): heterotrophs consuming an unspecified carbonaceous substrate
-
○ Ammonium oxidizing bacteria (AOB): autotrophs oxidizing ammonium to nitrite
-
○ Nitrite oxidizing bacteria (NOB): autotrophs oxidizing nitrite to nitrate
The model was implemented with default parameters (for more details, see supplementary materials 7). The MTS model was implemented with the same three guilds and the two generic parameters μmax and Vh described in Methods. As a working hypothesis, we assumed that the electron donor of the OHO guild is acetate, as it appears to be the most abundant volatile fatty acid in wastewater [48]. Simulation results are shown in Fig. 3, where the predictions of the MTS model are put into perspective by comparing them with simulation results obtained with a default calibration of parameters for the ASMN model (see supplementary materials 7). Despite overall similarity, the two simulations exhibit some differences. The main difference is that the three guilds grow simultaneously in the ASMN simulation and sequentially in the MTS model. This difference is linked to the structure of each model. In the ASMN model, growth depends on the combination of Monod-type affinity functions so that a population can grow as soon as its substrates are present in the medium. In the MTS model, the growth of heterotrophs leads to oxygen limitations that virtually prevent the growth of the two autotrophic guilds AOB and NOB in a similar way to that described in previous simulations. Moreover, the low exergonicity of the nitrite oxidation reaction requires that a sufficient amount of nitrite accumulates in the batch before NOB can compete with AOB for oxygen, leading to a sequential growth pattern and transient nitrite accumulation. This pattern has already been documented in piggery wastewater treatment [49]. This sequential growth phenomenon is also apparent in the oxygen concentration profiles, where the three stage consumption pattern is most apparent in the MTS simulation. Despite these differences, the simulations exhibit remarkable similarities. At the end of the simulation, the yield of each guild is similar in the two models: the final quantities of biomass in the batch according to ASMN are 108.4 gCOD.m−3 OHO, 50.63 gCOD.m−3 AOB and 45.07 gCOD.m−3 NOB, while the MTS model predicts 114.8 gCOD.m−3 OHO, 52.4 gCOD.m−3 AOB and 45.1 gCOD.m−3 NOB (Fig. 3). These results show that the abundances of each guild as predicted by thermodynamic rules are consistent with the growth yields measured in activated sludge and used for calibration in the ASMN model. The yields predicted by the MTS model depend on the energy dissipated by the metabolisms, which was computed using the empirical formula that Heijnen and collaborators [24] calibrated on experimental culture data. This is why the yields predicted by the MTS model match those of the ASMN model. In particular, the calibration process used to produce the empirical yield parameter of the ASMN model’s heterotrophic population captures the average yield of many thermodynamically constrained populations growing on diverse substrates. The average yield resulting from all these metabolisms is close to the yield of growth on acetate, which is one of the most abundant sources of carbon in such systems.
Concentration of main chemical species and biomass over time for a simplified activated sludge community in an aerated batch reactor. Left column shows the prediction from ASM; right column shows the predictions from the MTS model. The units used to express the concentration of the chemical species are the same as in the ASM models
Discussion
In this article, we present the dynamics of single and multiple microbial populations arising directly from the MTS theory [38] coupled to a thermodynamic and stoichiometric balance calculation framework developed in previous papers (reviewed in ref. [28]). While several approaches have been proposed in the past to link thermodynamic balance calculations and kinetics [28,24, 33,34,35, 50], the MTS theory differs fundamentally from all these previous approaches. What is completely novel is that it makes population dynamics emerge as the statistical outcome of all the individual division events described at the microscopic level, from fundamental and generic principles. Strikingly, despite the simplicity of the theory’s core principles, consistent microbial dynamics, successions and functional community assembly patterns were simulated without population-specific parameter calibration, for systems ranging from pure to mixed cultures. In this article, we therefore document that the kinetic equations arising from MTS theory intrinsically include many important generic properties to adequately model microbial population and community dynamics.
