Abstract
The historical conditions under which bacterial lineages evolve determine their functional traits, and consequently their contribution to ecosystem functions (EFs). Under significant trait conservatism, which is common in prokaryotes, phylogeny may track the evolutionary history of species and predict their functionality. Productive communities can arise from: (i) the coexistence of functional, and therefore phylogenetically distant lineages, producing high EF rates at large phylogenetic diversity (PD); (ii) the dominance of productive lineages that outcompete other clades, generating high EF at low PD. Community composition will modulate the PD–EF relationship: The effects of anciently divergent lineages, whose deeply conserved functions determine the occupancy of major niches, may differ from that of recently divergent lineages showing adaptations to current conditions. We hypothesized that, in our model Mediterranean ecosystem, EF can be explained both by competitive superiority of ancient lineages and functional complementarity of recent lineages. To test this hypothesis, we sequenced a phylogenetic marker targeting bacteria across 28 soil plots and quantified EF related to microbial productivity, decomposition and nutrient cycling. Plots accumulating recently divergent lineages consistently showed higher EF levels that were slightly modified by the accumulation of ancient lineages. We discuss the assembly processes behind these phylogenetic-scale disparities and the final outcome in terms of ecosystem functioning.
Similar content being viewed by others
Introduction
The relationship between biodiversity and ecosystem functioning (EF) is a classic topic in ecology that is increasingly incorporating the role of belowground biota [1,2,3]. Soil microbial communities are extraordinarily abundant and diverse components of terrestrial habitats that control essential EFs [4, 5], understood as the properties of ecosystems including both the pools of materials (e.g., stocks of carbon or nutrients) and the rates of processes (e.g., organic matter decomposition, chemical degradation or gas exchange with the atmosphere) [6]. Microbial regulation of EFs depends on a myriad of taxa that share intricate interactions among them and with their environment [1], and whose individual contribution to the overall function is most likely the result of complex combinations of functional traits ([7]; Fig. 1a).
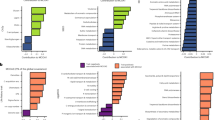
a Bacterial taxa’s individual contributions to a specific ecosystem function (EF) depends on a combination of functional traits that are conserved at varying depths across the phylogeny. EFs include both pools of materials and rates of processes. As an illustration, soil microbial productivity may depend on: T1: Oxidative respiration; T2: Growth efficiency in response to C substrates; T3: Use of glucose; T4: Use of lactose; T5: Use of glucuronamide [9, 20]. Exploring the community composition in terms of anciently or recently divergent lineages may enhance EF predictions. In the example, three ancient lineages mainly differ in their ability to respire aerobically or anaerobically (T1) and in their growth efficiency (T2), whereas eight recent lineages further distinctly use single C sources (T3–T5). For each taxon, EF is the sum of all trait values (i.e., non-blank cells). Numbers in the phylogenetic tree indicate branch lengths, in Million years. b Microbial communities with high EF levels can be assembled through niche or fitness differences, which respectively lead to high or low phylogenetic diversity (PD), and generating positive or negative PD–EF relationships. The phylogenetic depth of trait conservatism may also determine the magnitude of the PD–EF relationship. Left upper panel: The ability to consume glucose either using aerobic or anaerobic respiration may allow the coexistence of distant lineages, increasing PD and also EF due to the high functional complementarity of community members. Left lower panel: The ability to use distinct specific C substrates (e.g., lactose and glucuronamide) may allow the coexistence of closely related aerobic heterotrophs. Right panel: The best competitors in terms of growth efficiency in response to carbon sources overgrow and outcompete distant relatives, decreasing PD while increasing EF. The level of trait conservatism, e.g. use of glucose (upper panel) vs. glucuronamide (lower panel) will determine the strength of the negative PD–EF relationship. For each community, PD is the standardized mean phylogenetic distance and EF is the sum of all taxa values
The concept of biodiversity has rapidly expanded beyond the number of species to include the variety of life at every hierarchical level, from genes to ecosystems [8]. Phylogenetic trees capture the hierarchical architecture of nature by clustering together those lineages with similar genotypes and, when traits are evolutionarily conserved, similar phenotypes and even ecological niches [9, 10]. Thus, the phylogenetic proximity between two lineages may satisfactorily describe their functional proximity. This is the main reason why phylogenetic diversity (PD) has become a powerful predictor of EF [11,12,13,14,15]. However, other authors have shown circumstances under which PD poorly predicts EFs [16,17,18]. The most obvious instance is when the relevant trait for EF is randomly dispersed across the phylogeny, and therefore phylogenetic distances are not accurate predictors of either ecological differences or functions [19]. Despite this possibility, significant evolutionary trait conservatism seems to prevail across the tree of life, that is, close relatives tend to have more similar trait values than distantly related organisms, as has been shown both for eukaryotes [10] and prokaryotes [9, 20]. From the observation that traits impacting EFs tend to be phylogenetically conserved springs the key idea that the historical conditions under which lineages evolved might have long lasting effects (evolutionary legacy effects) that may have direct influence on the performance of ecosystems [21].
To better understand how the evolutionary legacy of the different lineages shapes EFs, several methods propose to look at the relationship between PD and EF (PD–EF) across the evolutionary time depicted in the phylogenetic tree [22,23,24,25]. Ecological communities are usually composed by a mixture of anciently and recently divergent lineages and such evolutionary disparity has been shown to be reflected in functional trait diversity [23, 26, 27]. Interestingly, the evolutionary conservatism of functional traits in bacteria occurs at different phylogenetic depths. Although genetically complex traits like oxygenic photosynthesis and methane oxidation are conserved very deeply in the phylogeny, other traits like the ability to use simple carbon substrates show evolutionary conservatism at very shallow clade depths [20, 28]. Thus, the coexistence of lineages whose ancestors diversified very anciently is expected to impact the EF in a different way than recently divergent lineages owing to their evolutionary legacies [21, 25], as exemplified in Fig. 1. Differences in the evolutionary depths represented in the ecological communities can determine the sign of the PD–EF relationship, as follows.
