Abstract
What happens to tap water when you are away from home? Day-to-day water stagnation in building plumbing can potentially result in water quality deterioration (e.g., lead release or pathogen proliferation), which is a major public health concern. However, little is known about the microbial ecosystem processes in plumbing systems, hindering the development of biological monitoring strategies. Here, we track tap water microbiome assembly in situ, showing that bacterial community composition changes rapidly from the city supply following ~6-day stagnation, along with an increase in cell count from 103 cells/mL to upwards of 7.8 × 105 cells/mL. Remarkably, bacterial community assembly was highly reproducible in this built environment system (median Spearman correlation between temporal replicates = 0.78). Using an island biogeography model, we show that neutral processes arising from the microbial communities in the city water supply (i.e., migration and demographic stochasticity) explained the island community composition in proximal pipes (Goodness-of-fit = 0.48), yet declined as water approached the faucet (Goodness-of-fit = 0.21). We developed a size-effect model to simulate this process, which indicated that pipe diameter drove these changes by mediating the kinetics of hypochlorite decay and cell detachment, affecting selection, migration, and demographic stochasticity. Our study challenges current water quality monitoring practice worldwide which ignore biological growth in plumbing, and suggests the island biogeography model as a useful framework to evaluate building water system quality.
Similar content being viewed by others
INTRODUCTION
Potable water supply infrastructure harbors complex microbial ecosystems, affecting the lives of 3.6 billion people living in urbanized areas [1,2,3]. In developed countries, the water supply networks in homes, schools, and hospitals (collectively termed as premise plumbing), have been identified as a hotspot for waterborne pathogens [1, 4]. In the United States, three plumbing-associated opportunistic pathogens (i.e., Legionella pneumophila, Mycobacterium avium, and Pseudomonas aeruginosa) are linked to ~41,000 infections per year, mostly affecting elderly populations and immunocompromised individuals [5, 6]. However, although drinking water is closely monitored for its biological quality based on indicator microorganisms in public supplies, active surveillance of water after entering a building is not included in current regulations [7]. In addition, unlike traditional fecal contamination, plumbing-associated pathogens are freshwater microorganisms that naturally occur in the biofilms of water systems, not necessarily correlated with occurrences of indicator microorganisms in current water quality monitoring [5, 8, 9]. A potential link has been reported between bacterial community composition and occurrences of opportunistic pathogens [10], suggesting the microbiome could be a prospective indicator to aid water quality surveillance and management.
One crucial characteristic of the in-building water supply system is water stagnation. In a city-wide water supply network, water flow seldom fully stops due to the aggregated water demand in urban areas. In buildings, however, water can frequently stay unused for extended periods of time due to individual’s water-use behaviors, thus allowing for lengthy incubation times for processes crucial to biological activity, such as the decay of disinfectants [11,12,13,14]. In addition, substantially smaller pipe diameters are used in buildings than in city infrastructure, which can result in over 10 times larger contact area per unit volume [1, 15, 16]. This further leads to an increased surface area between tap water and pipe-wall biofilms, and, hence, higher apparent reaction rates. Stagnation effects remain unaddressed in building designs. In fact, green-building design tends to focus on water conservation, which essentially extends water stagnation and accelerates water quality deterioration [16, 17]. Tracking stagnation effects on the bacterial communities in tap water can help to elucidate how tap water deviates from the city supply and to understand what magnitude of deviation is permissible. Outside of a small number of studies with community fingerprinting methods (i.e., T-RFLP or DGGE) that indicate community structure shifts [9, 18], detailed culture-independent field experiments focused on the effects of tap water stagnation on bacterial community composition have not been conducted.
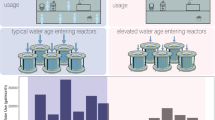
Here, we present a field experiment of bacterial community assembly within-building drinking water systems. The experiment was conducted twice in three student dormitory buildings at the University of Illinois at Urbana-Champaign (UIUC) during unoccupied times over a summer break before and after week-long stagnation (Fig. 1a). To map the bacterial communities to the pipe network structure, we sampled the post-stagnation water by collecting consecutive segments of flowing tap water (Fig. 1b). The samples were characterized using 16S rRNA gene amplicon sequencing, flow cytometry, and water chemistry analysis. In addition, we combined migration and demographic stochasticity modeling [19, 20] and reaction kinetics simulation to gain a mechanistic understanding of bacterial community assembly during tap water stagnation.
Sampling of building-impacted water from proximal to distal pipes. a Building impact to water microbiome was represented by allowing a pause in water use for ~6 days after a thorough purging with fresh city tap water. b Building-impacted water was sampled from three buildings, three faucets in each building from different floor levels, at two different time points. Water samples were taken consecutively at 100-mL intervals and pooled into four sites per faucet to represent the physical structure of pipes, namely distal-1 (first 100 mL) with, distal-2, (200–1000 mL), proximal-1 (1100–2000 mL), and proximal-2 (2100–3000 mL). One fresh city tap water sample was collected at each faucet at the start of the experiment
RESULTS
Drastic change in bacterial cell count and community composition after tap water stagnates in buildings
Tap water cell count significantly increased during stagnation, from less than 103 cells/ml (below detection in flow cytometry) to up to 7.8 × 105 cells/ml. After stagnation, the bacterial community in tap water exhibited decreased richness and evenness compared to fresh tap water samples, as measured by an array of alpha diversity matrices (i.e., the observed number of operational taxonomic units (OTUs) defined at 97% sequence similarity, Chao 1 index, Shannon index, etc.; Figure S1). Post-stagnation water community also had distinctive composition and structure from fresh tap water samples (Fig. 2). Beta-diversity measured by the Bray–Curtis dissimilarity showed that post-stagnation water communities formed separate groups from fresh city tap water (Fig. 2a). Principle coordinate analysis of Bray–Curtis dissimilarity and other beta-diversity metrics, including the Jaccard index and UniFrac distance (weighted and unweighted), also showed grouping between fresh and post-stagnation water samples (Figure S2). Post-stagnation water grouped more drastically from fresh tap water samples when measured by beta-diversity measurements considering abundance (Bray–Curtis dissimilarity and weighted UniFrac distance) than the presence/absence of taxa (the Jaccard index and unweighted UniFrac distance), suggesting that community difference was largely driven by differentiated abundance in common taxa.
