Abstract
Marine microbes have tremendous diversity, but a fundamental question remains unanswered: why are there so many microbial species in the sea? The idea of functional redundancy for microbial communities has long been assumed, so that the high level of richness is often explained by the presence of different taxa that are able to conduct the exact same set of metabolic processes and that can readily replace each other. Here, we refute the hypothesis of functional redundancy for marine microbial communities by showing that a shift in the community composition altered the overall functional attributes of communities across different temporal and spatial scales. Our metagenomic monitoring of a coastal northwestern Mediterranean site also revealed that diverse microbial communities harbor a high diversity of potential proteins. Working with all information given by the metagenomes (all reads) rather than relying only on known genes (annotated orthologous genes) was essential for revealing the similarity between taxonomic and functional community compositions. Our finding does not exclude the possibility for a partial redundancy where organisms that share some specific function can coexist when they differ in other ecological requirements. It demonstrates, however, that marine microbial diversity reflects a tremendous diversity of microbial metabolism and highlights the genetic potential yet to be discovered in an ocean of microbes.
Similar content being viewed by others
Introduction
Marine microorganisms play key roles in major biogeochemical processes [1], and the rapid development of sequencing tools has made it possible to uncover the extent of their community diversity [2]. The patterns of diversity correlate with the communities’ surrounding environment on different spatial and temporal scales [3], including the season [4], depth [5], or latitude [6]. As the ecological significance of microbial diversity for ecosystem function starts to be recognized [7], one of the fundamental questions that remains is why there are so many microbial species in the sea [2]. The observation of tremendous diversity was at the heart of the paradox of the plankton formulated by Hutchinson more than 50 years ago [8], which can be transposed to marine microbes as how an apparently limited range of resources can support an unexpectedly large number of microbial species. This high diversity was hypothesis to result from the presence in the environment of different taxa that are able to conduct the same set of metabolic processes [9, 10]. The maintenance of taxonomically diverse communities could thus be justified by the notion of functional redundancy. The idea of functional redundancy has long been assumed in microbial ecology and is often used in models, including those used to project responses to climate change. The assumption in so-called “black box” approaches is that a change in microbial community composition will not have consequences for microbial-mediated processes [9].
The paradigm of redundancy does not always hold in natural microbial communities, as shown recently in soil ecosystems [11]. In the sea, the predictable reoccurrence of microbial communities also suggests a low degree of strict functional redundancy [7, 12]. However, these marine studies were restricted to a taxonomic approach that did not take into account the entire functional potential of communities. Inversely, the use of genes annotated against databases [13] or predicted functional profiles [14] revealed high functional redundancy for marine microbes on a global scale. These studies, which rely on annotations obtained from cultured organisms, suggest that metabolic pathways have spread across taxa throughout evolution so that different microbial species conduct the same set of enzymatic reactions. Such a finding implies that different species can have similar niches and thus compete against each other. It does not resolve the “paradox of the marine microbes” because according to the competitive exclusion principle, species competing for the same resource cannot coexist, and diversity should decline if many species have the same functional traits. Thus, whether the enormous marine microbial diversity is characterized by a high level of functional redundancy is still unknown. Functional data from the field have seldom been used to validate or refute the hypothesis, and it has never been tested within a well-defined conceptual frame.
To be able to test the hypothesis of functional redundancy, the concept itself should be clearly defined. Functional redundancy can be defined as the coexistence of organisms that share the exact same set of functions and that can readily replace each other [3], (let’s call it “strict redundancy”). Alternatively, functional redundancy can be defined as the coexistence of organisms that share some specific function (e.g., two ammonia oxidizers) but may nevertheless differ in additional functions or other ecological requirements such as temperature preference (let’s call it “partial redundancy”). This is a much less stringent definition and often used in the literature on ecosystem stability [15]. In the present study, we aim at testing if there is a strict functional redundancy within marine microbial communities. Another important factor that should be taken in to account when testing functional redundancy is that communities have to be studied under common environmental conditions [9, 16], which may be difficult in nature. We dealt with this challenge by using the Banyuls Bay microbial observatory, which allowed us to repeatedly sample communities year after year under similar environmental conditions [4]. Our temporal study was extended to include a spatial factor and to test whether possible changes in community composition across regions, within the common environment of the Mediterranean Sea, alter the functional attributes of the communities. In the case of a strict redundancy, taxonomically different organisms that have the same set of functions could replace each other in the environment. We, therefore, postulate that a shift in community composition that alters community functions refutes the hypothesis of functional redundancy.
