Abstract
Symbiotic interactions between organisms create new ecological niches. For example, many insects survive on plant-sap with the aid of maternally transmitted bacterial symbionts that provision essential nutrients lacking in this diet. Symbiotic partners often enter a long-term relationship in which the co-evolutionary fate of lineages is interdependent. Obligate symbionts that are strictly maternally transmitted experience genetic drift and genome degradation, compromising symbiont function and reducing host fitness unless hosts can compensate for these deficits. One evolutionary solution is the acquisition of a novel symbiont with a functionally intact genome. Whereas almost all aphids host the anciently acquired bacterial endosymbiont Buchnera aphidicola (Gammaproteobacteria), Geopemphigus species have lost Buchnera and instead contain a maternally transmitted symbiont closely related to several known insect symbionts from the bacterial phylum Bacteroidetes. A complete genome sequence shows the symbiont has lost many ancestral genes, resulting in a genome size intermediate between that of free-living and symbiotic Bacteroidetes. The Geopemphigus symbiont retains biosynthetic pathways for amino acids and vitamins, as in Buchnera and other insect symbionts. This case of evolutionary replacement of Buchnera provides an opportunity to further understand the evolution and functional genomics of symbiosis.
Similar content being viewed by others
Introduction
Interaction between microbes and eukaryotes allows lineages to access new ecological niches and can promote biodiversity. Some of these symbiotic interactions evolve into obligate relationships in which neither lineage is able to persist without the other. Prominent examples of obligate symbiosis are found in sap-feeding insects that harbor bacterial endosymbionts, which synthesize and provide hosts essential nutrients that are limited in their diet of plant-sap [1, 2]. In exchange, bacterial symbionts are provided with a stable environment within specialized host cells called bacteriocytes, where synthesized nutrients are exchanged with the host [3]. Hosts have also developed mechanisms to vertically transmit obligate symbionts to offspring, thus ensuring transmission and persistence of symbionts [4, 5].
Over evolutionary time, the very features that have strengthened obligate symbiotic relationships may also present an evolutionary hazard. The small effective population size of vertically transmitted obligate symbionts promotes the accumulation of deleterious mutations resulting in reduced gene function and significant gene loss [6, 7]. As endosymbiont genomes continue to deteriorate, functions may become significantly compromised, resulting in reduced host fitness unless hosts can adapt and compensate for these deficits [8]. Despite showing signs of genomic degradation, obligate symbioses can be extremely stable, reflecting the host’s adaptations for regulating, maintaining, and transmitting their endosymbionts. One solution to the evolutionary degradation of obligate symbiont functions is the acquisition of novel symbionts with intact genomes and functional capabilities. In several lineages of sap-feeding insects, ancestral symbionts have been replaced or joined by a second obligate symbiont that also contributes to nutrient biosynthesis [8,9,10,11,12].
Often, host lineages have maintained the same bacterial associate over long evolutionary timescales without replacement. For example, aphids and their primary bacterial endosymbiont, Buchnera aphidicola (Gammaproteobacteria) have codiversified for 200 million years [13]. Buchnera genome sequences for several divergent aphid species confirm the conserved gene content and functional role of this obligate symbiont. Buchnera genomes are highly reduced, ranging in size from 416 to 644 kilobases (kb), and most ancestral genes have been lost [14,15,16]. Preserved genes include those underlying the synthesis and transport of essential amino acids (EAA) required by aphids. Buchnera’s role in amino acid provisioning has been elucidated through genome sequencing of both the symbiont and host genome of Acyrthosiphon pisum (pea aphid) [17, 18]. For Buchnera genes that are retained, elevated rates of mutation accumulation have resulted in lower thermal stability of encoded proteins relative to homologous proteins in other bacteria [19].