Bacterial community composition in building-impacted water varies by the location within pipes. a Non-metric multidimensional scaling plot showing Bray–Curtis dissimilarities between building-impacted water samples, fresh city tap water, and biofilms from water meters. Building-impacted water formed distinctive groups separate from water or biofilm communities in the distribution systems. Samples separated in groups by the location within pipes, but not by different buildings, floors, or time points. b Phylum-level community composition of city tap water and building water supplies in one replicate and water meter biofilm community. Proteobacteria was separated into alpha-, beta-, and gamma-Proteobacteria. Bacterial phyla with abundance less than 1% were binned into “others”
In terms of community composition, the fresh tap water samples harbored bacterial phyla Proteobacteria, Cyanobacteria (mostly classified to “Candidatus” Melainabacteria), Bacteroidetes, Planctomycetes, Actinobacteria, and Firmicutes; yet the post-stagnation tap water was dominated by Proteobacteria (Fig. 2b). A transition in community composition was also apparent at finer taxonomic resolutions (Figure S3). Consistent with our previous studies of the municipal water distribution system supplying these buildings [21], OTUs related to the Proteobacteria families including Erythrobacteraceae, Sphingomonadaceae, Rhodocyclaceae, and Comamonadaceae were abundant (Figure S4 and Table S1). These families are aerobic heterotrophs frequently observed in the drinking water ecosystem [22,23,24]. Several OTUs related to the Methylocystis genus were detected (e.g., OTU0005, whose relative abundance ranked 5th in fresh tap water and 4th in post-stagnation water samples); most representative isolates reported from this genus so far were obligate methanotrophs, with the exception of facultative methanotrophic isolates with clear preference of methane [25, 26]. The dissolved methane from the source water, a methanogenic Mahomet aquifer, might support their growth [27,28,29]. Notably, an OTU related to 'Candidatus' Melainabacteria was ranked the second abundant in fresh tap water (OTU00026), yet declined in rank in post-stagnation water samples. This recently-named 'Candidatus' phylum sibling to Cyanobacteria has been detected in subsurface aquifer, sediment, and the human gut microbiome. Genome reconstruction suggests this clade to be non-photosynthetic, anaerobic, motile, and obligately fermentative [30]. With increasing detection of Candidatus Melainabacteria in culture-independent 16S rRNA gene surveys of drinking water supply systems, such as Champaign, Illinois [29], Ohio River Basin [31], and New Orleans, Louisiana [32], further studies on their roles in the engineered water environment are needed.
Remarkably, community composition in the same faucet between temporal replicates was highly reproducible. We specifically tested the reproducibility of the spatial trajectory of abundant taxa (>1% in at least one sample) using a Spearman correlation, which resulted in a median of 0.78, and reached 0.9–1.0 for certain taxa. This consistency suggests that minimal technical variation occurred between replications, and the community assembly processes in building plumbing could be modeled and predicted.
Because tap water is in contact with inner surfaces of water pipes during stagnation, we hypothesized that the difference in pre- and post-stagnation communities resulted from seeding by biofilms formed in pipes. Due to the logistical constraints of sampling within-building pipe biofilms, we used previously published data of biofilms from household water meters in the same water supply (WM biofilms) as a proxy [29]. Comparing post-stagnation water communities to biofilms collected from household water meters, we found a high degree of similarity between post-stagnation samples at the distal pipes (i.e., the first 100 mL water coming out of tap) and biofilm communities.
Post stagnation, water community composition is consistent across buildings, yet varies by proximity to the city water supply
Sampling across buildings, floors, and faucets revealed that bacterial communities in post-stagnation water communities vary by distance from the faucets (Fig. 2a). In a non-metric multidimensional scaling (nMDS) plot based on Bray–Curtis dissimilarity, communities from the distal pipes (first 100 mL) grouped distinctively from proximal pipes (1100–2000 mL and 2100–3000 mL). Samples from the transitional sections (200–1000 mL) were scattered across these two groups (Fig. 2a). Other sample characteristics, including building ID, floor number, and faucet ID were not associated with any obvious patterns in the community composition data (Figure S5). Analysis of similarity (ANOSIM) [33, 34] confirms that sample types (i.e., fresh city water and distance from faucets) yielded the highest amount of variance explained among all factors (R = 0.508; p = 0.01). Building and floor were also significantly associated with community composition (p = 0.0049 for floor and p = 0.03 for buildings), but the amount of variance explained was lower compared to sample type (R = 0.028 for floor and R = 0.039 for building). Faucet (p = 0.21) and temporal replicates (p = 0.182) did not significantly affect the community structure. In addition, building-impacted tap water communities progressed towards higher similarity to biofilm communities as samples moved from proximal to distal structures (faucet).
Pipe diameter and area-to-volume ratio gradients in the water supply networks
We reasoned that correlation between community composition and distance to faucets resulted from the gradient in pipe diameters inherent to building pipe architecture. By engineering guidelines, building water supplies are designed in a tree-like structure, in which pipe diameter decreases as the pipes approach the water-use apparatus. In the buildings we surveyed, the pipe diameter ranged from 1.25 to 2 inches in source proximal vertical pipes (Branch Level I), 0.5 to 1 inch in the next horizontal pipes (Branch Level II), and were 0.375 inch in the final distal pipes (Branch Level III, Fig. 3a). This results in a sharp increase in surface-area-to-volume ratio as the pipe branch deepens (Fig. 3b), which could lead to drastically different dynamic profiles in the bulk volume for the same surface reactions.