Materials and methods
Sampling
Surface seawater (3 m) was collected monthly from January 2012 to February 2015 (40 samples) by using a 10-L Niskin bottle at the SOLA station (42°31′N, 03°11′E) in the Bay of Banyuls sur Mer (France) in the northwestern Mediterranean. The frequency of the sampling varied slightly because of ship availability and occasional poor weather conditions. The water was kept in high-density polyethylene carboys in the dark until being processed in the laboratory (within 1.5 h). A volume of 5 L was prefiltered through 3-μm pore-size polycarbonate filters (Millipore, Billerica, MA, USA), and the microbial biomass was collected on 0.22-μm pore-size GV Sterivex cartridges (Millipore) and stored at −80 °C until nucleic acid extraction.
In-situ temperature and salinity were obtained using a Seabird CTD SBE9/11. Concentrations of nitrate, nitrite, phosphate, and silicate were determined with a Skalar auto-analyzer following a previously described protocol [17]. Dissolved ammonium was determined by spectrophotometry at 630 nm following conversion to indophenol via a monochloroamine intermediate [18]. Chlorophyll a concentrations were measured from 1 L of seawater collected on a GF/F filter at low pressure ( < 0.2 bar) as in ref. [7]. The physicochemical parameters (Extended Data Table 1) were provided by the Service d’Observation en Milieu Littoral (SOMLIT).
DNA extraction and metagenome sequencing
The nucleic acid extraction method followed the protocol described by Hugoni et al. [4] and consisted of cell lysis with freshly prepared lysozyme solution (20 mg/mL) applied directly to Sterivex cartridges, a second incubation with proteinase K (20 mg/mL), followed by extraction using the AllPrep DNA/RNA kit (Qiagen, Hilden, Germany).
The Nextera XT DNA Sample Preparation Kit (Illumina, San Diego, CA, USA) was used to fragment DNA and ligate adapters. The DNA quality was checked with the Agilent High Sensitivity kit (Agilent Technologies, Santa Clara, CA, USA). Samples were sequenced on eight lanes of a HiSeq 2500 “High-Output” paired-end run (2 × 100 bp) (Illumina). Sequencing produced a total of 2,984,444,036 reads (Table S2). Raw sequences were archived in the EBI repository under accession number PRJEB26919.
Read filtering and metagenome assembly
Raw paired-end Illumina sequences were preprocessed by removing Nextera adapters with the bbduck program from the BBTools package (12.10.2015 release) (http://jgi.doe.gov/data-and-tools/bbtools/). Reads were then trimmed using Trimmomatic (v. 0.33, [19]) based on their quality and length (LEADING:28 TRAILING:28 SLIDINGWINDOW:4:15 MINLEN:30), generating a read length of ca. 85 bp. A total of 34 to 112 million reads per sample remained after filtering (Extended Data Table 2). For each metagenome, high-quality reads were individually assembled with IDBA-UD [20] with the default iterative k-mer assembly with the k-mer length increasing from 20 to 100 bp in steps of 20, the pre-correction option, and with both pair-end reads (-r entry) and single-end reads (--long entry).
Comparison of the metagenomic data
The methods listed below are represented in Supplementary Fig. S6.
Comparison without a priori knowledge
To define the functional attributes of the microbial communities, we targeted the entire set of reads (method 1) and the entire set of predicted proteins (method 2) for comparison purposes. We avoided over-simplifying the complex microbial communities by describing the full community-aggregated functional attributes in an approach similar to the concept of community-aggregated traits [21].