First, using a stoichiometric approach, we show how the model jointly captures the influence of all substrates simultaneously (see Eq. 5). However, the simulation results also illustrate how growth dynamics actually appear to be limited by one substrate at a given time. This emerging property of the model recalls Liebig’s law of the minimum, which states that the growth rate of an organism requiring multiple different resources is controlled by the scarcest resource only. This situation indeed corresponds to the widely accepted intuitive understanding of the way microbial cultures behave in the laboratory, and has often been investigated, tested and confirmed [51]. Liebig’s hypothesis is thus widely used to model microbial dynamics, sometimes implicitly, as in the most simple, mono-substrate expression of Monod’s equation, sometimes explicitly, as in multi-substrate implementations of the Monod model where the limiting substrate has to be selected at each time step by computing the minimum of all resource-dependent growth factors to tune down the maximum growth rate [52]. Without introducing the Liebig hypothesis, considering the growth dependence of multiple substrates in Monod based models often leads to inconsistent growth patterns that require extensive adaptation of model structure [53,54,55]. In our case, the MTS model can produce sequential growth limitation either by electron donor, acceptor or by nutrient. The exponential nature of the relationship makes tuning factors “rise” from insignificant to significant in a narrow concentration range (as seen in Fig. 1), thus allowing pronounced limitation switches. Under the simple conditions simulated, virtual cultures thus exhibit “Liebig like behavior” that is obtained parsimoniously, as an emerging property of the model. The modeling framework we propose is therefore a simple and elegant way to jointly capture the effect of electron donor, acceptor and nutrient concentrations on microbial dynamics, without infringing Liebig’s hypothesis.
Second, the simulations involving several populations in competition either for electron donor or acceptor result in community assembly patterns structured like microbial redox towers (see Fig. 2), as observed in many natural habitats such as the anoxic hypolymnia of eutrophic lakes [45]. These types of patterns have already been generated using Monod-based kinetic models [30, 56]. However, our contribution differs significantly from previous works regarding the scientific conclusions that can be drawn from the simulations.
Gonzalez-Cabaleiro’s model [57] uses a modified Herbert-Pirt equation in which the growth yield is a function of thermodynamic variables. The kinetic equation used to describe substrate consumption rate is a Monod-like function. This model is therefore a juxtaposition of a framework for the calculation of the thermodynamic balance and a phenomenological description of the dynamic growth process that results in ecological successions. Since kinetic parameters were not calibrated in Gonzalez–Cabaleiro’s article, their simulations indeed show the phenomenological dependence of ecological successions on thermodynamic variables. However their simulations provide no support for using Monod’s law itself as a way to reproduce this pattern. Indeed, since Monod’s law is an empirical equation, it carries no hypotheses per se; any similarly shaped curve would have provided the same simulation results. In contrast, in the case of the MTS model, the thermodynamic variables are used to compute the division probability at the level of each individual. The MTS theory then provides a mechanistic explanation of the influence of energy on microbial division. The sum of all individual division events then propagates at the scale of the population, resulting in the MTS growth equation. Therefore, both the role of thermodynamic variables and the hypotheses on how they are mechanistically linked to the growth rate receive support from the simulation. Our simulations therefore show that the mechanistic explanation of the influence of energy on microbial division at the heart of the MTS theory is sufficient to explain ecological successions.
An interesting parallel can then be drawn with Jin and Bethke’s work [33], since both their model and the MTS model propose a linkage between microbial kinetics and thermodynamic variables, based on two totally different theories (a probabilistic reasoning for the MTS model, non-equilibrium thermodynamics for Jin and Bethke’s model). Both models are fundamentally different since Jin and Bethke’s model expresses the rate of catabolism, while the MTS model expresses the rate of biomass synthesis. Jin and Bethke’s model therefore cannot account for the limitation of growth by nutrients as the MTS model does (Fig. 1c). However, the fact that two theory-based kinetic models provide alternative formulations of microbial growth is interesting, and the careful comparison of the models may provide fruitful insights into microbial growth kinetics.
Regarding the simulation of a simplified activated sludge ecosystem, the MTS model was shown to make predictions qualitatively similar to those of an engineering model (namely, implementation of the ASM). To implement the system whose simulation is depicted in Fig. 3, the ASM model requires identification of nine kinetic parameters along with three growth yield parameters, whose values have been carefully adjusted during decades of experimentation [58]. In contrast, the kinetics obtained in the MTS approach emerges from a theoretical construct in which the only two parameters (µmax and Vh) common to all guilds were considered. While allowing population-specific values for Vh could indeed improve the quantitative accuracy of MTS simulations (for an illustration see supplementary materials 8), we considered default parameters’ values in the simulations displayed in Fig. 3 to show that our model is able to generate consistent dynamics with a remarkably small number of parameters, suggesting that the energy/rate dynamic equation resulting from the MTS theory already has interesting predictive abilities in the absence of parameter fitting.