Almost all aphids have Buchnera [20]; known exceptions are limited to a few members of the Asian tribe Cerataphidini that instead harbor an extracellular fungal symbiont [21,22,23]. We have found a second case of Buchnera loss: members of the aphid genus Geopemphigus have lost their ancient Buchnera symbiont and have replaced it with a symbiont from the bacterial phylum Bacteroidetes. In this study, we sequence a representative genome of this novel symbiont and reconstruct metabolic pathways. Phylogenetic analysis of several Geopemphigus species and their symbionts suggests that (a) the replacement event occurred in the ancestor of Geopemphigus aphids and (b) the new symbiont is closely related to Bacteroidetes endosymbionts found in some other insects. We find that several aphid genes hypothesized as important in the Buchnera symbiosis are retained in Geopemphigus genomes, suggesting that it may utilize ancestral genetic tools for interactions with its relatively novel symbiotic partner. Our results show that Geopemphigus aphids have lost Buchnera and replaced it with a new bacterial symbiont that has undergone convergent evolution with Buchnera, serving as a functional substitute for the lost ancient symbiont.
Materials and methods
Biological samples
We obtained live samples of Geopemphigus species living on Pistacia texana in Langtry, TX, USA, as well as preserved samples from this locality and from Pistacia mexicana in localities in Mexico (Table S1). These samples represent four Geopemphigus species, including undescribed species. Geopemphigus species are confined to New World Pistacia that occurs mostly in Mexico [24, 25]; thus, our samples include a substantial representation of diversity in this aphid clade.
Lack of Buchnera in Geopemphigus species
To verify the absence of Buchnera in Geopemphigus aphids, we amplified a region of 16S rRNA sequence using a pair of universal bacterial primers (10 F/1057 R) and a pair targeted to amplify only Buchnera sequence (10 F/757 R). We PCR-screened multiple species of Geopemphigus aphids (Table S1) as well as Acyrthosiphon pisum samples, containing Buchnera, using the two primer sets to determine bacterial DNA template quality and presence of Buchnera, respectively. DNA template quality was also determined based on PCR amplification of a region of host mitochondrial COI (primers LepF/LepR) [26]. Samples with successful amplification of the universal 16S rRNA region and mitochondrial sequence and failure to amplify the Buchnera-specific 16S rRNA region were considered to potentially lack the ancient endosymbiont of aphids.
Illumina sequencing to detect symbiont genomes
Galls formed on Pistacia texana containing Geopemphigus sp. individuals were collected from Langtry, TX, USA (Table S1). Aphids from a single gall were collected and pooled together for DNA extraction using a Qiagen DNeasy Kit. Since each gall is initiated by a single parthenogenetic aphid, aphids and symbionts within a gall are expected to be clonal. An Illumina sequencing library was prepared for the isolated genomic DNA and sequenced using Illumina HiSeqV4000 PE150 to generate 4.4 GB of shotgun metagenomic sequence data. Using CLC Genomics workbench v9, adapter-trimmed paired-reads were quality-filtered and assembled to produce 27,428 contigs (PRJNA414766). Contigs representing symbiont genome sequences were identified based on relative coverage, as obligate symbiont genomes typically occur at higher copy number than host genomes. Assembled contigs were used to generate a BLAST database for nucleotide and protein searches (blastn and blastx, respectively) against several reference bacterial genomes (Table S2) to identify additional bacterial reads. A blastn search using Buchnera genome sequences was also performed to detect this symbiont.
Genome annotation and metabolic reconstruction
The assembled symbiont genome was annotated using the RAST Prokaryotic Genome Annotation Server (http://rast.nmpdr.org/). Annotations of transfer RNAs (tRNAs) were verified using tRNAscan-SE [27,27,]. Metabolic pathways were reconstructed using RAST annotations and verified based on Ecocyc and KEGG annotation [28,29,30]. Inferred metabolic functions were compared to several reference genomes representing free-living bacteria and bacterial endosymbionts of insects (Table S2).
Molecular phylogeny
To determine the potential origins of this symbiotic relationship, we collected sequence data for additional Geopemphigus samples (Table S1) to generate a bacterial phylogeny. We used Bacteroidetes-specific primers [31] to sequence 1399 bp of 16S rRNA for each Geopemphigus sample. Generated sequences were aligned with 16S rRNA sequences from several bacterial species (Table S2) using MUSCLE 3.8.31 and verified manually. Bacterial 16S rRNA phylogeny was inferred with maximum likelihood using RAxML v7.2.8 with 1000 bootstrap replicates.