Communities with high levels of EFs may result from the coexistence of functionally distinct lineages (niche differences) and/or from the presence of one or a few productive lineages that outcompete the rest (fitness differences) [15, 29]. Functional differences allow species to coexist through the segregation of their niches and, at the same time, to deliver different functions to the ecosystem. Thus, niche differences simultaneously increase both species diversity and EFs. Niche differences with an ancient origin, as for example the evolution of aerobic vs. anaerobic respiration (Fig. 1a), determine the occupancy of major niches [20]. Therefore, communities containing those functionally different lineages will have complementary functions (i.e., high EF values) performed by extremely distant lineages in the phylogenetic tree (i.e., high PD values) (left upper panel, Fig. 1b). On the other extreme, niche differences evolved recently, as for example the efficiency to grow on different carbon sources (Fig. 1a), will also lead to positive, but weaker PD–EF relationships because the phylogenetic distance between functionally distinct lineages will be necessarily smaller (left lower panel, Fig. 1b). Supporting this scenario, experimental evolution tests with bacteria show positive biodiversity–EF relationship in communities assembled with specialist lineages because of the enhanced niche complementarity [2]. That is, lineages may coexist by exploiting a narrower range of resources and, this situation can provide further functions to the ecosystem because more specialized lineages are expected to be more efficient on their own ground than generalists are (see [30] for a discussion of this effect called jack-of-all-trades is a master of none).
Communities composed by a few but very productive lineages may also show elevated levels of EFs. These productive lineages are very effective in the use of resources and tend to exclude other lineages. When such competitive superiority is phylogenetically conserved [31,32,33], the salient picture can be a negative PD–EF relationship owing to the low PD of the whole community (Fig. 1). Again, the strength of such relationship will depend on the evolutionary origin of the relevant trait. When the trait conferring fitness advantage is phylogenetically conserved at ancient times, a significant number of members of a single lineage may outcompete entire clades, leading to extremely negative PD–EF relationships (right upper panel, Fig. 1b). However, the relationship will be weaker if the trait is conserved at shallower clade depths (right lower panel, Fig. 1b). Evidence showing high EF levels at negative PD have been related to the high fitness of Proteobacteria under carbon-enriched conditions [14]. Numerous members of this ancient clade respond to high contents of soil organic carbon [34], performing very efficiently particular EFs while outcompeting multiple distantly related clades [35]. This process eventually leads to the overrepresentation of Proteobacteria and remarkably reduces the PD of the soil bacterial communities [36].
Ecosystems with large environmental and trait variability provide ideal scenarios to test the relationship between biodiversity and EF [37]. Biodiversity effects on EF in spatially heterogenous ecosystems may depend on both complementary effects at a local place and turnover in species composition and dominance across places at a broader scale [38]. Evidence in mono- vs multispecies plant experiments indicate that EF is simultaneously mediated by niche differences in plots with functionally diverse species and by fitness differences in plots with functionally similar species [39]. In patchy environments, like those shaped by ecological facilitation between plant species, the presence of stippled plant patches composed by a nurse plant and its facilitated species increases the heterogeneity of (organic and water) resources at the landscape level. We have previously shown that plant patchiness begets soil environmental and bacterial trait variability [40]. Although more bacterial ecological strategies coexist in the open spaces between plants, the milder conditions underneath plant patches increase the functional and phylogenetic similarities among coexisting bacteria by intensifying the competitive dominance of a single clade [40]. The high dominance of the productive Proteobacteria reduces the PD of the whole bacterial community resulting in high EF levels at very low PD [14]. Interestingly, however, the low levels of PD are accompanied with a richness increment [40]. Such an increment in bacterial richness is consistent with an effective niche partitioning occurring at shallow clade depths. Assuming niche conservatism, niche partitioning will promote coexistence, especially among distantly related species whose niches are less overlapped [25, 41]. In line with this mechanism, Goldfarb et al. [35] have suggested that specialization on C substrates, a trait conserved at shallow clade depths [20], may contribute to the PD of soil bacterial communities. A phylogenetic signature of this process would be a positive PD–EF relationship at shallow clade depths.
Based on these observations, we expect that in our model system fitness differences will mainly operate among ancient lineages, whereas niche differences will prevail at shallower phylogenetic scales. To test this hypothesis, we combine metrics of diversity that capture the phylogenetic composition of soil bacterial communities over the evolutionary time so as to better explain a set of microbially driven EFs. We detect phylogenetic-scale disparities in the soil microbial diversity–EF relationship, and discuss them in terms of the prevalence of anciently and recently divergent lineages.
Methods
Study site
The study site, which is located in Alacant (SE Spain; UTM 30 N 689062, 4238201), is described in detail elsewhere [42]. In this semi-arid Mediterranean region (240 mm mean annual rainfall, 20 °C mean annual temperature), we sampled gypsum soils covered with a patchy shrub steppe dominated by the gypsophyte legume Ononis tridentata L. that facilitates the establishment of other plant species [13]. In a total area of ca. 0.2 ha, we collected samples from two microsites: (1) plant patches, that is, an O. tridentata individual and the group of plants growing underneath its canopy, and (2) gaps, that is, the open space between multispecific plant patches, where individuals of gypsum-tolerant plant species grow scattered [13]. Out of a total 43 plant species recorded in the sampling area, 18 were detected growing both in the patches and gaps [13]. We collected 30 soil samples, 15 underneath plant patches and 15 in adjacent gaps. Each soil sample was a mixture of five surface soil samples (0–2 cm) that were randomly taken from the area of each patch or gap. Samples were transported to the laboratory on ice and sieved through a < 1 mm mesh. Further details are given in Goberna et al. [40].