A reaction kinetics model for microbial dispersal and disinfectant decay predicts that smaller pipe diameters favor higher dispersal from biofilms and longer sub-inhibitory disinfectant exposure. a Branch levels from proximal to distal branches. b Pipe diameters from buildings surveyed in this study vary by branch levels. c Specific area of pipes varies by branch level. d Dynamic profiles of free chlorine (sodium chlorite) concentration simulated for branch level I, II, and III within one floor show an increased overall decay rate in narrower (distal) pipes. e Cell count dynamics simulated from dispersal and growths/deaths shows faster overall increase in narrower (distal) pipes
Modeling diameter effects on ecologically relevant physical processes
We used first-order reaction models to simulate how pipe diameter affects the dynamics of cell count and disinfectant concentrations through microbial dispersal and disinfectant reduction. These two surface reactions have been reported to occur on biofilm-covered pipes in both full-scale water distribution systems and laboratory experiments [35,36,37,38]. The growth/death of bacterial cells in the water column was modeled dynamically in tandem with the seeding from biofilms and inhibition from disinfectants. To simulate the diameter effect on physical and chemical processes, we simplified the scenario as a monoculture with uniform detachment rate, growth rate, and sensitivity to disinfectants. Reactions were mapped to the one of the buildings in our experiment to represent building pipe geometry. Disinfectant consumption in the water column was also simulated, but the extent was much smaller than the wall reactions. We ran the simulation over 7 days at varying pipe diameters, with the initial conditions set the same as the fresh city tap water. The results showed that both the disinfectant and cell count were mainly affected by the pipe diameter, and were minimally different in axial and radial directions (i.e., reaction-limited). Within one floor, the simulation showed a monotonic increase of disinfectant consumption rate (Fig. 3d) and cell count (Fig. 3e) from large-diameter (proximal) to narrow pipes (distal). Notably, cell counts in the distal pipes were shown to be 1000 times higher than the main vertical pipe by the end of a week-long stagnation. Conversely, disinfectant was consumed to a very-low concentration (<0.2 mg/L) in the 0.375-inch, final distal tap within 78 h, yet remained above the same concentration to 162 h in the 1-inch, main vertical pipe, suggesting a much shorter exposure to inhibitory levels of the disinfectant in the narrower pipes.
Cell count peaked and disinfectant was depleted in the first 100 mL of water out of the taps
This model is supported by cell count and disinfectant measurements in the experiment. Despite the low abundance of microbial cells in city tap water (<1000 cells/mL), the cell count in building water supplies increased to 104–106 cells/mL after the week-long pause in water use. Cell counts correlated strongly to the plumbing structure (Fig. 4a) and decreased as samples transitioned from the distal (close to faucets) to the proximal structures (close to the city supplies), whereas disinfectant concentration showed the opposite trend and was completely depleted at the first 100 mL (Fig. 4b). This simple model was supported by statistical analysis. When the cell count data from each building was subjected to forward selection of explanatory variables including flow volume (representation of physical structure of pipes), floor, and building in a general linear model, flow volume was the first explanatory variable chosen among all tested variables (p < 2.2 × 10−16 as opposed to p > 0.001 for other factors tested). Other variables were not of comparable significance (floor: p = 0.0025; building p = 0.0031). Similarly, flow volume was the predictor that explained the highest variation for free chlorine data (data not shown).
Community assembly model shows a decreasing role of city water supply in post-stagnation water at distal pipes
Ideally, we would like to fully explain the assembly of post-stagnation water communities, teasing apart migration from two possible sources, i.e., fresh city tap water and pipe biofilms, stochasticity of growth/death during stagnation time, and niche assembly in the different pipe environments. However, this is difficult to test in the field, because tracking biofilms in buildings actively in use would require destructive methods (e.g., demolition of buildings). Therefore, we asked a simpler question instead: can the post-stagnation water communities be assembled solely from migration and demographic stochasticity of city water supply? We built a mainland-island model between fresh and post-stagnation water samples, treating the mean OTU abundance of fresh tap water samples from all the different buildings, floors, and faucets in the experiment as the city-wide mainland, and the post-stagnation tap water from each pipe locations as islands (Fig. 5a).
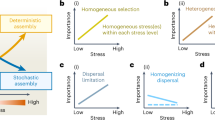
Modeling the role of city water supply to building-impacted water. Results of neutral model fitting shows an increase in the deviation from city water supplies as water approaches the faucets, in both OTUs defined at 97 and 99% sequence similarity (a). b–e Fitting of neutral processes (i.e., migration and demographic stochasticity) from city water supply to the building tap water islands in the flow segments of first 100 mL (b), 200–1000 mL (c), 1100–2000 mL (d), and 2100–3000 mL (e)
The result showed a higher model fit in proximal pipes (Fig. 5d, e R2 = 0.41 and 0.48 for proximal-1 and proximal-2 samples, respectively) than in distal pipes (Fig. 5b, c R2 = 0.21 and 0.19 for distal-1 and distal-2 samples, respectively). The model fit is consistent for OTUs defined at 97 and 99% sequence similarity (Fig. 5a), which further supports direct migration and demographic processes. This result suggested that city water supply contribute largely to the water community assembly in proximal pipes closer to the city, whereas other processes (i.e., selective growth and migration from biofilms) could drive community assembly in distal pipes. This agreed with our reaction and dispersal simulation (Fig. 3), which shows lower disinfectant levels in distal pipes throughout the stagnation time (Fig. 3d) and higher dispersal from biofilms in distal pipes (Fig. 3e). We also tested a biofilm mainland with water meter biofilm data as a proxy, which showed higher model fit in distal pipes than proximal pipes (Figure S6).