Read-based approach (method 1)
The high-quality reads of the 40 metagenomes were compared to assess the pairwise similarity with the Commet software [22]. The method allows an all-against-all comparison of the non-assembled reads based on shared k-mers. To verify the results, a similar analysis was conducted with the MetaFast software [23]. The results given by these two tools were similar (Supplementary Fig. S7).
Predicted genes approach (method 2)
Gene prediction for contigs ≥ 1 kb was performed using MetaGeneAnnotator [24] and generated a total of 6.4 million gene-coding sequences. A catalog of genes was then built by clustering the predicted gene-coding sequences using CD-HIT (v. 4.6, [25] (parameters: -g 1, -c 0.95, -aS 0.90) at 95% identity as done earlier [13]. Sequences shorter than 100 bp were discarded. The resulting catalog contained 1,568,213 non-redundant predicted genes for the SOLA site.
To build an abundance matrix of gene-coding sequences, a total of 30,750,000 high-quality reads were randomly selected (without replacement) from each metagenome. These reads were mapped to our predicted SOLA gene catalog using the SOAPaligner [26] with options -M 4 (find best hits), -l 30 (seed length), -r 1 (random assignment of multiple hits), and -v 5 (maximum number of mismatches). Mapped reads were filtered using a minimum mapping quality of 10 and were counted to form an abundance matrix. The abundance matrix was then normalized to the gene length as in ref. [13].
Comparison with a priori knowledge
Annotation-based analysis (method 3)
All predicted translated genes were compared to the KEGG database [27] using UBLAST [28] with an e-value threshold of 1e-3 and a percentage of identity of 60%. Annotation-based abundance tables were then constructed by keeping only the annotated genes from the full gene-coding matrix. Out of the 1,568,213 predicted genes detected at the SOLA site only 283,094 could be annotated against KEGG.
16S rRNA based approaches
Taxonomic annotation (method 4)
The 16S rRNA gene sequences were identified by comparing all HQ reads to the SILVA (v.123, [29]) 16S rRNA database with BLASTn (identity ≥ 90% and length > 80 bp). A total of 343,234 16S rRNA sequences were detected in the 40 samples. An OTU table was built by clustering reads at a 97% sequence similarity against the SILVA sequence collection. For further analysis, all samples were resampled down to an equal number of sequences.
Predictive approaches: PICRUST and FAPROTAX (method 5)
The 16S rRNA contigs were identified with BLASTn (e-value < 1e-5, identity ≥ 97%) against the SILVA 16S rRNA database (v. 123, [29]). A total of 1563 contigs with a length > 300 bp were detected in the 40 samples. For each sample, the randomly drawn high-quality reads (see above) were mapped to the 16S rRNA contigs with BWA [30] (mem algorithm and minimum mapping quality of 10). Mapped reads were counted to form an OTU table. The community composition originating from the abundance table obtained from contigs was similar to the one obtained directly with all reads (Supplementary Fig. S8).
The reference sequences of the contig-based 16S rRNA OTU table were taxonomically affiliated against the greengenes database with at a cutoff of 97% as recommended for using PICRUST [31]. In PICRUST, the data was normalized by the known/predicted 16S rRNA copy number abundance (normalize_by_copy_number.py) before KEGG categories were predicted (predict_metagenomes.py and categorize_by_function.py). The Nearest Sequenced Taxon Index (NSTI), which measures the average distance between OTUs and their nearest sequenced genome representatives was computed. The average NSTI value (0.16 ± 0.04) is comparable to the one found in soils and considered as good [31].
For comparison we also used the software FAPROTAX [14]. FAPROTAX extrapolates taxonomic microbial community profiles into putative functional profiles based on a database of cultured microorganisms. The prediction was obtained from the normalized contig-based 16S rRNA OTU table annotated against the greengenes database (collapse_table.py, http://www.zoology.ubc.ca/louca/FAPROTAX/lib/php/index.php?section = Home).
Ocean Sampling Day (OSD) sequence analysis
The OSD sampling stations used in this study (69 samples) and the corresponding environmental data are presented in the Supplementary Table S3. Workable metagenome sequences were retrieved form the online OSD repository. The detail of the sequence pre-processing is available online (https://github.com/MicroB3-IS/osd-analysis/wiki/Guide-to-OSD-2014-data).