Soil bacterial sequencing and phylogenetic reconstruction
Procedures used for soil DNA extraction, PCR amplification of a bacterial phylogenetic marker, targeted metagenomics, and subsequent sequence processing can be found in Goberna et al. [40]. Briefly, DNA extractions from 1 g soil were performed with the UltraCleanTM Soil DNA isolation kit (MO BIO Laboratories, Carlsbad, CA, USA). We PCR amplified the 16S ribosomal RNA (rRNA) gene, using the universal bacterial primers 8F [43] and 534R [44] including 454 sequencing adaptors and 8-bp barcodes. Purified tagged amplicons were quantified using the Quant-iTTM PicoGreen dsDNA Kit (Invitrogen) and pooled in equimolar amounts. Two samples that failed to amplify were discarded. Pyrosequencing was performed with the Roche 454 GS-FLX system using Titanium chemistry. After demultiplexing, trimming of sequencing adaptors and primers, we removed short and low-quality sequences and chimeras. We defined operational taxonomic units (OTUs) at a maximum identity level of 97% and obtained 3290 OTUs using the complete-linkage clustering method [45] that were aligned using the Infernal aligner [46] after discarding singletons. Sequences representative of each OTU were assigned to bacterial taxa using the Naïve Bayesian Classifier [47]. The relative abundance of each OTU in each plot was calculated based on the total number of reads in the same plot, and subsequently corrected by the estimated number of 16S rRNA gene copies [48]. Sequences were deposited in the European Nucleotide Archive (https://www.ebi.ac.uk/ena/data/view/PRJEB4887; [40]). In all soil bacterial communities, we detected the same 12 bacterial phyla, excluding those representing < 0.2% of the total community. Proteobacteria and Actinobacteria were the most abundant phyla both in patches and gaps, accounting for 60–80% of the community. Most phyla were equally abundant in both environments but Proteobacteria that were overrepresented underneath plant patches and Acidobacteria, Gemmatimonadetes, Cyanobacteria and Firmicutes in the open spaces [40].
We reconstructed the phylogenetic relationships among soil bacteria using RAxML 7.3.0 [49] with the GTRGAMMA substitution model after screening out hypervariable regions according to the Lane mask [50]. We constrained tree topology to match the phylum-level relationships of the mega-tree built from the Silva database [51], so as to reduce the large phylogenetic uncertainty resulting from the usage of short pyrosequencing reads. We selected a phylogenetic tree among the best of 1000 iterations, and used Archaeoglobus profundus the outgroup. We calibrated the phylogenetic tree so that branch lengths represent evolutionary time (in Myr) by using eight dated nodes (Supplementary Information S1) according to Sheridan et al. [52] and Marin et al. [53]. Calibration was run with the treePL software using the thorough and prime options [54]. This algorithm uses a penalized likelihood approach to estimate the divergence times and allows for different rates in different branches. The method optimizes the smoothing parameter that determines how much the rate differences among the tree are penalized based on cross-validations. We run calibrations with smoothing parameters ranging from 0.1 to 1000 and found 10 to be the optimum smoothing value [54].
Microbial EFs
We measured six parameters as indicators of microbially driven EFs, including:
(i) Pools of soil (1) microbial biomass C (MBC), which we used as a surrogate of total microbial biomass, and measured using the fumigation extraction procedure [55]; and (2) ATP, as an indicator of the total microbial activity, which we extracted with the procedure by Webster et al. [56] and quantified with the luciferine–luciferase assay using luminometry [57].
(ii) Rates of processes, including soil: (1) Organic matter decomposition, which we measured as the microbial basal respiration (BR) during 28-day aerobic incubation assays of root-free soils to quantify the mineralization of organic C into CO2 through gas chromatography [58]; (2) C cycling, which we inferred from β-glucosidase activity, by quantifying the enzymatic hydrolysis of β-glucoside bonds of carbohydrates. β-Glucosidases often catalyze the rate-limiting step in cellulose degradation and have been suggested as key regulators of the whole cellulolytic process [59]; (3) P cycling, which we estimated from alkaline phosphatase activity by measuring the enzymatic break down of phosphoric esters into phosphoric acid and alcohols [60]. This reaction is directly responsible for the replenishment of the soil pool of inorganic P; (4) N cycling, which we measured as the urease activity that is responsible for the enzymatic cleavage of urea into ammonia and CO2. All rates of enzymatic hydrolysis were measured colorimetrically, based on the production of p-nitrophenol for β-glucosidase and phosphatase [61, 62] and NH4+ for urease [63]. Details on all protocols can be found in Navarro-Cano et al. [13] and data in Supplementary Information S2.
PD and EFs at different phylogenetic depths
We tested the relationship between the bacterial PD and EFs across the evolutionary time using the three approaches that follow, and analyzing altogether our 28 plots encompassing a large variability in environmental and EF parameters.
We first applied two simple metrics traditionally used in community phylogenetics that recover the outcome of the interactions at different phylogenetic levels: (a) standardized mean phylogenetic distance (MPD), which recovers all the interactions between the taxa in the phylogeny, and therefore across all the phylogenetic depths of the tree and (b) standardized mean nearest taxon distance (MNTD) that only reflects interactions between pairs of closely related species, and therefore at the shallowest phylogenetic depth [64, 65]. MPD and MNTD were computed using the picante package for R [66] as follows: MPD = (MPDobs – MPDrand)/sdMPDrand, where MPDobs is the average of all pairwise phylogenetic distances between the taxa in the local community weighed by their abundances, MPDrand is the average of MPD calculated in n randomly constructed communities (i.e., after randomly shuffling all taxa labels) considering the regional pool of taxa (i.e., the sum of all taxa identified in all plots), and sdMPDrand is the standard deviation of MPDrand ([67]; Supplementary Information S3). MNTD was obtained as MPD but pairwise phylogenetic distances were calculated between the nearest taxa [67]. A factorial linear model was applied to test whether MPD and MNTD significantly explained each EF. To verify that the PD–EF relationship is mediated by high levels of environmental heterogeneity, we added in further statistical models the factor Microsite (i.e., Patch vs. Gap) that is the main source of environmental variation in our system. EF parameters were log transformed for all analyses to normalize the residuals of the model.