DISCUSSION
Our analysis of bacterial communities in building-impacted tap water revealed a spatially structured community within the plumbing system following stagnation. These spatially structured communities were highly reproducible. We hypothesized that the assembly of these communities was mediated by the physical structure of pipes (i.e., pipe diameter). This hypothesis was first supported by a reaction-transport model, which showed that pipe diameter deterministically affected two ecologically relevant physical/chemical processes, namely the dispersal of bacteria from pipe surface biofilms and the decay of disinfectants. A neutral community assembly model was then applied to show that an island biogeography model accounting for migration and demographic stochasticity from the city water supply could explain the community composition at the proximal pipes. The variation explained by the city water supply declined as the physical proximity of the water samples approached the faucets (distal pipes), which co-occurred with a narrowing pipe diameter. Due to sampling difficulty, we were unable to obtain data and separate selection from the stochastic processes arising from another possible mainland, the pipe-wall biofilms, and had to compromise using water meter biofilm communities from a previous study as a proxy. Despite that, these results together connected a design parameter (i.e., diameter) of the physical structure of building water supplies to the microbial community therein, and showed that the microbiome in tap water could be modeled, predicted, and included in building design criteria.
Currently, to manage the public health risks associated with waterborne diseases, water treatment and monitoring rely on the detection of biological contaminants based on plate counting (e.g., heterotrophic plate count, coliforms, or Escherichia coli) [39, 40], which has limited power to reflect the biotic complexity of a building’s water supply system and any microbiological problems associated with it. The mainland-island model applied in this study to 16S rRNA gene sequence data showed that tap water microbial communities can be used to infer the level of deviation from the city supply due to in situ biofilm development.
The microbiome of building water supplies can be used as a probe for monitoring infrastructure health. While sampling pipes requires excavation, which is cost-prohibitive or unfeasible, water microbiomes are present in a flowing medium that is easily accessible. Guided by the mainland-island model, buildings in a city can be rigorously sampled and cataloged, a background level of building-induced deviation can be established, and anomalies can be detected. Microbial biosensors could be used to address long-standing issues in infrastructure management, like disentangling the extent of contamination risk posed by municipal water supplies or by private infrastructure [15]. With the aid of our model, this problem can be tackled by testing whether the level of deviation from city-water microbiome exceeds an expected range.
Our results showed that small-diameter pipes at the distal ends of building water supplies harbored the highest cell counts and deviated most from the city-water supply microbiome. This highlights small-diameter pipes as a site for biological regrowth. Small-diameter pipes are widely used in plumbing at the connections to residential water-use devices, such as shower hoses, kitchen hoses, dish washers, and ice makers, and medical devices such as dental water lines. However, this risk cannot be managed by simply increasing the size of pipes. Larger diameter pipes could actually lead to increased levels of water stagnation and consequent water quality deterioration [15, 41]. Instead, this invites innovative solutions. In the near term, hospitals and extended care facilities, where plumbing-associated infections are more common [42], may want to consider upgrading their end-point disinfection to counteract hypochlorite decay and address within-pipe cell growth. Precise flushing of smaller-diameter pipes, rather than the whole building, could also be included in building management algorithms to prevent stagnation while minimizing water waste. In the long term, small-diameter pipes can be made with biofilm-inhibitive materials [43, 44]. While water conservation is a major green-building design focuses water stagnation and degradation of water quality should be taken into account as well [17]. Future conservation strategies could redirect water flow to non-potable use based on bacterial community-wide response to stagnation time and pipe diameter.
METHODS
Study sites
We conducted the experiment in three 4-storey buildings in Champaign, Illinois. The water supply in the area sourced from groundwater aquifers and treated with a conventional treatment process. The treatment processes, and the diversity of the suspended and biofilm communities in this system has been reported in our previous study [21]. The system carried stable free chlorine as residual disinfectants during the whole time of this study.
Three residence halls T, V, and S were chosen for their design and availability. All potable water in these buildings were supplied with water directly from the water main, without water tanks and additional storage structures involved. This is the most basic type of indoor water supply design and we chose it to eliminate interference from building architectural complexity. These buildings were managed by University Housing at UIUC. The T Hall and V Hall were built in 1955 and S Hall in 1963 as part of the on-campus residence at UIUC. In each building, we controlled access of the second, third and fourth floor during the experiment.
Water supply system design
We accessed the original blueprints and project manual for the studied buildings to acquire design data. Missing details due to common knowledge in the trade were supplemented by estimation from architects at University Housing. Level I branches were designed as 2” to 1” from Floor 1 to 4, connected to Level II branches of 3/4”–1/2”, and then Level III branches of 3/8”. The specific area in the pipe sections increased as the branch deepens, from 1.05–1.57 cm−1 in Level I to 4.2 cm−1 in Level III lines. The volume of water in each section can also be estimated based on branch diameter and length. Common to all the floors, the Level III branch (3/8”) could hold 21 mL water, and the Level II branch 1212 mL. The capacity of Level III branches differed by floor and ranged 1389 mL to 3125 mL from 4th floor to 2nd floor.
Sample collection
We treated the buildings as natural laboratories to study the effect of stagnation on the microbial communities in water. The samples were collected from cold water faucets at the handwashing basins in the dormitory bathrooms. Before the experiment, aerators as part of the faucet accessory structure were removed to eliminate possible interference on the community profiles [45]. Prior to the designed stagnation, tap water in the studied pipelines were replenished by running tap water for 30 min, and then the fresh city tap water (pre-stagnation) samples were collected on each floor to represent tap water without the influence of stagnation. After that, the sites were controlled for 5–6 days with controlled access. At the end of the stagnation period, the post-stagnation samples were collected at different flow volume by letting tap water flow out at minimum flow rate. Samples were taken as every 100 mL for the first liter and every 500 mL in the 2100–3000 mL for free chlorine and total cell count measurement, and were pooled as distal-1 (first 100 mL), distal-2 (200–1000 mL), proximal-1 (1100–2000 mL), and proximal-2 (2100–3000 mL) for community analysis. This sampling method was intended for harvesting a consistent level of bacterial cells for DNA extraction, yet we acknowledge that the sampling volume was uneven and might introduce biases towards detection of rare taxa. In all buildings, biological replicates of fresh city tap water (pre-stagnation) and post-stagnation samples were taken as one faucet per floor from three floors in each building, and then repeated once for temporal replication. Another replication for inter-faucet difference was designed and taken as three faucets per floor from three floors in one building.