For the functional analysis, the comparison of the metagenomic data was conducted with the Commet software as described above for the SOLA data (method 1). For the taxonomic analysis, the 16S rRNA sequences extracted from the metagenomes were retrieved from the EBI repository (https://www.ebi.ac.uk/metagenomics/projects/ERP009703;jsessionid = 038DD38F02117EA5AF698E7C2996778F). The protocol used by the EBI to extract 16S rRNA sequences and build OTU tables can be found here: https://www.ebi.ac.uk/metagenomics/pipelines/2.0. Briefly, 16S rRNA are identified among metagenomic reads with rRNASelector [32] and the OTU table is build with the closed-reference OTU picking protocol in QIIME [33] by clustering reads against a reference sequence collection.
Statistics
Similarity
Similarity between community compositions was computed for the taxonomic composition, the predicted gene composition, the KEGG Orthology (KO) composition and the phylogenetic composition. For the taxonomic, gene and KO composition the Bray-Curtis similarity was computed based on resampled tables using the vegan package in R [34]. The OTU table was square root transformed to reduce the asymmetry of the species distribution [35] and chloroplastic sequences were removed. For the phylogenetic diversity, the 16S rRNA sequences were aligned against complete sequences (length > 15,500 bp) from the greengenes database with SINA [36]. The resulting alignment was checked and corrected manually and was inserted into an optimized tree according to the maximum parsimony criteria without allowing any changes to the existing tree topology with the ARB tool [37]. The UNIFRAC distance (weighted and unweigthed) was computed with the phyloseq package [38] in R.
In order to compare community composition across seasons, a multidimensional scaling (MDS) analysis was conducted based on Bray-Curtis dissimilarity with the phyloseq [38] package in R.
Significance
Linear relationships were tested with ANOVA in R. The significance of changes in gene abundance between seasons was tested with the Welshes t-test implemented in STAMP (P < 0.05) [39]. The list of genes used in the analysis is presented in Supplementary Table S4. The significance of the difference in community composition between seasons was tested with a PERMANOVA (adonis function, vegan package in R).
Results and discussion
We repeatedly sampled the coastal Mediterranean Sea over 3 years and characterized the microbial communities by sequencing 40 metagenomes in depth (Supplementary Table S1 and Supplementary Fig. S1). The metagenomes were used to describe the overall functional attributes of the communities. Rather than arbitrarily selecting specific functional genes to define community-level functions, which would over-simplify the complexity of microbial communities characterized by numerous microorganism-to-microorganism interactions, we used the entire set of metagenome reads, or all predicted proteins, as markers for the overall functionality of the communities. The 16S ribosomal gene, a standard taxonomic marker, was used to describe the composition of the communities.
Our data show that across seasons, the similarity in the communities’ functional attributes was strongly correlated with the similarity in the bacterial community composition between samples (R2 = 0.77, Supplementary Fig. 1a), as well as with the phylogenetic composition (Supplementary Fig. S2). In the case of a strict redundancy, taxonomically different organisms that have the same set of functions could replace each other. In that case, a change in taxonomic composition would not correspond to a change in overall community function. Our demonstration that shifts in the taxonomic structure of microbial communities are associated with shifts in community-level functions thus refutes the hypothesis of strict redundancy.