After inspecting whether EFs were explained with the metrics accounting for different phylogenetic depths, we tested whether these EFs correlated with the diversification metrics identifying anciently (ES1) and recently (ES2) divergent lineages in each plot (Supplementary Information S4). We used the ELDERness (Evolutionary Legacy of DivERsity) approach that compares the polynomial function representing the observed accumulation of lineages across the time depicted in the phylogeny against a straight line describing lineage accumulation under a constant diversification model [25]. The intersection point between the polynomial function and the straight line separates two areas. The first area (ES1), from time 0 to the time where intersection occurs, represents the accumulation of ancient lineages. The second area (ES2), from the intersection to the present time, indicates the accumulation of recent lineages. Real communities often show complex patterns of diversification by accumulating (i) both ancient and recent lineages, or (ii) lineages accumulating at intermediate evolutionary ages (see [25] for a detailed explanation of the ELDERness surfaces). The analyses were run with the ELDERness code for R [25]. A factorial linear model was applied to test whether absolute values of ES1 and ES2 significantly explained each EF. A second set of statistical models included the factor Microsite as above.
Finally, to detect the particular phylogenetic depths at which EFs were significantly correlated with taxa composition, we used the Beta-Diversity Through Time (BDTT) approach (Supplementary Information S5). This approach allows testing whether plots differing in their taxa composition also differ in their levels of particular EFs across several temporal slices in the phylogeny of bacteria [23]. We established 40 temporal slices of 100 million years each, from 0 to 4000 Myr, and calculated the compositional turnover (i.e., beta-diversity) between communities at each time. We subsequently correlated the beta-diversity matrix with the distance matrix of each of the six EFs measured. The Bray–Curtis index, which accounts for abundance data, was used to calculate beta-diversity and Euclidean distance to calculate EF distances. Matrix correlation was performed through Mantel tests and statistical significance computed after 1000 permutations of taxa names in the phylogeny. Analyses were run with the help of the BDTT code for R [23]. All analyses were performed using R 3.4.2 [68].
Results
The traditional metrics that captures the PD across all the phylogenetic depths of the tree (MPD) indicates a negative correlation between PD and EFs. That is, plots with low PD tend to have higher levels of microbial biomass, activity, organic matter decomposition and cycling of organic C, N and P compounds (MPD in Table 1). The other metrics, reflecting the PD at the shallowest clade depths (MNTD) failed to explain most EFs considered in this study (MNTD in Table 1). Only for N cycling, did MNTD show positive significant effects (Table 1).
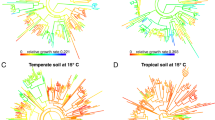
To explore the PD–EF relationship based on ancient and recently divergent lineages, we calculated the ELDERness surfaces. In all soil bacterial communities, we detected a complex pattern of diversification, with positive ES1 (mean = 81.4 range = [0.3, 368.9]) and negative ES2 values (mean = −2342.4, range = [−3081, −582.6]). This is the result of high rates of lineage accumulation both at the beginning and end of the evolutionary time spanned by the bacterial phylogenetic tree separated by a plateau with low levels of lineage accumulation (see an example of a lineage accumulation plot in Fig. 2). This branching pattern was common to all our study plots, but soils in open spaces tended to accumulate more anciently divergent lineages than soils underneath plant patches as shown by the larger ES1 values in gaps (t13.2 = 3.47, p < 0.05; Fig. 3a). On the contrary, gaps showed a significantly lower accumulation of recent lineages with their smaller ES2 values (t17.1 = 4.925, p < 0.05; Fig. 3b).
Accumulation of lineages in a study plot along the evolutionary time of the bacterial phylogeny. Each point reflects the divergence time of a given node in the phylogeny (x axis) vs. the number of lineages subtending that node (y axis). The solid line represents the expected accumulation of lineages under a constant speciation model. ES1 and ES2, respectively, represent the areas between the curve of accumulation of anciently and recently divergent lineages and the constant speciation line
EFs were significantly explained by the interaction between the abundance of anciently (ES1) and recently divergent lineages (ES2) (see significant ES1 × ES2 interaction in Table 2). The analysis of the statistical interaction showed that higher levels of all EFs were systematically detected in plots with high accumulation of recent lineages (Supplementary Information S6). However, such an increase was modulated by the accumulation of ancient lineages in a distinct manner for the EFs studied. The largest levels of microbial biomass and activity, as well as the fastest rates of decomposition and P cycling were found in plots enriched in recent lineages and having a low accumulation of ancient lineages. The highest rates of C cycling were achieved at high levels of recent lineages irrespective of ancient lineages, whereas N cycling increased in plots combining high accumulation of both recent and ancient lineages (Supplementary Information S6).
We verified that the environmental heterogeneity underlies the strength of the PD–EF relationship by incorporating to our statistical models the effect of the Microsite (patch vs. gap). This factor was highly significant in all models and overrode the power of simple metrics, such as MPD and MNTD, to predict EF levels (Supplementary Information S7). However, ES1 and/or ES2 were significant (or marginally significant) EF predictors even after accounting for the effect of the microsite for microbial activity, decomposition and C cycling.