Aphid host genes for symbiosis
We searched all assembled contigs for several aphid host genes that are upregulated in bacteriocytes and are suggested to play an important role in symbiosis between Buchnera and aphids. Specifically, we performed a blastx to search for genes previously identified as transferred from bacterial genomes to the aphid genome as well as for novel aphid-specific orphan genes encoding secretion proteins expressed within bacteriocytes [32,33,34]. Proteins encoded by identified laterally transferred genes were aligned with sequences from aphid genomes [17, 35,36,37] and with homologs from bacterial species available in GenBank. Subsequently, we performed maximum likelihood phylogenetic analysis for each protein alignment (as above) to determine the evolutionary relationships of any laterally transferred or novel aphid genes in Geopemphigus aphids.
Fluorescence in situ hybridization microscopy
Geopemphigus specimens were also collected and preserved for whole-mount FISH to localize the novel endosymbiont within the host. Samples were prepared and hybridized as previously described [31, 38]. Briefly, the samples were fixed with Carnoy’s solution, bleached in 6% H202–80% EtOH, and hybridized with fluorescent probes CFB319 and PRO319, which target Bacteroidetes and Proteobacteria 16S rRNA, respectively. Samples were treated with hydrogen peroxide (H2O2) to reduce autofluorescence of insect tissues [38]. Host DNA was counterstained using DAPI. Images were visualized and captured using a Nikon Eclipse TE2000-U epifluorescence microscope.
Results
Absence of Buchnera and detection of novel Bacteroidetes symbiont
In a previous study aimed at using Buchnera gene sequences to determine phylogenetic relationships of 70 aphid species representing most major aphid lineages, the only sample that failed to amplify for Buchnera genes was a Geopemphigus species [20]. However, amplification failure alone is a weak evidence for absence of Buchnera, since it could result from sequence changes at primer binding sites. Because Buchnera genomes typically outnumber aphid genomes by >20 times within total genomic DNA samples, deep shotgun sequencing gives clear evidence of presence or absence of this symbiont. Illumina shotgun sequencing of total DNA from two Geopemphigus species yielded no reads that corresponded to Buchnera sequences, confirming its absence from those samples. Shotgun sequencing from Geopemphigus sp. S2 generated three large (1053 kb, 186 kb, and 69 kb) and two small contigs (4874 bp and 3624 bp), all identified as bacterial genome sequences from the phylum Bacteroidetes (PRJNA414766). Average sequence coverage of the three large contigs was ×236, while the two small contigs occurred at ×505 (for 4874 bp contig) and ×272 (for 3624 bp contig) average sequence coverage.
Using primers designed to selectively detect Bacteroidetes, the novel symbiont was also found to be present in other Geopemphigus species from additional localities (Table S1). The detection of this symbiont based on 16S rRNA sequences for multiple species of Geopemphigus suggests that this symbiont was acquired prior to the diversification of this aphid group. To determine whether Buchnera was also absent from other Geopemphigus samples, we attempted to amplify Buchnera 16S rRNA genes using primers to conserved sites. PCR amplification of an insect mitochondrial gene was used to verify template quality. All Geopemphigus DNA samples gave positive amplification of the insect mitochondrial gene and Bacteroidetes 16S rRNA gene, but failed to amplify Buchnera sequences, supporting the absence of Buchnera.
Genome structure of the novel Bacteroidetes symbiont
To determine the genome of the Bacteroidetes symbiont from Geopemphigus sp. S2, we used Illumina sequencing of DNA from whole aphid bodies, combined with additional molecular and bioinformatic approaches. We assembled a complete genome for the symbiont. This bacterial species is closely related to other insect endosymbionts from Flavobacteriaceae (Bacterioidetes) and has a reduced genome compared to genomes of free-living Bacteroidetes (Fig. 1; Table S2). The symbiont has a 1.32 Mb genome with a GC-content of 37.0% and encodes 1611 protein coding genes, 771 of which were classified as encoding hypothetical proteins (Table S2). Additionally, the symbiont retains three rRNAs, corresponding to a single copy each of the 5S, 16S, and 23 S rRNAs, and 35 tRNAs, which is near the minimal number needed for translating all codons.