Similarity in functional attributes is related to similarity in community composition and shows reproducible patterns over time. a The relationship between the similarity in communities’ overall functional attributes and community composition (R2 = 0.8, P < 0.001, F test for the overall significance of the linear regression). b Pairwise comparisons of the overall functional attributes for communities sampled during a 3-year period at the SOLA station in the coastal northwest Mediterranean Sea. The similarity in functional attributes was measured by a direct metagenome-to-metagenome comparison of the sequence content, which gave results similar to the ones obtained by using all predicted proteins (Supplementary Fig. S3). The similarity between communities (1- Bray-Curtis dissimilarity) was estimated from the composition of 16S rRNA genes
Our findings also demonstrate that communities that have many taxa in common also share many functional traits. The same result was obtained when the abundances of all predicted bacterial genes were used as markers for community-level functional profiles (Supplementary Fig. S3a). Our data support Finlay and colleagues’ early statement that “the concept of redundancy of microbial species has little meaning” [40]. However, recent reports showed high functional redundancy for marine microbes at a global scale [13, 14]. These previous studies were based on the indirect characterization of functional profiles [14], which were extrapolated from the similarity between taxonomic annotations of environmental samples and cultured microorganisms, or based on annotated genes only [13], which again originate from cultured microorganisms. These earlier studies thus relied on a limited number of genes. The fact that these genes could be found in various communities indicates the possible presence of a partial functional redundancy. Partial redundancy could be defined as the coexistence of organisms that share some specific function (e.g., two ammonia oxidizers) but may nevertheless differ in other ecological requirements.
For comparison purposes, we applied two different indirect approaches used earlier to infer function from taxonomy (FAPROTAX [14] and PICRUSt [31]) and found a poor correlation between similarity in community composition and similarity in community functions (R2 = 0.16 for both, Supplementary Fig. S3b, c). Functional profiles were also built from our metagenomic data by directly annotating reads against the KEGG database. Comparisons of the community vs. functional similarity showed that the KEGG-based gene composition did not explain the taxonomic composition as well as the all read-based composition (R2 = 0.58, Supplementary Fig. S3d). Our data demonstrate the need to consider the entire set of predicted genes within communities, which is essential to simultaneously target all community-level functions. We thus highlight the importance of focusing on multi-functionality [41], or community-aggregated functional profiles [21], in microbial ecosystems in which species are highly connected with complementarity across functions [42]. A without a priori method is essential to achieve the community-level trait-based approach required to make significant progress in understanding the role of microbes in the environment. Inversely, the use of taxonomy-based predicted functions, or annotated genes only, limits the mapping of the functional properties of natural communities by relying on databases built from cultured organisms. As they are derived from only a few cultured organisms from the marine environment, the existing databases contain a limited proportion of the proteins potentially found in nature [43]. In our case, an average of only 22% of the predicted genes could be annotated against the KEGG database over the 3 years of the study (Supplementary Table S2). Thus, studies based on only known genes cannot reflect the true associations existing between community function, community composition, and the environment.
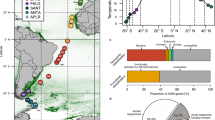
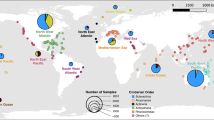
To validate our results regarding the relationship between the community structure and functional attributes, our temporal study was extended to incorporate a spatial scale. To focus on geography alone, without any interference due to seasonality, we used global data from Ocean Sampling Day (OSD) [44] that originated from samples taken concurrently during the week of the summer solstice in June 2014. We focused on the Mediterranean Sea to remain within the hypothetical frame of a single biotope, and found that there was a significant correlation between the similarity in communities’ overall functional attributes and similarity bacterial community composition (Fig. 2a). Our test was subsequently extended to Atlantic Ocean samples, which show more variations in environmental conditions due to the wider range of latitudes sampled (Supplementary Fig. S4). Again, there was a significant correlation between the overall functional properties of the communities and the bacterial community composition (Fig. 2b). However, at the ocean scale, the correlation was lower, and a larger number of samples had dissimilar functional properties and variable community similarities. We propose that the looser relationship observed for the wider geographical scale is a reflection of the different biotopes sampled (i.e., different salinity, temperature, and day length in the Atlantic Ocean samples (Supplementary Table S3)) rather than an indicator of true functional redundancy. When different biotopes are studied simultaneously, the lack of a relationship could highlight the presence of different microbial ecotypes able to conduct a subset of similar functions under different environmental conditions [45].