To detect the particular phylogenetic depths at which EFs were significantly correlated with taxa composition, we calculated the β-diversity at different time slices across the phylogenetic tree. Significant correlations between β-diversity and EF distances indicate that communities harbouring similar bacterial lineages tend to display similar levels of EFs at all the phylogenetic depths between 0 and 2900 Myr (Fig. 4a and Supplementary Information S8). The highest correlations between EF and lineage composition were obtained for communities sharing either anciently divergent (2100–2900 Myr) or recently divergent lineages (i.e., 200–600 Myr), whose phylogenetic composition varied remarkably (Fig. 4b). The anciently divergent lineages at the phylogenetic depth of 2500 Myr corresponded to eight extant phyla and the most recent common ancestors of the clades Chloroflexi-OD1-TM7 and Planctomycetes–Verrucomicrobia–Gemmatimonadetes–Acidobacteria–Nitrospira (Fig. 4b). At 600 Myr, the total 16 bacterial phyla present in our study system were represented and had experienced a remarkable diversification (Fig. 4b).
a Phylogenetic-scale disparities in the relationship between soil bacterial diversity and ecosystem functions (correlation between β-diversity and decomposition distance) at different temporal slices. Significant and nonsignificant correlations are indicated with orange and grey circles, respectively. The time points at which the highest correlations are obtained are shown with horizontal dashed lines and projected to the phylogenetic tree. b Bacterial phylogenetic tree depicting the evolutionary relationships between 16 phyla present in the study system. At 2500 Myr, coloured circles indicate eight extant phyla and the most recent common ancestors of the clades Chloroflexi-OD1-TM7 and Planctomycetes–Verrucomicrobia–Gemmatimonadetes–Acidobacteria–Nitrospira
Discussion
Soil is an exceptionally heterogeneous habitat where a huge number of bacteria compete with their neighbours for space and resources [69]. Survival of bacteria in hyperdiverse communities raises the same problem posed by the “paradox of the plankton”. As Hutchinson [70] early noted, the coexistence of a huge number of competing species is possible under spatio-temporal heterogeneity of resources and conditions, positive interactions or differential predation. Ultimately, all these factors will determine species coexistence by altering the balance between forces equalizing niche differences and forces reducing fitness differences [71]. Each type of force leaves a different phylogenetic signature in the community structure [72] that we have used to explain the relationship between the PD and key EFs performed by soil bacteria.
The rationale behind the use of the PD as a proxy for EF initially stemmed from the simple idea that distantly related species have had more time to diverge functionally and, therefore, to provide a wider variety of EFs [11, 12, 15]. In this study, we found that this relationship is much more complex and requires the correct understanding of the outcome of ecological interactions occurring at different phylogenetic depths. The evolutionary legacy represented in soil bacterial communities leaves signatures of functional differences across the phylogenetic tree that can be used in an integrative way to understand the mechanisms behind EFs. More specifically, we detected that important EFs related to microbial productivity, decomposition and nutrient cycling required the use of metrics sensitive to both processes occurring anciently and recently in the history of bacterial evolution.
MPD, a traditional metrics that is sensitive to all the phylogenetic depths [73], indicates that bacterial PD is negatively correlated with the levels of all our estimated EFs. Supporting this interpretation, we also found, by using a beta-diversity through time approach, that communities resembling in their bacterial composition display similar levels of EFs across all the phylogenetic depths throughout 2900 Myr. This pattern held across EF, which were positively correlated (Pearson r > 0.7, p < 0.05). The negative PD–EF relationship detected with MPD could be broadly interpreted as fitness differences between productive and unproductive clades reducing PD and increasing ecosystem productivity [14, 74]. Although MPD cannot discern at which phylogenetic depths these processes are occurring, the maximum correlations between beta-diversity and EF were detected at ancient (2100–2900 Myr) and recent (200–600 Myr) times, suggesting a temporal dimension in the PD–EF relationship. We tried to focus at shallow phylogenetic depths by using the MNTD, which exclusively attends to phylogenetic distances between pairs of closest relatives. MNTD was, however, not able to detect PD–EF relationships for most EFs. This result probably reflects the fact that working at the tips of the phylogeny introduces a lot of uncertainty derived from the low phylogenetic resolution we currently have at the OTU level. Only for N cycling, did we detect a positive correlation between MNTD and EF, which could be theoretically assigned to niche differences occurring at shallow phylogenetic depths. Simple PD metrics such as MPD and MNTD have been shown to be worse predictors of EFs than more complex metrics that capture the patterns of diversification along the whole phylogeny [25]. This is particularly the case under complex branching scenarios, such as those occurring in our soil bacterial communities. By analyzing the branching patterns depicted in the phylogenetic tree through time, we could delimit how ancient (ES1) and recent lineages (ES2) contributed to each EF as follows.
We detected a general pattern indicating that the levels of all the EFs increased with the accumulation of recently divergent lineages in the community. The contribution of recent lineages to a positive PD–EF relationship agrees with the scenario in which niche differences mediated by traits conserved at shallow phylogenetic depths allow the coexistence of species (left lower panel, Fig. 1b). This process is coherent with the response of bacterial lineages to different organic sources that is phylogenetically conserved at shallow phylogenetic depths [20, 33]. Such differences in resource consumption arise not only from the various chemical structure of the array of organic substrates present in soil (e.g., organic P compounds) but also from the differential catalytic efficiency of enzymes that belong to the same family (e.g., phosphatases) depending on their biological origin [75]. The more effective niche filling with differentiated recent lineages would ultimately explain the correlation between high PD and EFs [25]. Such a consistent pattern was finely tuned by the accumulation of ancient lineages in the community, but in various ways for different EFs. Microbial productivity, in terms of biomass and activity, and the rates of decomposition were lowered in the presence of ancient lineages, what would be coherent with the scenario where fitness differences among deeply branching clades drive community assembly (right upper scenario, Fig. 1b). In particular, the presence of ancient lineages, like Proteobacteria and Actinobacteria, which are superior competitors that respond to experimental carbon inputs, may produce the competitive exclusion of less productive clades such as Acidobacteria, Planctomycetes or Verrucomicrobia [34, 35]. This process would simultaneously explain low PD and high productivity [14]. Similar responses of main bacterial phyla as those described after carbon addition to soil have been described following the amendment with other nutrients [76, 77]. However, we detected that EFs associated with the hydrolysis of organic C, N or P compounds showed idiosyncratic patterns regarding the contribution of ancient lineages. Further research on the complexity and evolutionary conservatism of the traits involved in bacterial rates of nutrient cycling is needed to assign these patterns to fitness or niche differences at the deep phylogenetic level.