Clearly, the community dynamics obtained by simulation corresponds to those of an idealized functional partitioning of the community resulting from a first layer of energetic and stoichiometric drivers. As implemented in this article, the MTS model obviously does not account for the whole range of phenomena that influence microbial community dynamics and community structure in real complex environmental or engineered settings like those encountered in activated sludge plants. Other processes such as inhibition due to compound toxicity, indirect pH and temperature adaptation effects [59], non-metabolic inter-species interactions, spatial organization [60], etc. need to be added to obtain a more realistic picture of genuine complex microbial community dynamics. However, including such phenomena in the model was beyond the scope of the present study. The microbial dynamics simulated in this article were indeed not the result of an effort to reproduce specific experimental patterns through the calibration of empirical equations. Rather, the most salient message is to document and analyze the consistency of generic growth dynamics and community assembly patterns as emerging directly from a new kinetic theory of microbial growth relying on first principles coupled with thermodynamic and stoichiometric calculations.
To that extent, the simulations reported here illustrate a set of key properties of MTS dynamic models. To our knowledge, a microbial population dynamics model per se has never before exhibited all these properties, that is, without the need for additional hypotheses or specific parameter calibration. What is of the utmost importance and constitutes the novelty of our contribution, is the fact that these properties are obtained parsimoniously through the combination of fundamental and generic principles translated into mathematical equations, and not from the calibration of population-specific parameters. More generally, we advocate the need in microbial ecology to propose new theoretical abstractions to grasp a whole category of phenomena in an inclusive picture. We believe that such approaches pave the way for a new class of microbial ecology and engineering models, built on more robust theoretical foundations and exhibiting enhanced predictive abilities.
References
Whitman WB, Coleman DC, Wiebe WJ. Prokaryotes: the unseen majority. Proc Natl Acad Sci USA. 1998;95:6578–83.
Falkowski PG, Fenchel T, Delong EF. The microbial engines that drive Earth’s biogeochemical cycles. Science. 2008;320:1034–9.
Rodríguez J, Lema JM, Kleerebezem R. Energy-based models for environmental biotechnology. Trends Biotechnol. 2008;26:366–74.
Verstraete W. Microbial ecology and environmental biotechnology. ISME J. 2007;1:4–8.
Widder S, Allen RJ, Pfeiffer T, Curtis TP, Wiuf C, Sloan WT et al. Challenges in microbial ecology: building predictive understanding of community function and dynamics. ISME J. 2016;10: 2557–68.
Monod J. The growth of bacterial cultures. Annu Rev Microbiol. 1949;3:371–94.
Contois. D. E. Kinetics of Bacterial Growth: Relationship between Population Density and Specific Growth Rate of Continuous Cultures. Journal of General Microbiology, 1959;21:40–50.
Andrews. J.F. A mathematical model for the continuous culture of microorganisms utilizing inhibitory substrates. Biotechnol Bioeng. 1968;10:707–23.
Louca S, Jacques SMS, Pires APF, Leal JS, Srivastava DS, Parfrey LW, et al. High taxonomic variability despite stable functional structure across microbial communities. Nat Ecol Evol. 2016;1:15.
Louca S, Parfrey LW, Doebeli M. Decoupling function and taxonomy in the global ocean microbiome. Science. 2016;353:1272–7.
Raes J, Letunic I, Yamada T, Jensen LJ, Bork P. Toward molecular trait-based ecology through integration of biogeochemical, geographical and metagenomic data. Mol Syst Biol. 2011;7:473–3.
Kaiser K, Wemheuer B, Korolkow V, Wemheuer F, Nacke H, Schöning I, et al. Driving forces of soil bacterial community structure, diversity, and function in temperate grasslands and forests. Sci Rep. 2016;6:33696–6.
Nelson MB, Martiny AC, Martiny JBH. Global biogeography of microbial nitrogen-cycling traits in soil. Proc Natl Acad Sci USA. 2016;113:8033–40.
Ju F, Guo F, Ye L, Xia Y, Zhang T. Metagenomic analysis on seasonal microbial variations of activated sludge from a full-scale wastewater treatment plant over 4 years. Environ Microbiol Rep. 2014;6:80–9.