Water quality of pre-stagnation samples and post-stagnation samples (measured collectively) were compared (Table S2). Other than near depletion of free chlorine reported in Results (0.3 ± 0.3 mg-Cl/L, p < 0.001), we found a rise in water temperature to room temperature and release of plumbing metals (i.e., zinc and copper from brass fittings), which is expected from water stagnation and supports a successful water-use control. Other parameters were stable, including below-regulation levels of copper and lead (Lead and Copper Rule, lead < 15 ppb and copper < 1.5 ppm).
Biomass collection and DNA extraction
Water samples were collected at the sampling site with steam-sterilized polypropylene carboys or bottles (Fisher scientific), and transferred to the laboratory immediately, then concentrated by 0.22 µm filter retention in a biological safety hood prepared by standard sterilization with UV and ethanol wiping. The filters used in this study were prepackaged sterilized polyethersulfone filter units (Millipore). Because of the need for other tests, the filtration volume for pre-stagnation was 40 liter per sample, and the volume of post-stagnation water samples were 80 mL for S1, 880 mL for S2, and 980 mL for S3 and S4 samples. Based on the flow cytometry data described in Results, this is estimated to retain 1.8 × 107–5.1 × 108 cells/ mL. Filters were removed from the filter unit with sterile scalpels and preserved in sterile falcon tubes −80 °C prior to DNA extraction. DNA was extracted using the Schmidt’s protocol [46] and purified with Promega Wizard DNA cleanup system (Promega) and was stored at −80 °C. The protocol was selected based on a previous publication that evaluated the different protocols for DNA extraction of drinking water samples from distribution systems [47].
Illumina sequencing
Sequencing analysis was conducted with the Illumina dual-index strategy. The extracted DNA was amplified with the following bacterial specific forward 515 F and 806 R (Read1-TATGGTAATTGTGTGCCAGCMGCCGCGGTAA; Read2-AGTCAGTCAGCCGGACTACHVGGGTWTCTAAT; Linker -ATTAGAWACCCBDGTAGTCCGGCTGACTGACT) [48]. The amplified products were purified with Promega PCR clean-up systems. Sequencing was performed at the W.M. Keck Center, part of the Roy J. Carver Biotechnology Center at the University of Illinois at Urbana-Champaign.
Data sharing
The sequencing data generated in this study have been deposited to NCBI SRA under PRJNA383368 (Biosamples: SAMN06763497-SAMN06763620).
Sequence processing
All sequences were curated with Mothur v1.33.3 [49]. The sequences from the dormitory experiment have been assembled into contigs assembled from Illumina pair-ended sequences targeting the V4 region (515 F/806 R). The water meter biofilm sequences were previously denoised sequences targeting V4–V5 region (515 F/907 R) and were trimmed (using pcr.seqs command in Mothur) to the same size as the dormitory samples. Following that, unique sequences were picked and aligned to SILVA Gold bacteria alignment [50]. Unique sequences were used for OTU picking at 97% similarity. Taxonomy classification was conducted using the SILVA online server. The sequences were subsampled to the lowest depth (1067 seqs/sample) to achieve an even sampling depth for diversity analysis.
Community differences between samples were calculated as Bray–Curtis dissimilarity. The similarity matrix was visualized by nMDS. A stress value below 0.2 was obtained, indicating a good 2D representation of the multidimensional matrix [51]. Multivariate tests were performed with distance-based methods analysis of similarities (ANOSIM), [33, 52]. These tests were performed in the PRIMER-6.0 package (PRIMER-E, Plymouth, UK) according to the authors’ manual [53]. Alpha Diversity analysis and additional beta-diversity analysis using various diversity matrices were conducted with R packages phyloseq (Version 1.23.1) [54] and vegan (Version 2.4–5) [55], and described in detail in Supplementary Materials.
Flow cytometry
The samples for flow cytometry were collected in gamma-irradiation sterilized falcon tubes (Fisher) and dechlorinated on site with filter-sterilized 1% sodium thiosulfate. The staining and flow cytometry measurement was conducted according to a published method for drinking water samples [56, 57]. The samples were transported to lab within 1 h of collection, stored in 4 °C refrigerator temporarily, and measured within 8 h after collection. Bacterial cells were stained with 10 µL/mL SYBR Green (Invitrogen) and incubated in dark at room temperature for 20 min. Flow cytometry was conducted with a Partec CyFlow space flow cytometer equipped with a 50 mW solid-state laser (488 nm). The fluorescence signals were collected with a green fluorescence channel FL1 = 536 mm and was also set as the trigger for the signals. Electronic gating for cell signals was performed with the Flowmax software (Partec). Cell signals were separated from noise by forward fluorescence signals and side scattering [58, 59].
Water chemistry measurement
Water samples were tested for time-sensitive parameters on site, including temperature and free chlorine. The free chlorine is measured using the N, N-diethyl-p-phynylenediamine (DPD) chlorine test kit (Hach, CO), a method approved by EPA for water utilities to monitor chlorine disinfectants. Other water chemistry were measured at the laboratory after 0.22-µm-filtration of the first liter. The measured parameters include inductively coupled plasma (ICP) metals, orthophosphate, anions (nitrate, nitrite, and sulfate), ammonia, non-volatile organic carbon (NVOC), and alkalinity. The chemical analyses were conducted at the Illinois State Water Survey in accordance with US EPA methods.