Altogether, our results indicate that EF can be both increased or decreased with PD depending on the phylogenetic scale we attend. At first glance, these processes seem mutually exclusive but this situation has been already predicted by theoretical models. Indeed, theory predicts that even when EFs are explained by a few productive, competitive lineages, many other species are needed to keep high levels of EF if competitive hierarchy changes with time or space [38]. According to this prediction, the high spatial heterogeneity of resources and conditions in our study system mediates the PD–EF relationship by allowing different taxonomic groups to succeed. For example, some classes within the phylum Proteobacteria (Gammaproteobacteria and Betaproteobacteria) or families within Actinobacteria (Micrococcaceae) have been shown to outgrow and reduce the amount of available resources for their competitors [33, 35]. At the community level, the phyla Proteobacteria and Actinobacteria would be the dominant lineages outcompeting the rest of clades in the community. However, the competitive superiority of these clades would not reduce the species richness in the community because spatial variability in the available resources would change the competitive hierarchy within the dominant phyla promoting coexistence and, ultimately, richness [78].
Both fitness differences leading to competitive superiority at deep and niche differences at recent phylogenetic depths are shaped by evolution, making more complex the relationship between phylogeny and coexistence, and therefore the PD–EF relationship [79]. We admit that our interpretation of the mechanisms underlying the patterns observed remains speculative and assumes niche conservatism [25]. Assembling experimental communities with anciently and recently divergent lineages where both the outcome of competitive interactions and the levels of EFs were registered would help supporting or refuting the putative mechanisms we have described. Elusive relationships require complex models and therefore the inclusion of several phylogenetic metrics uncovering ancient and recent evolutionary histories will help improving our predictions on how the evolutionary legacy of soil bacterial communities explain EFs.
References
Bardgett RD, Van der Putten WH. Belowground diversity and ecosystem functioning. Nature. 2014;515:505–511.
Gravel D, Bell T, Barbera C, Bouvier T, Pommeir T, Venail P, et al. Experimental niche evolution alters the strength of the diversity-productivity relationship. Nature. 2011;469:89–92.
Midgley GF. Biodiversity and ecosystem function. Science. 2012;335:174–175.
Van der Heijden MGA, Bardgett RD, van Straalen NM. The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett. 2008;11:296–310.
Wardle DA, Bardgett RD, Klironomos JN, Setälä H, Van der Putten WH, Wall DH. Ecological linkages between aboveground and belowground biota. Science. 2004;304:1629–1633.
Hooper DU, Chapin ES III, Ewel JJ, Hector A, Inchausti P, Lavorel S, et al. Effects of biodiversity on ecosystem functioning: a consensus of current knowledge. Ecol Monogr. 2005;75:3–35.
Díaz S, Purvis A, Cornelissen JHC, Mace GM, Donoghue MJ, Ewers RM, et al. Functional traits, the phylogeny of function, and ecosystem service vulnerability. Ecol Evol. 2013;3(9):2958–2975.
Wilson EO. Biodiversity. Washington D.C., USA: National Academy Press; 1988.
Goberna M, Verdú M. Predicting microbial traits with phylogenies. ISME J. 2016;10:959–967.
Blomberg SP, Garland T Jr, Ives AR. Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution. 2003;57:717–745.
Flynn DF, Mirotchnick N, Jain M, Palmer MI, Naeem S. Functional and phylogenetic diversity as predictors of biodiversity–ecosystem-function relationships. Ecology. 2011;92:1573–1581.
Cadotte MW. Experimental evidence that evolutionarily diverse assemblages result in higher productivity. Proc Natl Acad Sci USA. 2013;110:8996–9000.
Navarro-Cano JA, Goberna M, Valiente-Banuet A, Montesinos-Navarro A, García C, Verdú M. Plant phylodiversity enhances soil microbial productivity in facilitation-driven communities. Oecologia. 2014;174:909–920.
Pérez-Valera E, Goberna M, Verdú M. Phylogenetic structure of soil bacterial communities predicts ecosystem functioning. FEMS Microbiol Ecol. 2015;91:fiv031.
Srivastava DS, Cadotte MW, MacDonald AAM, Marushia RG, Mirotchnick N. Phylogenetic diversity and the functioning of ecosystems. Ecol Lett. 2012;15:637–648.
Gravel D, Bell T, Barbera C, Combe M, Pommier T, Mouquet N. Phylogenetic constraints on ecosystem functioning. Nat Comm. 2012;3:1117.
Cadotte MW, Davies JT, Peres-Neto PR. Why phylogenies do not always predict ecological differences. Ecol Monogr. 2017;87:535–551.
Venail P, Gross K, Oakley TH, Narwani A, Allan E, Flombaum P, et al. Species richness, but not phylogenetic diversity, influences community biomass production and temporal stability in a re-examination of 16 grassland biodiversity studies. Funct Ecol. 2015;29:615–626.
de Bello F, Šmilauer P, Diniz-Filho JAF, Carmona CP, Lososová Z, Herben T, et al. Decoupling phylogenetic and functional diversity to reveal hidden signals in community assembly. Methods Ecol Evol. 2017;8:1200–1211.