The similarity in overall community functional attributes and community composition in the Mediterranean Sea (a) and Atlantic Ocean (b). All communities were sampled during the 2014 summer solstice. The similarity in overall community functional attributes was measured by a direct metagenome-to-metagenome comparison of the sequence content. The similarity between communities (1- Bray-Curtis dissimilarity) was estimated from the composition of 16S rRNA genes
In the coastal northwest Mediterranean, the functional similarity between two communities was highest when samples were taken 1 year apart and lowest when samples were taken 6 months apart (Fig. 1b), which reflects the strong seasonality of the northwest Mediterranean. Both the functional and community composition showed significant differences between seasons (Fig. 3) (R2 = 0.5 and R2 = 0.35, respectively, PERMANOVA, P < 0.01).
The seasonal structure of the microbial communities. a The structure of the overall community functional attributes across different seasons. b The structure of the microbial community composition across different seasons. The nonmetric multidimensional scaling plots are based on a Bray-Curtis dissimilarity calculated from the composition of all predicted proteins (a) or the composition of 16S rRNA genes (b)
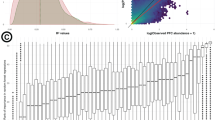
To test if we could identify known functional processes that varied with seasons, we selected key oceanic marker genes [13] among the genes that could be annotated in our dataset (Supplementary Table S4). Close to 100 marker genes showed significant differences based on the season (Supplementary Fig. S5), and 5 out of the 13 selected processes varied significantly (Fig. 4). Winter showed an enrichment in genes related to prokaryotic carbon fixation, nitrogen metabolism, and manganese-related metabolism. Spring showed an enrichment in genes associated with iron-related metabolism, and summer had more genes associated with flagellar assembly. Flagellar assembly was previously shown to be more common in surface waters compared to deeper waters [5] and is related to organisms swimming toward particles to feed. Genes related to anoxygenic photosynthesis had the lowest abundance in winter (Fig. 4), which is in agreement with earlier results based on infrared microscopy [46].
We also found that the richness of the functional profiles from the metagenomic data was significantly correlated with the richness of bacterial communities across the samples (R2 = 0.69, P < 0.01) (Fig. 5). This observation suggests that microbial communities that are taxonomically richer also have a richer array of functional genes. Such a relationship has also been shown in soil using an approach based on the diversity on annotated genes [11]. For marine microbes, there is little information available, but a decline in the diversity of protein-coding gene categories with depths coincided with a decline in taxonomic diversity [47], and higher 16S rRNA diversity reflected a higher diversity of annotated gene transcripts [48]. Our results showing a relationship between community composition and function are important in the context of the debate in ecology regarding whether to disentangle taxonomic diversity from direct changes in function when evaluating the impact of biodiversity on ecosystem function.
At a single site in the northwest Mediterranean, we detected more than 1 million predicted genes that could not be annotated and demonstrated that this huge number of unknown potential microbial proteins corresponded to unique features rather than to redundant functional attributes. Our data showed that non-annotated genes must be accounted for to obtain a correct interpretation of function-based data, which has strong implications for functional studies in microbial ecology. When such large numbers of non-redundant and unknown predicted genes are documented from a single marine site, it suggests that the catalog of genes reported from the global ocean [13] represents a huge genetic reservoir of unknown proteins.
References
Falkowski PG, Fenchel T, Delong EF. The microbial engines that drive Earth’s biogeochemical cycles. Science. 2008;320:1034–9.
Sogin ML, Morrison HG, Huber JA, Welch DM, Huse SM, Neal PR, et al. Microbial diversity in the deep sea and the under explored “rare biosphere”. Proc Natl Acad Sci USA. 2006;103:12115–20.
Fuhrman JA, Cram JA, Needham DM. Marine microbial community dynamics and their ecological interpretation. Nat Rev Micro. 2015;13:133–46.
Hugoni M, Taib N, Debroas D, Domaizon I, Jouan Dufournel I, Bronner G, et al. Structure of the rare archaeal biosphere and seasonal dynamics of active ecotypes in surface coastal waters. Proc Natl Acad Sci USA. 2013;110:6004–9.
DeLong EF, Preston CM, Mincer T, Rich V, Hallam SJ, Frigaard NU, et al. Community genomics among stratified microbial assemblages in the ocean’s interior. Science. 2006;311:496–503.