De Filippis F, Genovese A, Ferranti P, Gilbert JA, Ercolini D. Metatranscriptomics reveals temperature-driven functional changes in microbiome impacting cheese maturation rate. Sci Rep. 2016;6:1–2.
Huttenhower C, Gevers D, Knight R, Abubucker S, Badger JH, Chinwalla AT, et al. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486:207–14.
Hansson L. Why ecology fails at application: should we consider variability more than regularity? OIKOS. 2003;100:624–7.
Lawton JH. Are there general laws in ecology? OIKOS. 1999;84:177–92.
McGill BJ, Enquist BJ, Weiher E, Westoby M. Rebuilding community ecology from functional traits. Trends Ecol Evol. 2006;21:178–85.
Simberloff D. Community ecology: is it time to move on? (An American Society of Naturalists Presidential Address). Am Nat. 2004;163:787–99.
Odum E. P. The strategy of ecosystem development. Science, New series, 1969;164:262--70.
McCarty PL. Thermodynamics of biological synthesis and growth. Air Water Pollut. 1965;9:621–39.
Heijnen JJ, Dijken JPV. In search of a thermodynamic description of biomass yields for the chemotrophic growth of microorganisms. Biotechnol Bioeng. 1991;39:833–58.
Heijnen JJ, Kleerebezem R. Bioenergetics of microbial growth. In: Flickinger MC, editors. Encyclopedia of industrial biotechnology: bioprocess, bioseparation and cell technology. John Wiley & Sons, Inc; 2010. pp. 1–24.
McCarty PL. Thermodynamic electron equivalents model for bacterial yield prediction: modifications and comparative evaluations. Biotechnol Bioeng. 2007;97:377–88.
Roels JA. Application of macroscopic principles to microbial metabolism. Biotechnol Bioeng. 1980;103:2–59. discussion 51
von Stockar U, Liu J. Does microbial life always feed on negative entropy? Thermodynamic analysis of microbial growth. Biochim Biophys Acta. 1999;1412:191–211.
Kleerebezem R, Van Loosdrecht MCM. A generalized method for thermodynamic state analysis of environmental systems. Crit Rev Environ Sci Technol. 2010;40:1–54.
von Stockar U, Vojinović V, Maskow T, Liu J. Can microbial growth yield be estimated using simple thermodynamic analogies to technical processes? Chem Eng Process: Process Intensif. 2008;47:980–90.
Gonzalez-Cabaleiro R, Ofiteru ID, Lema JM, Rodriguez J. Microbial catabolic activities are naturally selected by metabolic energy harvest rate. ISME J. 2015;9:2630–41.
González-Cabaleiro R, Lema JM, Rodríguez J, Kleerebezem R. Linking thermodynamics and kinetics to assess pathway reversibility in anaerobic bioprocesses. Energy Environ Sci. 2013;6:3780–80.
González-Cabaleiro R, Lema JM, Rodríguez J. Metabolic energy-based modelling explains product yielding in anaerobic mixed culture fermentations. PLoS ONE. 2015;10:1–17.
Jin Q, Bethke CM. A new rate law describing microbial. Respiration. 2003;69:2340–8.
Noguera DR, Brusseau GA, Rittmann BE, Stahl DA. A unified model describing the role of hydrogen in the growth of desulfovibrio vulgaris under different environmental conditions. Biotechnol Bioeng. 1998;59: 732–746.
Hoh CY, Cord-Ruwisch R. A practical kinetic model that considers endproduct inhibition in anaerobic digestion processes by including the equilibrium constant. Biotechnol Bioeng. 1996;51:597–604.
Button DK. Nutrient uptake by microorganisms according to kinetic parameters from theory as related to cytoarchitecture. Microbiol Mol Biol Rev. 1998;62:636–45.
Liu Y. A simple thermodynamic approach for derivation of a general Monod equation for microbial growth. Biochem Eng J. 2006;31:102–5.
Desmond-Le Quéméner E, Bouchez T. A thermodynamic theory of microbial growth. ISME J. 2014;8:1747–1751.
Battley E. The development of direct and indirect methods for the study of the thermodynamics of microbial growth 1. Thermochim Acta. 1998;309:17–37.
Milo R, Phillips R. Cell biology by the number. Garland Science 2015:358.
Eyring H. The activated complex in chemical reactions. J Chem Phys. 1935;3:107–15.
Hiatt WC, Grady CPL. An updated process model for carbon oxidation, nitrification, and denitrification. Water Environ Res. 2008;80:2145–56.