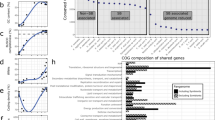
Comparison of genome size and G + C content for several species of Proteobacteria (circles) and Bacteroidetes (triangles) representing free-living (black), facultative (gray), and obligate (white) insect endosymbiont bacteria, which shows that the novel Geopemphigus symbiont “Candidatus Skilesia alterna” is more similar to other obligate endosymbiont genomes with reduced genome size and lower G + C% content
Metabolic functions of the novel Bacteroidetes symbiont
To determine the functional role of the Bacteroidetes symbiont, we reconstructed metabolic pathways based on genome annotations. Analysis of amino acid biosynthetic pathways revealed that the novel Bacteroidetes symbiont encodes nearly all of the enzymes needed to synthesize the ten EAAs (Fig. 2). The novel symbiont retains ilvE, the gene encoding the branched-chain amino acid aminotransferase that completes that final step of the pathway for biosynthesis of the branched-chain amino acids (valine, leucine, and isoleucine). Buchnera lacks this terminal transaminase, though the aphid host possesses a homolog of the gene that is upregulated in bacteriocytes [3]. The novel symbiont appears to be unable to synthesize methionine de novo, but maintains the ability to import this amino acid. Additionally, the novel symbiont still encodes genes for the synthesis and metabolism of most non-EAAs (Fig. 2). The genes encoding alanyl-tRNA synthetase and four hypothetical proteins are encoded on the smallest (3624 bp) contig.
A large fraction of genes related to biotin, folate, and riboflavin biosynthesis remain in the genome. Genes that encode copper efflux pumps (csoR/copZA) as well as magnesium and cobalt efflux pumps (corC) are retained in the symbiont and may be used to avoid levels toxic to the cell. A subset of genes that encode the twin-arginine translocation system (tatAC) and bacterial-signal-recognition particle and receptor (ftsY and ffh) are also present. Genes for several outer membrane proteins were also identified.
In this symbiont, the genes encoding glycolysis seem operational while gluconeogenesis may either be nonfunctional or catalyzed by an unknown enzyme. The symbiont maintains a minimal pentose phosphate pathway in which only the nonoxidative branch of the pathway is intact. Genes encoding the tricarboxylic acid cycle (TCA) cycle are all intact. The genes sucA and sucB are located on the 4874 bp contig, which occurs at twice the sequencing depth (×505 average coverage) compared to the large symbiont contigs, indicating that this contig represents a duplicated region or a plasmid.
The bacterial symbiont retains genes for DNA replication, transcription, and translation. Genes for DNA polymerase subunits (holA, dnaE, dnaN, dnaQ, and dnaX) are retained in the genome. In terms of transcription and translation function, one copy of each of the three ribosomal RNAs is retained, along with 28 large-subunit ribosomal proteins and all 21 small-subunit ribosomal proteins, as well as translation initiation and elongation factors. Genes underlying transcription include rpoA, rpoB, rpoC, and rpoD. Similar to other symbionts with small genomes, heat-shock proteins involved in protein folding and stability (groEL, groES, dnaJ, dnaK, grpE, htpG) are retained in the genome. Several genes for DNA recombination and repair are still present in the genome including genes encoding DNA polymerase III and recombination repair (recABC and uvrD). The symbiont has also recBCD, genes involved in homologous recombination that are maintained in all sequenced Buchnera. DNA mismatch repair genes mutL and mutS are present, but not mutH, which is also absent from other related Bacteroidetes endosymbionts.
The Bacteroidetes symbiont maintains several genes for cell wall production and other genes involved in cell structure and division. Specifically, the symbiont has lost the ability to synthesize phospholipids, but retains genes responsible for the production of lipid A, fatty acids, and peptidoglycan. Maintained genes that have been shown to be involved in peptidoglycan recycling include pepD and ddl as well as ldcA and amiD [39], the latter two which have previously identified as bacterial genes that have been horizontally transferred to the aphid host genome and are highly expressed in aphid bacteriocytes [32, 33]. Additionally, the presence of rod-shape determining genes mreBCD and rodA suggests that the symbiont may be rod-shaped. We observe loss of genes related to cell motility and stress response, similar to other endosymbionts surviving in the stable nutrient-rich environment of a host cell.