Numerical simulation of chlorine concentration and cell count
The dynamic change of chlorine concentration and cell count in pipes of one building during a period of seven days was simulated with COMSOL Multiphysics® software (Version 4.3a), using the transport of dilute species module. Details on the governing equations and parameters are provided in the Supplementary Materials. Briefly, because the process we studied was water stagnation without movement in a building distribution system, only diffusion was taken consideration into the mass transport (i.e., no convection). Disinfectant consumption was modeled as sodium hypochlorite reactions on pipe surfaces and bulk liquid [11, 60], and first-order reactions were assumed based on literature. Bulk reaction and wall reaction were modeled as Rc=−kbc and N0,c=−kwc, respectively, in which kb is the bulk reaction constant, kw is the wall reaction constants, and c represents sodium hypochlorite concentration.
Bacterial cell counts in the water phase were modeled as results from dispersal from biofilms, growth in bulk liquid, natural deaths, and chlorine inactivation [59,60,61,62]. The detachment of cells was modeled as a first-order process [60], and the detached cells were subjected to disinfectant inactivation that follows an exponential form [59]. Together the detachment–inactivation process is modeled as \({N}_{0,{\mathrm{X}}} = {n} \times {\mathrm{exp}}\left( {\frac{{ - {ac}}}{{1 + {bc}}}} \right)\), in which n represents detachment rate, a and b are inactivation kinetics constants, and c represents sodium hypochlorite concentration.
A few simplifications were involved in the model. The cell count was modeled as a monoculture rather than mixed-species to reflect the physical process. The growth substrate was modeled as one carbon species, dissolved methane, based on our previous study that showed methanotrophy as the primary production in the local distribution system [29]. The substrate was modeled as a constant supply.
Modeling migration and demographic stochasticity
The Sloan version of neutral model was used to explore the role of neutral processes in the assembly of indoor plumbing water communities and the contribution of plumbing-associated biofilms and city-supplied water to the in-plumbing water communities in [63]. We assumed the mainland communities to be stable (i.e., no speciation), because the experiment time (~6 days) was relatively short for the level of taxonomic resolution we were observing, i.e., OTUs defined based on 97% similarity of 16S rRNA genes.
This implementation was inspired by previous implementations of UNTB on human lung microbiome [64] and zebrafish gut microbiome [65]. Briefly, the post-stagnation water communities from each volume fraction were treated as the island communities, and the biofilm or city tap water communities as the mainland communities. For each taxon shared between the island and mainland communities, the expected frequency of detection in the local communities from dispersal and random growths/deaths (i.e., neutral processes) was computed from the abundance in the mainland communities, following a beta function [63]. Then, the overall fit of the model to the experimental data was assessed by comparing the residual sum of squares. The fitting was performed with a non-linear regression package, Minpack.lm, in R [65, 66]. Confidence intervals were calculated with the Hmisc package in R [67]. Link to the code was provided in Supplementary Materials.
References
National Research Council. Drinking water distribution systems: assessing and reducing risks. Washington D.C.: National Academies Press; 2006.
Proctor CR, Hammes F. Drinking water microbiology—from measurement to management. Curr Opin Biotechnol. 2015;33:87–94.
Szewzyk U, Szewzyk R, Manz W, Schleifer K-H. Microbiological safety of drinking water. Annu Rev Microbiol. 2000;54:81–127.
Centers for Disease Control and Prevention (CDC) Surveillance for waterborne disease outbreaks associated with drinking water and other nonrecreational water—United States, 2009–2010. MMWR. 2013;62:714–20..
Falkinham JO, Hilborn ED, Arduino MJ, Pruden A, Edwards MA, Edwards MA. Epidemiology and ecology of opportunistic premise plumbing pathogens: Legionella pneumophila, Mycobacterium avium, and Pseudomonas aeruginosa. Environ Health Perspect. 2015;123:749–58.
Naumova EN, Liss A, Jagai JS, Behlau I, Griffiths JK. (2016). Hospitalizations due to selected infections caused by opportunistic premise plumbing pathogens (OPPP) and reported drug resistance in the United States older adult population in 1991–2006. J Public Health Policy 37: 1–14.
US EPA O. National priorities: impacts of water conservation on water quality in premise plumbing and water distribution systems. https://www.epa.gov/research-grants/national-priorities-impacts-water-conservation-water-quality-premise-plumbing-and Accessed 19 Jul 2017).
Feazel LM, Baumgartner LK, Peterson KL, Frank DN, Harris JK, Pace NR. Opportunistic pathogens enriched in showerhead biofilms. Proc Natl Acad Sci. 2009;106:16393–9.
Wang H, Edwards M, Falkinham JO, Pruden A. Molecular survey of the occurrence of Legionella spp., Mycobacterium spp., Pseudomonas aeruginosa, and amoeba hosts in two chloraminated drinking water distribution systems. Appl Environ Microbiol. 2012;78:6285–94.
Ji P, Rhoads WJ, Edwards MA, Pruden A. Impact of water heater temperature setting and water use frequency on the building plumbing microbiome. ISME J. 2017;11:1318–30.
Biswas P, Lu C, Clark RM. A model for chlorine concentration decay in pipes. Water Res. 1993;27:1715–24.
Alcocer-Yamanaka VH, Tzatchkov VG, Arreguin-Cortes FI. Modeling of drinking water distribution networks using stochastic demand. Water Resour Manag. 2012;26:1779–92.
Hallam N, West J, Forster C, Simms J. The potential for biofilm growth in water distribution systems. Water Res. 2001;35:4063–71.
Zhou SL, McMahon TA, Walton A, Lewis J. Forecasting daily urban water demand: a case study of Melbourne. J Hydrol. 2000;236:153–64.
Ji P, Parks J, Edwards MA, Pruden A, Berry D, Xi C.et al. Impact of water chemistry, pipe material and stagnation on the building plumbing microbiome. PLoS ONE. 2015;10:e0141087.