Martiny AC, Treseder K, Pusch G. Phylogenetic conservatism of functional traits in microorganisms. ISME J. 2013;7:830–838.
Cavender-Bares J, Ackerly DD, Hobbie SE, Townsend PA. Evolutionary legacy effects on ecosystems: biogeographic origins, plant traits, and implications for management in the Era of global change. Annu Rev Ecol Evol Syst. 2016;47:433–462.
Davies TJ, Urban M, Rayfield B, Cadotte MW, Peres-Neto PR. Deconstructing the relationships between phylogenetic diversity and ecology: a case study on ecosystem functioning. Ecology. 2016;97:2212–2222.
Groussin M, Mazel F, Sanders J, Smillie C, Lavergne S, Thuiller W, et al. Unraveling the processes shaping mammalian gut microbiomes over evolutionary time. Nat Comm. 2017;8:14319.
Mazel F, Davies TJ, Gallien L, Renaud J, Groussin M, Münkemüller T, Thuiller W. Influence of tree shape and evolutionary time-scale on phylogenetic diversity metrics. Ecography. 2015;39:913–920.
Yguel B, Jactel H, Pearse IS, Moen D, Winter M, Hortal J, et al. The evolutionary legacy of diversification predicts ecosystem function. Am Nat. 2016;188:398–410.
Herrera CM. Historical effects and sorting processes as explanations for contemporary ecological patterns: character syndromes in Mediterranean woody plants. Am Nat. 1992;140:421–446.
Valiente-Banuet A, Vital A, Verdú M, Callaway R. Modern Quaternary plant lineages promote diversity through facilitation of ancient Tertiary lineages. Proc Natl Acad Sci USA. 2006;103:16812–16817.
Martiny JBH, Jones SE, Lennon JT, Martiny AC. Microbiomes in light of traits: a phylogenetic perspective. Science. 2015;350:aac9323-1–8.
Carroll IT, Cardinale BJ, Nisbet RM. Niche and fitness differences relate the maintenance of diversity to ecosystem function. Ecology. 2011;92:1157–1165.
Pianka ER. Evolutionary ecology. 7th edn – ebook. 2011.
Goberna M, Navarro-Cano JA, Verdú M. Opposing phylogenetic diversity gradients of plant and soil bacterial communities. Proc R Soc London B. 2016;283. https://doi.org/10.1098/rspb.2015.30031825.
Mayfield MM, Levine JM. Opposing effects of competitive exclusion on the phylogenetic structure of the communities. Ecol Lett. 2010;13:1085–1093.
Morrissey EM, Mau RL, Schwartz E, Caporaso JG, Dijkstra P, van Gestel N, et al. Phylogenetic organization of bacterial activity. ISME J. 2016;10:2336–2340.
Fierer N, Bradford MA, Jackson RB. Toward an ecological classification of soil bacteria. Ecology. 2007;88:1354–1364.
Goldfarb KC, Karaoz U, Hanson CA, Santee CA, Bradford MA, Treseder KK, et al. Differential growth responses of soil bacterial taxa to carbon substrates of varying chemical recalcitrance. Front Micro. 2011;2:1–10.
Goberna M, García C, Verdú M. A role for biotic filtering in driving phylogenetic clustering in soil bacterial communities. Glob Ecol Biogeog. 2014;23:1346–1355.
Hodapp D, Hillebrand H, Blasius B, Ryabov AB. Environmental and trait variability constrain community structure and the biodiversity-productivity relationship. Ecology. 2016;97(6):1463–1474.
Isbell F, Cowles J, Dee LE, Loreau M, Reich PB, Gonzalez A, et al. Quantifying effects of biodiversity on ecosystem functioning across times and places. Ecol Lett. 2018;21(6):763–778.
Cadotte MW. Functional traits explain ecosystem function through opposing mechanisms. Ecol Lett. 2017;20:989–996.
Goberna M, Navarro-Cano JA, Valiente-Banuet A, García C, Verdú M. Abiotic stress tolerance and competition related traits underlie phylogenetic clustering in soil bacterial communities. Ecol Lett. 2014;17:1191–1201.
Wiens JJ, Ackerly DD, Allen AP, Anacker BL, Buckley LB, Cornell HV, et al. Niche conservatism as an emerging principle in ecology and conservation biology. Ecol Lett. 2010;13:1310–1324.
Goberna M, Pascual JA, García C, Sánchez J. Do plant clumps constitute microbial hotspots in semi-arid Mediterranean patchy landscapes? Soil Biol Biochem. 2007;39:1047–1054.
Turner S, Pryer KM, Miao VPW, Palmer JD. Investigating deep phylogenetic relationships among Cyanobacteria and plastids by small subunit rRNA sequence analysis. J Eukaryot Microbiol. 1999;46:327–338.
Muyzer G, De Waal EC, Uitterlinden AG. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol. 1993;59:695–700.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, et al. The ribosomal database project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res. 2009;37:D141–145.
Nawrocki EP, Eddy SR. Query-dependent banding (QDB) for faster RNA similarity searches. PLoS Comput Biol. 2007;3:e56.
Wang Q, Garrity GM, Tiedje JM, Cole JR. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73:5261–5267.
Kembel SW, Wu M, Eisen JA, Green JL. Incorporating 16S gene copy number information improves estimates of microbial diversity and abundance. PLoS Comp Biol. 2012;8:e1002743.
Stamatakis A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics. 2006;22:2688–2690.
Lane DJ. 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M, editors. Nucleic acid techniques in bacterial systematics. New York, NY: Wiley; 1991. p. 115–175.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucl Acids Res. 2013;41:D590–D596. (D1)
Sheridan PP, Freeman KH, Brenchley JE. Estimated minimal divergence times of the major bacterial and archaeal phyla. Geomicrobiol J. 2003;20:1–14.