Evolutionary origins of the novel Bacteroidetes symbiont
To infer the evolutionary relationships of this symbiosis, we sequenced symbiont 16S rRNA for several additional Geopemphigus species (Table S1). While Geopemphigus samples failed to amplify using Buchnera-specific primers, PCR using Bacteroidetes-specific primers resulted in amplification of bacterial 16S rRNA sequences with greater than 99% identity to sequences retrieved from deep shotgun sequencing of the original samples. Using an alignment of 16S rRNA sequences from both free-living bacteria and insect-bacterial endosymbionts, we inferred a maximum likelihood phylogeny to determine the evolutionary origins of the novel symbiont. The symbiont is nested within a clade of bacterial endosymbionts in the phylum Bacteroidetes and is highly divergent from Buchnera aphidicola, which belongs to the phylum Proteobacteria (Fig. 3). Analyses suggest that the closest relatives (with complete genomes publicly available) of the new symbiont are Uzinura diaspidicola (CP003263) and Walczuchella monophlebidarum (CP006873), which are both obligate endosymbionts of scale insects [40, 41].
Evolutionary relationships of the novel Geopemphigus symbiont within a clade of bacterial endosymbionts in the phylum Bacteroidetes. Buchnera aphidicola, the symbiont of most aphids, belongs to the phylum Proteobacteria. Phylogeny is constructed using maximum likelihood analysis of 1399 bp of 16S rRNA
Aphid host genes for symbiosis
Several bacterial genes have been hypothesized to play roles in the Buchnera-aphid symbiosis, based on their inferred functions and their high expression in bacteriocytes. These include several genes acquired through horizontal gene transfer from bacteria to aphid ancestors [32, 33]. Our sequencing data included sequences from the host aphid genome with an estimated average coverage of ×22, and we searched the contigs for homologs of A. pisum genes involved in symbiosis with Buchnera. BLASTp searches of all assembled contigs identified homologs for several of these A. pisum genes including amiD, bLys, and rlpA, though not ldcA. Genes ldcA and amiD are putatively involved in peptidoglycan recycling [39], while bLys is functions in cell wall degradation.
The gene amiD encodes an outer membrane lipoprotein, and has been found in A. pisum (ACYPI006531) and other in aphid genomes. The second exon of amiD shares 60% protein sequence similarity (length 193 aa) with a protein encoded in the Geopemphigus genome, and phylogenetic analysis places the Geopemphigus copy with copies from other aphid species (Figure S1A). bLys, a chimeric protein consisting of a fusion of a eukaryotic peptidase and a bacterial lysozyme, is also upregulated in A. pisum bacteriocytes [33] and has a homolog in Geopemphigus that clusters with aphid copies in phylogenetic analyses (Figure S1B). The A. pisum genome contains five homologs of the gene rlpA, encoding a rare lipoprotein; these copies represent duplications of a gene originating through lateral transfer from bacteria [32, 33]. Using A. pisum copies as queries, we found two related genes in the Geopemphigus genome (Figure S1C).
A. pisum also possesses aphid-specific genes, BCR1-8 and SP1-4, 5a, 6, which encode secretion proteins localized to and highly expressed in bacteriocytes. BLASTp searches using these A. pisum proteins as queries retrieved significant hits for only SP2 and SP4, which seem to remain intact in the Geopemphigus sp. S2 genome. Amino acid sequence alignments of SP2 (181 aa) and SP4 (465 aa) between A. pisum and Geopemphigus sp. S2 show 33.7% and 62.8% sequence similarity, respectively; both cluster with A. pisum proteins in phylogenetic analyses (Figure S1D). Thus, the novel Bacteroidetes symbiont retains numerous characteristic aphid genes that are hypothesized to play a role in the symbiosis between aphids and Buchnera and that are specifically expressed in bacteriocytes in A. pisum.
Localization of the novel Bacteroidetes symbiont
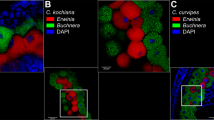
The presence of the Bacteroidetes symbiont in Geopemphigus sp. S2 was confirmed using FISH microscopy. Bacteroidetes symbionts are present both within maternal tissues (Fig. 4) and also within developing embryos, in which they are undergoing different stages of symbiont packaging (Fig. 4f, g, dashed arrows). The arrangements of the Bacteroidetes symbionts within apparent bacteriocytes and embryos resemble the Buchnera packaging observed in A. pisum [5]. We also see what appear to be large maternal bacteriocytes surrounded by the embryos (Fig. 4f, g, solid arrows).