Müller B, Bryant LD, Matzinger A, Wüest A. Hypolimnetic oxygen depletion in eutrophic lakes. Environ Sci Technol. 2012;46:9964–71.
Hutchinson GE. A treatise on limnology: geography, physics, and chemistry. pt. 1. Geography and physics of lakes. John Wiley & Sons Inc; 1957. p. 1016.
Boehrer B, Schultze M. Stratification lakes. Rev Geophys. 2008;46:1–27.
Hauduc H, Gillot S, Rieger L, Ohtsuki T, Shaw a, Takàcs I, et al. Activated sludge modelling in practice: an international survey. Water Sci Technol. 2009;60:1943–51.
Henze M, Grady CPL, Gujer W, Marais GVR, Matuso T. Activated sludge model no. 1. IAWQ Scientific and Technical Report No. 1; 1987.
Rössle WH, Pretorius Wa. A review of characterisation requirements for in-line prefermenters paper 2: process characterisation. Water SA. 2001;27:413–22.
Rajagopal R, Rousseau P, Bernet N, Beline F. Combined anaerobic and activated sludge anoxic/oxic treatment for piggery wastewater. Bioresour Technol. 2011;102:2185–92.
Westerhoff HV, Hellingwerf KJ, Van Dam K. Thermodynamic efficiency of microbial growth is low but optimal for maximal growth rate. Proc Natl Acad Sci USA. 1983;80:305–9.
Kovarova-Kovar K, Egli T. Growth kinetics of suspended microbial cells: from single-substrate-controlled growth to mixed-substrate kinetics. Microbiol Mol Biol Rev. 1998;62:646–66.
Droop MR. The nutrient status of algal cells in continuous culture. J Mar Biol Assoc UK. 1974;54:825.
Bajpai-Dikshit J, Suresh AK, Venkatesh KV. An optimal model for representing the kinetics of growth and product formation by Lactobacillus rhamnosus on multiple substrates. J Biosci Bioeng. 2003;96:481–86.
Bapat PM, Bhartiya S, Venkatesh KV, Wangikar PP. Structured kinetic model to represent the utilization of multiple substrates in complex media during rifamycin B fermentation. Biotechnol Bioeng. 2006;93:779–90.
Nikolajsen K, Nielsen J, Villadsen J. Structured modeling of a microbial system: III. Growth on mixed substrates. Biotechnol Bioeng. 1991;38:24–29.
Bethke CM, Sanford Ra, Kirk MF, Jin Q, Flynn TM. The thermodynamic ladder in geomicrobiology. Am J Sci. 2011;311:183–10.
González-Cabaleiro R, Ofiţeru ID, Lema JM, Rodríguez J. Microbial catabolic activities are naturally selected by metabolic energy harvest rate. ISME J. 2015;9:2630–41.
Gujer W, Morgens I, Mino T, Van Loosdrecht MCM. Activated sludge models asm1, asm2, asm2d and asm3. London: IWA Publishing; 2000.
Shammas N. Interactions of temperature, pH, and biomass on the nitrification process. Water Pollut Control Fed. 1986;58:52–59.
Meister M, Winkler D, Rezavand M, Rauch W. Integrating hydrodynamics and biokinetics in wastewater treatment modelling by using smoothed particle hydrodynamics. Comput Chem Eng. 2017;99:1–2.
Acknowledgements
The authors thank D. Bart Haegeman for the very useful comments he made on a preliminary version of this manuscript. The authors are also grateful to Région Ile de France for funding Hadrien Delattre’s PhD in the framework of the DIM R2DS project. The authors thank the French “Agence Nationale de la Recherche” for its financial support through the “THERMOMIC” project ANR-16-CE04-0003-01.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Delattre, H., Desmond-Le Quéméner, E., Duquennoi, C. et al. Consistent microbial dynamics and functional community patterns derived from first principles. ISME J 13, 263–276 (2019). https://doi.org/10.1038/s41396-018-0272-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0272-0
This article is cited by
-
Functional convergence in slow-growing microbial communities arises from thermodynamic constraints
The ISME Journal (2023)
-
Hydrothermal plumes as hotspots for deep-ocean heterotrophic microbial biomass production
Nature Communications (2021)
-
Proto-dolomite formation in microbial consortia dominated by Halomonas strains
Extremophiles (2019)