Ladau J, Sharpton TJ, Finucane MM, Jospin G, Kembel SW, O’Dwyer J, et al. Global marine bacterial diversity peaks at high latitudes in winter. ISME J. 2013;7:1669–77.
Galand P, Salter I, Kalenitchenko D. Microbial productivity is associated with phylogenetic distance in surface marine waters. Mol Ecol. 2015;24:5785–95.
Hutchinson GE. The paradox of the plankton. Am Nat. 1961;95:137–45.
Allison SD, Martiny JBH. Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci USA. 2008;105:11512–9.
Yin B, Crowley D, Sparovek G, De Melo WJ, Borneman J. Bacterial functional redundancy along a soil reclamation gradient. Appl Environ Microbiol. 2000;66:4361–5.
Fierer N, Ladau J, Clemente JC, Leff JW, Owens SM, Pollard KS, et al. Reconstructing the microbial diversity and function of pre-agricultural tallgrass prairie soils in the United States. Science. 2013;342:621–4.
Fuhrman JA, Hewson I, Schwalbach MS, Steele JA, Brown MV, Naeem S. Annually reoccurring bacterial communities are predictable from ocean conditions. Proc Natl Acad Sci USA. 2006;103:13104–9.
Sunagawa S, Coelho LP, Chaffron S, Kultima JR, Labadie K, Salazar G, et al. Structure and function of the global ocean microbiome. Science. 2015;348:1261359.
Louca S, Parfrey LW, Doebeli M. Decoupling function and taxonomy in the global ocean microbiome. Science. 2016;353:1272–7.
Jurburg SD, Salles JF. Functional redundancy and ecosystem function—the soil microbiota as a case study. L. Yueh-Hsin, J.A. Blanco, R. Shovonlal (Eds.), Ecosystems—Linking Structure and Function, InTech Open Science, Rijeka (2015), pp. 29-42.
Bradford MA, Fierer N. The biogeography of microbial communities and ecosystem processes: implications for soil and ecosystem models. In: Wall DH, Bardgett RD, (eds). Soil Ecology and Ecosystem Services.. Oxford: Oxford University Press; 2012. p. 424.
Tréguer P, Le Corre P. Manuel d’analyse des sels nutrifs dans l’eau de mer: utilisation de l’Autoanalyzer II Technicon (R). Université de Bretagne Occidentale, Brest, France; 1975.
Solorzano L. Determination of ammonia in natural waters by the phenolhypochlorite method. Limnol Oceanogr. 1969;14:799–801.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014; 30:2114-20
Peng Y, Leung HC, Yiu S-M, Chin FY. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics. 2012;28:1420–8.
Fierer N, Barberán A, Laughlin DC. Seeing the forest for the genes: using metagenomics to infer the aggregated traits of microbial communities. Front Microbiol. 2014; 5:614.
Maillet N, Collet G, Vannier T, Lavenier D, Peterlongo P. COMMET: comparing and combining multiple metagenomic datasets. IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Belfast, United Kingdom. 2014:94–8.
Ulyantsev VI, Kazakov SV, Dubinkina VB, Tyakht AV, Alexeev DG. MetaFast: fast reference-free graph-based comparison of shotgun metagenomic data. Bioinformatics.2016; 32:2760-7.
Noguchi H, Taniguchi T, Itoh T. MetaGeneAnnotator: detecting species-specific patterns of ribosomal binding site for precise gene prediction in anonymous prokaryotic and phage genomes. DNA Res. 2008;15:387–96.
Li W, Godzik A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics. 2006;22:1658–9.
Li R, Yu C, Li Y, Lam T-W, Yiu S-M, Kristiansen K, et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics. 2009;25:1966–7.
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucl Acids Res. 2015; 44:457-62.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26:2460–1.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucl Acids Res. 2013;41:D590–6.
Li H, Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 2009;25:1754–60.
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol. 2013;31:814–21.