Images of a gall formed by Geopemphigus sp. S2 on Pistacia texana and b Geopemphigus sp. S2 feeding within the gall. c–e Detection of endosymbionts in Geopemphigus sp. S2 using fluorescent in situ hybridization (FISH) microscopy of a whole-mount insect (ventral view with head at left). c DAPI staining for DNA (blue), mostly present in insect cell nuclei, d FISH for Buchnera (Proteobacteria) (green) showing absence from abdomen (insect cuticle gave some autofluorescence), e FISH for Bacteroidetes (red) showing presence of bacteria within embryos in mother’s abdomen (some autofluorescence of insect cuticle), and f a merged image from c–e. g FISH image of a second whole-mount Geopemphigus sp. S2 female (dorso-lateral view with head at lower right corner). Merged FISH images show localization of the novel symbiont within developing embryos (dashed arrow) and predicted maternal bacteriocytes (solid arrow). White scale bar is 200 μm
Proposed name of the novel Bacteroidetes symbiont: “Candidatus Skilesia alterna”
The novel symbiont in Geopemphigus aphids is evolutionarily distinct from any known bacterial species, with a 16S rRNA gene 96.8% identical to an unnamed Bacteroidetes bacterium from the scale insect Cryptococcus ulmi (DQ133549) and 91.6% identical to 16S rRNA sequence from the bacterial endosymbiont of Llaveia axin axin (Bacteroidetes: “Candidatus Walczuchella monophlebidarum”; CP006873). We propose the name “Candidatus Skilesia alterna” for the novel endosymbiont found in Geopemphigus aphids. The name Skilesia is in honor of Jack Raymond Skiles, for his efforts in research, education, and preservation of the biodiversity of the Trans-Pecos region of West Texas, where Geopemphigus samples were collected. In the 1970s, Skiles established the Judge Roy Bean Visitor Center and Botanical Gardens in Langtry, Texas, where he developed a living collection of local plants, including Pistacia texana. He and his family continue efforts to preserve the natural history and biodiversity of West Texas. We propose the species name alterna, meaning one after another, to describe the replacement of Buchnera with the new symbiont. Distinctive features of this organism include its restriction to living in Geopemphigus aphids and the following distinct sequences in the 16S rRNA genes (numbering corresponding to Escherichia coli (strain K12) standard numbering, 5’ to 3’): 5’-GGCTCCTGG-3’ at positions 261 to 269, 5’-GGCATAACC-3’ at 398 to 406, 5’-ACTTATTCGTGGCACA-3’ at 529–542, 5’-TCGTGT-3’ at 992 to 997, and 5’-GGGCGCACG-3’ at 1298 to 1306.
Discussion
In this study, we report a novel symbiont in Geopemphigus aphids, provisionally named “Ca. Skilesia alterna”, and provide evidence that this novel symbiont has functionally and physically replaced Buchnera, the ancient symbiont present in almost all aphids. Over evolutionary time, obligate, vertically transmitted symbiont genomes lose genes and shrink [15]. If symbiont capabilities erode to the point where they no longer provide functions needed by hosts, the hosts may acquire additional symbionts or even completely replace ancient symbionts with new symbionts [10, 12, 42]. Because Geopemphigus is nested within the clade of aphids known to harbor Buchnera, we can reject the hypothesis that Geopemphigus represents an ancient lineage that diverged from other aphids prior to the acquisition of Buchnera.
Like Baizongia species, Geopemphigus species are in the tribe Fordini and form galls on host plants of the genus Pistacia (Fig. 4a), but Geopemphigus occurs on New World Pistacia trees, whereas Baizongia occurs on Old World Pistacia trees. Based on phylogenetic trees inferred for both aphid hosts and bacterial symbionts, the novel Bacteroidetes symbiont was acquired after the split of Geopemphigus from Baizongia pistaciae, which lives on Pistacia lentiscus, but before the diversification of Geopemphigus species, which live on the New World species, P. texana and P. mexicana [24, 25]. These Pistacia-dwelling aphids likely diverged along with their host plants following geographic isolation, with each aphid forming species-specific galls [43]. Pistacia is at least 50 million years old, and the texana-mexicana lineage forms an early branch in the Pistacia phylogeny [44,45,46]. Thus, the new Bacteroidetes symbiont could have colonized its host as long ago as 50 million years.