Marin J, Battistuzzi FU, Brown AC, Hedges SB. The timetree of prokaryotes: new insights into their evolution and speciation. Mol Biol Evol. 2017;34:437–446.
Smith SA, O’Meara BC. treePL: divergence time estimation using penalized likelihood for large phylogenies. Bioinformatics. 2012;28(20):2689–2690.
Vance ED, Brookes PC, Jenkinson DS. An extraction method for measuring soil microbial biomass C. Soil Biol Biochem. 1987;19:703–707.
Webster JJ, Hampton GJ, Leach FR. ATP in soil: a new extractant and extraction procedure. Soil Biol Biochem. 1984;16:335–342.
Alef K, Nannipieri P. Methods in applied soil microbiology and biochemistry. London: Academic Press; 1995.
Nannipieri P, Grego S, Ceccanti B. Ecological significance of the biological activity in soil. In: Bollag JM, Stotzky G, editors. Soil biochemistry. New York, NY: Marcel Dekker; 1990. p. 293–355.
Yeoman CJ, Han Y, Dodd D, Schroeder CM, Mackie RI, Cann IKO. Thermostable enzymes as biocatalysts in the biofuel industry. Adv Appl Microbiol. 2010;70:1–55. https://doi.org/10.1016/S0065-2164(10)70001-0. e-pub
Tabatabai MA. Soil enzymes. In: Bottomley PS, Angle JS, Weaver RW, editors. Methods of soil analysis, Part 2. Microbiological and biochemical properties. Madison: SSSA Book Series, no 5. Soil Science Society of America; 1994. p. 775–833.
Eivazi F, Tabatabai MA. Glucosidases and galactosidases in soils. Soil Biol Biochem. 1988;20:601–606.
Tabatabai MA, Bremner JM. Use of p-nitrophenyl phosphate for assay of soil phosphatase activity. Soil Biol Biochem. 1969;1:301–307.
Kandeler E, Gerber H. Short-term assay of soil urease activity using colorimetric determination of ammonium. Biol Fert Soils. 1988;6:68–72.
Kraft NJB, Cornwell WK, Webb CO, Ackerly DD. Trait evolution, community assembly, and the phylogenetic structure of ecological communities. Am Nat. 2007;170:271–283.
Tucker CM, Cadotte MW, Carvalho SB, Davies TJ, Ferrier S, Fritz SA, et al. A guide to phylogenetic metrics for conservation, community ecology and macroecology. Biol Rev. 2017;92:698–715.
Kembel SW, Cowan PD, Helmus MR, Cornwell WK, Morlon H, Ackerly DD, et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics. 2010;26:1463–1464.
Webb CO, Ackerly DD, McPeek MA, Donoghue MJ. Phylogenies and community ecology. Annu Rev Ecol Syst. 2002;33:475–505.
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. 2017. http://www.R-project.org/.
Schloss PD, Handelsman J. Towards a census of bacteria in soil. PLoS Comput Biol. 2006;2(7):e92 https://doi.org/10.1371/journal.pcbi.0020092.
Hutchinson GE. The paradox of the plankton. Am Nat. 1961;95:137–145.
Chesson P. Mechanisms of maintenance of species diversity. Annu Rev Ecol Syst. 2000;31:343–366.
HilleRisLambers J, Adler PB, Harpole WS, Levine MS, Mayfield MM. Rethinking community assembly through the lens of coexistence theory. Annu Rev Ecol Evol Syst. 2012;43:227–248.
Qian H, Chen S, Zhang JL. Disentangling environmental and spatial effects on phylogenetic structure of angiosperm tree communities in China. Sci Rep. 2017; 7. https://doi.org/10.1038/s41598-017-04679-5.
Turnbull LA, Isbell F, Purves DW, Loreau M, Hector A. Understanding the value of plant diversity for ecosystem functioning through niche theory. Proc R Soc B. 2016;283:20160536.
Tazisong IA, Senwo ZN, He Z. Phosphatase hydrolysis of organic phosphorus compounds. Adv Enz Res. 2015;3:39–51.
Leff JW, Jones SE, Prober SM, Barberán A, Borer ET, Firn JL, et al. Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe. Proc Natl Acad USA. 2015;112:10967–10972.
Zhou J, Jiang X, Wei D, Zhao B, Ma M, Chen S, et al. Consistent effects of nitrogen fertilization on soil bacterial communities in black soils for two crop seasons in China. Sci Rep. 2017;7:3267.
Gallien L. Intransitive competition and its effects on community functional diversity. Oikos. 2017;126:615–623.
Godoy O, Kraft NJB, Levine JM. Phylogenetic relatedness and the determinants of competitive outcomes. Ecol Lett. 2014;17:836–844.
Acknowledgements
Thanks to B Yguel, J Hortal and E Pérez-Valera for helping with the ELDERness models. Financial support was provided by the Spanish Ministry of Economy and Competitiveness (CGL2014-58333-P; CGL2016-81706-REDT; CGL2017-89751-R) and the Generalitat Valenciana (SEJI/2017/030). MG acknowledges support by the Ramón y Cajal Programme of the Spanish Ministry of Economy and Competitiveness.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Goberna, M., Verdú, M. Phylogenetic-scale disparities in the soil microbial diversity–ecosystem functioning relationship. ISME J 12, 2152–2162 (2018). https://doi.org/10.1038/s41396-018-0162-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0162-5
This article is cited by
-
Reconsidering functional redundancy in biodiversity research
npj Biodiversity (2023)
-
Transitivity and intransitivity in soil bacterial networks
The ISME Journal (2023)
-
Microbial assembly, interaction, functioning, activity and diversification: a review derived from community compositional data
Marine Life Science & Technology (2019)