Nguyen C, Elfland C, Edwards M. Impact of advanced water conservation features and new copper pipe on rapid chloramine decay and microbial regrowth. Water Res. 2012;46:611–21.
Rhoads WJ, Pruden A, Edwards MA. Survey of green building water systems reveals elevated water age and water quality concerns. Environ Sci Water Res Technol. 2016;2:164–73.
Lautenschlager K, Boon N, Wang Y, Egli T, Hammes F. Overnight stagnation of drinking water in household taps induces microbial growth and changes in community composition. Water Res. 2010;44:4868–77.
Hubbell SP. The unified neutral theory of biodiversity and biogeography (MPB-32). New Jersey; Princeton University Press: 2001.
Sloan WT, Woodcock S, Lunn M, Head IM, Curtis TP. Modeling taxa-abundance distributions in microbial communities using environmental sequence data. Microb Ecol. 2007;53:443–55.
Hwang C, Ling F, Andersen GL, LeChevallier MW, Liu W-T. Microbial community dynamics of an urban drinking water distribution system subjected to phases of chloramination and chlorination treatments. Appl Environ Microbiol. 2012;78:7856–65.
Lautenschlager K, Hwang C, Liu W-T, Boon N, Köster O, Vrouwenvelder H, et al. A microbiology-based multi-parametric approach towards assessing biological stability in drinking water distribution networks. Water Res. 2013;47:3015–25.
Soto-Giron MJ, Rodriguez-R LM, Luo C, Elk M, Ryu H, Hoelle J, et al. Biofilms on hospital shower hoses: characterization and implications for nosocomial infections. Appl Environ Microbiol. 2016;82:2872–83.
Vaz-Moreira I, Nunes OC, Manaia CM. Diversity and antibiotic resistance patterns of Sphingomonadaceae isolates from drinking water. Appl Environ Microbiol. 2011;77:5697–706.
Lindner AS, Pacheco A, Aldrich HC, Costello Staniec A, Uz I, Hodson DJ. Methylocystis hirsuta sp. nov., a novel methanotroph isolated from a groundwater aquifer. Int J Syst Evol Microbiol. 2007;57:1891–1900.
Belova SE, Kulichevskaya IS, Bodelier PLE, Dedysh SN. Methylocystis bryophila sp. nov., a facultatively methanotrophic bacterium from acidic Sphagnum peat, and emended description of the genus Methylocystis (ex Whittenbury et al. 1970) Bowman et al. 1993. Int J Syst Evol Microbiol. 2013;63:1096–104.
Flynn TM, Sanford RA, Ryu H, Bethke CM, Levine AD, Ashbolt NJ, et al. Functional microbial diversity explains groundwater chemistry in a pristine aquifer. BMC Microbiol. 2013;13:146.
Hackley KC, Panno SV, Anderson TF. Chemical and isotopic indicators of groundwater evolution in the basal sands of a buried bedrock valley in the midwestern United States: implications for recharge, rock-water interactions, and mixing. Geol Soc Am Bull. 2010;122:1047–66.
Ling F, Hwang C, LeChevallier MW, Andersen GL, Liu W-T. Core-satellite populations and seasonality of water meter biofilms in a metropolitan drinking water distribution system. ISME J. 2016;10:582–95.
Di Rienzi SC, Sharon I, Wrighton KC, Koren O, Hug LA, Thomas BC, et al. The human gut and groundwater harbor non-photosynthetic bacteria belonging to a new candidate phylum sibling to Cyanobacteria. Elife. 2013;2:e01102.
Stanish LF, Hull NM, Robertson CE, Harris JK, Stevens MJ, Spear JR, et al. Factors influencing bacterial diversity and community composition in municipal drinking waters in the Ohio river basin, USA. PLoS One. 2016;11:e0157966.
Hull NM, Holinger EP, Ross KA, Robertson CE, Harris JK, Stevens MJ, et al. Longitudinal and source-to-tap New Orleans, LA, U.S.A. drinking water microbiology. Environ Sci Technol. 2017;51:4220–9.
Chapman MG, Underwood AJ. Ecological patterns in multivariate assemblages: information and interpretation of negative values in ANOSIM tests. Mar Ecol Prog Ser. 1999;180:257–65.
Anderson MJ, Walsh DCI. PERMANOVA, ANOSIM, and the Mantel test in the face of heterogeneous dispersions: what null hypothesis are you testing? Ecol Monogr. 2013;83:557–74.
Boe-Hansen R, Albrechtsen H-J, Arvin E, Jørgensen C. Bulk water phase and biofilm growth in drinking water at low nutrient conditions. Water Res. 2002;36:4477–86.
Lehtola MJ, Miettinen IT, Keinänen MM, Kekki TK, Laine O, Hirvonen A, et al. Microbiology, chemistry and biofilm development in a pilot drinking water distribution system with copper and plastic pipes. Water Res. 2004;38:3769–79.
Norton CD, LeChevallier MW. A pilot study of bacteriological population changes through potable water treatment and distribution. Appl Environ Microbiol. 2000;66:268–76.
Rossman LA, Clark RM, Grayman WM. Modeling chlorine residuals in drinking-water distribution systems. J Environ Eng. 1994;120:803–20.
Allen MJ, Edberg SC, Reasoner DJ. Heterotrophic plate count bacteria—what is their significance in drinking water? Int J Food Microbiol. 2004;92:265–74.
Reasoner DJ, Blannon JC, Geldreich EE, Barnick J. Nonphotosynthetic pigmented bacteria in a potable water treatment and distribution system. Appl Environ Microbiol. 1989;55:912–21.
Sarin P, Snoeyink VL, Bebee J, Jim KK, Beckett MA, Kriven WM, et al. Iron release from corroded iron pipes in drinking water distribution systems: effect of dissolved oxygen. Water Res. 2004;38:1259–69.