Lee J-H, Yi H, Chun J. rRNASelector: a computer program for selecting ribosomal RNA encoding sequences from metagenomic and metatranscriptomic shotgun libraries. J Microbiol. 2011;49:689–91.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335–6.
Dixon P, Palmer M. VEGAN, a package of R functions for community ecology. J Veg Sci. 2003;14:927–30.
Legendre P, Legendre LF. Numerical ecology. Amsterdam, The Netherlands, vol. 24. Elsevier; 2012.
Pruesse E, Peplies J, Glöckner FO. SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics. 2012;28:1823–9.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Buchner A, et al. ARB: a software environment for sequence data. Nucl Acids Res. 2004;32:1363–71.
McMurdie PJ, Holmes S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. ploS one. 2013;8:e61217.
Parks DH, Tyson GW, Hugenholtz P, Beiko RG. STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics. 2014;30:3123–4.
Finlay BJ, Maberly SC, Cooper JI. Microbial diversity and ecosystem function. Oikos. 1997;80:209–13.
Hector A, Bagchi R. Biodiversity and ecosystem multifunctionality. Nature. 2007;448:188–90.
Giovannoni SJ, Cameron Thrash J, Temperton B. Implications of streamlining theory for microbial ecology. ISME J. 2014;8:1553–65.
Rinke C, Schwientek P, Sczyrba A, Ivanova NN, Anderson IJ, Cheng J-F, et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature. 2013;499:431–7.
Kopf A, Bicak M, Kottmann R, Schnetzer J, Kostadinov I, Lehmann K, et al. The ocean sampling day consortium. Gigascience. 2015;4:27.
Sintes E, Bergauer K, De Corte D, Yokokawa T, Herndl GJ. Archaeal amoA gene diversity points to distinct biogeography of ammonia-oxidizing Crenarchaeota in the ocean. Environ Microbiol. 2013;15:1647–58.
Ferrera I, Borrego CM, Salazar G, Gasol JM. Marked seasonality of aerobic anoxygenic phototrophic bacteria in the coastal NW Mediterranean Sea as revealed by cell abundance, pigment concentration and pyrosequencing of pufM gene. Environ Microbiol. 2014;16:2953–65.
Bryant JA, Stewart FJ, Eppley JM, DeLong EF. Microbial community phylogenetic and trait diversity declines with depth in a marine oxygen minimum zone. Ecology. 2012;93:1659–73.
Gilbert JA, Field D, Swift P, Thomas S, Cummings D, Temperton B, et al. The taxonomic and functional diversity of microbes at a temperate coastal site: a ‘multi-omic’ study of seasonal and diel temporal variation. PLoS one. 2010; 5:e15545.
Acknowledgements
Raw sequences were archived in the EBI repository under accession number PRJEB26919. The work of PEG was supported by the Agence Nationale de la Recherche (ANR) through the projects EUREKA (ANR-14-CE02-0004-01). We thank the captain and crew of the Nereis II, Eric Maria, and Louise Oriol for assisting with the collection and analysis of samples over the time series. We extend our acknowledgments to all the researchers that were involved in working with the time series over the years.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Galand, P.E., Pereira, O., Hochart, C. et al. A strong link between marine microbial community composition and function challenges the idea of functional redundancy. ISME J 12, 2470–2478 (2018). https://doi.org/10.1038/s41396-018-0158-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0158-1
This article is cited by
-
Contrasting resistance of prokaryotic plankton biomass and community composition to experimental nutrient inputs in a coastal upwelling system (NW Spain)
Hydrobiologia (2024)
-
Mulches assist degraded soil recovery via stimulating biogeochemical cycling: metagenomic analysis
Applied Microbiology and Biotechnology (2024)
-
Soil bacterial responses to disturbance are enlarged by altitude in a mountain ecosystem
Journal of Soils and Sediments (2023)
-
Earthworm and soil microbial communities in flower strip mixtures
Plant and Soil (2023)
-
Spatiotemporal Dynamics of Bacterial Taxonomic and Functional Profiles in Estuarine Intertidal Soils of China Coastal Zone
Microbial Ecology (2023)