Soda EA, Barskey AE, Shah PP, Schrag S, Whitney CG, Arduino MJ, et al. Vital signs: health care–associated legionnaires’ disease surveillance data from 20 states and a large metropolitan area—United States, 2015. MMWR. 2017;66:584–9.
Gu H, Lee SW, Buffington SL, Henderson JH, Ren D. On-demand removal of bacterial biofilms via shape memory activation. ACS Appl Mater Interfaces. 2016;8:21140–4.
Arciola CR, Campoccia D, Speziale P, Montanaro L, Costerton JW. Biofilm formation in Staphylococcus implant infections. A review of molecular mechanisms and implications for biofilm-resistant materials. Biomaterials. 2012;33:5967–82.
Cross DF, Benchimol A, Dimond EG. The faucet aerator–a source of pseudomonas infection. N Engl J Med. 1966;274:1430–1.
Schmidt TM, DeLong EF, Pace NR. Analysis of a marine picoplankton community by 16S rRNA gene cloning and sequencing. J Bacteriol. 1991;173:4371–8.
Hwang C, Ling F, Andersen GL, LeChevallier MW, Liu W-T. Evaluation of methods for the extraction of DNA from drinking water distribution system biofilms. Microbes Environ. 2012a;27:9–18.
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol. 2013;79:5112–20.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–6.
Clarke KR, R.M. W. Change in marine communities: an approach to statistical analysis and interpretation. 2nd ed. Plymouth: PRIMER-E; 2001.
Clarke KR. Non-parametric multivariate analyses of changes in community structure. Austral Ecol. 1993;18:117–43.
Anderson MJ, Gorley RN, Clarke KR. (2008). PERMANOVA+ for PRIMER: Guide to software and statistical methods. 2008 214.
McMurdie PJ, Holmes S. phyloseq:an R Package for reproducible interactive analysis and graphics of microbiome census data. PLoS One. 2013;8:e61217.
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, et al. vegan: community ecology package. R Package Version. 2017;2:4–5.
Hammes F, Berney M, Wang Y, Vital M, Köster O, Egli T. Flow-cytometric total bacterial cell counts as a descriptive microbiological parameter for drinking water treatment processes. Water Res. 2008;42:269–77.
Berney M, Hammes F, Bosshard F, Weilenmann H-U, Egli T. Assessment and interpretation of bacterial viability by using the LIVE/DEAD BacLight Kit in combination with flow cytometry. Appl Environ Microbiol. 2007;73:3283–90.
Arevalo J. (2007). Modeling free chlorine and chloramine decay in a pilot distribution system. Electron Theses Diss. http://stars.library.ucf.edu/etd/3066 Accessed 12 Feb 2017.
Albrich JM, Hurst JK. Oxidative inactivation of Escherichia coli by hypochlorous acid: rates and differentiation of respiratory from other reaction sites. FEBS Lett. 1982;144:157–61.
Bester E, Wolfaardt G, Aznaveh N, Greener J. Biofilms’ role in planktonic cell proliferation. Int J Mol Sci. 2013;14:21965–82.
Billen G, Servais P, Ventresque C, Bouillot P. Functioning of biological filters used in drinking-water treatment: the chabrol model. J Water Supply Res Technol AQUA. 1992;41:231–41.
Dukan S, Levi Y, Piriou P, Guyon F, Villon P. Dynamic modelling of bacterial growth in drinking water networks. Water Res. 1996;30:1991–2002.
Sloan WT, Lunn M, Woodcock S, Head IM, Nee S, Curtis TP. Quantifying the roles of immigration and chance in shaping prokaryote community structure. Environ Microbiol. 2006;8:732–40.
Venkataraman A, Bassis CM, Beck JM, Young VB, Curtis JL, Huffnagle GB, et al. Application of a neutral community model to assess structuring of the human lung microbiome. MBio. 2015;6:e02284–14.
Burns AR, Stephens WZ, Stagaman K, Wong S, Rawls JF, Guillemin K et al. (2015). Contribution of neutral processes to the assembly of gut microbial communities in the zebrafish over host development. ISME J. e-pub ahead of print, https://doi.org/10.1038/ismej.2015.142.
Elzhov T V., Mullen KM, Spiess A-N, Bolker B. (2013). Minpack.lm: R Interface to the Levenberg-Marquardt Nonlinear Least-Squares Algorithm Found in MINPACK, Plus Support for Bounds. https://cran.r-project.org/web/packages/minpack.lm/index.html Accessed 7 Jan 2016).
Harrell F. (2015). Hmisc: Harrell Miscellaneous. https://cran.r-project.org/web/packages/Hmisc/index.html (Accessed January 7, 2016).
Acknowledgements
We would like to thank staff at University Housing at UIUC for providing assistance in water sampling and construction history. We thank Dr. Vern Snoeyink for meaningful discussions. The authors declare no conflicts of interest. Fangqiong Ling is partially supported by Alfred P Sloan Foundation Microbiome of the Built Environment Postdoctoral Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ling, F., Whitaker, R., LeChevallier, M.W. et al. Drinking water microbiome assembly induced by water stagnation. ISME J 12, 1520–1531 (2018). https://doi.org/10.1038/s41396-018-0101-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0101-5
This article is cited by
-
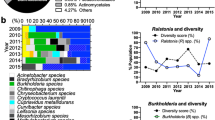
Responses of drinking water bulk and biofilm microbiota to elevated water age in bench-scale simulated distribution systems
npj Biofilms and Microbiomes (2024)
-
A revolutionary solution for on-the-go water disinfection
Nature Water (2024)
-
Walking-induced electrostatic charges enable in situ electroporated disinfection in portable water bottles
Nature Water (2024)
-
Occurrence of viable but non-culturable (VBNC) pathogenic bacteria in tap water of public places
Frontiers of Environmental Science & Engineering (2024)
-
Transition effects in an unchlorinated drinking water system following the introduction of partial reverse osmosis
Nature Water (2023)