This novel symbiont has undergone substantial gene loss and genome size reduction as compared to related free-living bacteria (Fig. 1; Table S2). Gene loss is a convergent feature of heritable obligate symbionts [1, 47, 48]. Comparisons of genome size, gene content, and GC content suggest that the Bacteroidetes symbiont is in an intermediate stage of genome degradation along the irreversible path towards obligate symbiosis (Fig. 1, Table S2). Based on metabolic pathway reconstruction, we infer that a primary function of this symbiont is EAA biosynthesis, indicating functional convergence with the now absent Buchnera symbiont (Fig. 2). Based on FISH microscopy, “Ca. Skilesia alterna” is retained within host-derived bacteriocytes and embryos are inoculated during development; these features parallel the colonization and transmission of Buchnera in A. pisum, a model for obligate symbiosis [4, 5].
The novel symbiont in Geopemphigus falls within a clade consisting of obligate insect symbionts, in hosts ranging from cockroaches, to leafhoppers, spittlebugs, and cicadas to scale insects (Fig. 3); however, the novel symbiont has a genome with less extreme reduction and AT-bias (Fig. 1). One hypothesis for this distribution of this bacterial symbiont clade among hosts is that it has somehow been horizontally transferred among diverse insect groups, although highly specialized obligate symbionts typically are believed not to be able to transfer into a divergent host species. An alternative hypothesis for this distribution is that there are environmental bacteria not yet characterized that serve as a source for repeated infections of insects, with some evolving to become obligate symbionts. This second scenario resembles that supported for Sodalis species, which are symbiotic in a wide array of insect host groups [49], a range that appears to result from repeated infections by an ancestral species resembling Sodalis precaptaevis, which was isolated from an opportunistic wound from a tree branch [50] and appears to have preadaptations for becoming symbiotic in insect hosts [51]. Under this latter scenario, the Geopemphigus symbiont may have a more recent origin, explaining its less reduced genome.
Previously, the only aphids known to lack Buchnera were several species within the tribe Cerataphidini (subfamily Hormaphidinae) [21, 22, 52], a group not closely related to Geopemphigus (subfamily Fordinae) [20, 53]. For example, some species of Cerataphis and Tuberaphis lack Buchnera and instead contain an extracellular fungal symbiont that lives in the body cavity [1, 21, 22, 54]. Interestingly, Cerataphidini and Geopemphigus both inhabit warm regions, unlike most aphid species including most Hormaphidinae, which live in temperate regions with cold winters. Since Buchnera, like many obligate heritable symbionts, is sensitive to high temperatures [55, 56], it is possible that aphids living in hot climates are more prone to acquire novel symbionts more tolerant of heat [53].
Whether Geopemphigus bacteriocytes are homologous to those housing Buchnera in other aphids is not known, and not resolved by our FISH microscopy (Fig. 4). A species of Tuberaphis that lacks Buchnera was shown to maintain cells homologous to the bacteriocytes of other aphids [5], showing that these cell types do not require colonization by Buchnera. The secondary symbiont, Serratia symbiotica, can replace Buchnera within A. pisum following experimental elimination of Buchnera; S. symbiotica colonizes the cytoplasm of bacteriocytes where it can fulfill the essential role of Buchnera [57]. Likewise, multiple symbiont types can colonize bacteriocytes of some adelgids [58]. But, novel co-obligate symbionts are sometimes restricted to novel types of bacteriocytes distinct from those harboring the more ancient partners [10, 59]. Interestingly, based on sequence data from the host aphid genome, Geopemphigus retains a set of genes homologous to A. pisum genes that are predominantly or solely expressed in bacteriocytes, that were acquired by an ancestral aphid from bacterial sources, and that are hypothesized to be critical in the interaction with Buchnera. Potentially these genes have been co-opted in the novel symbiosis with the Bacteroidetes symbiont; information on where these are expressed in Geopemphigus would be useful for understanding whether this novel symbiosis exploits genetic and cellular machinery that evolved for the Buchnera symbiosis.
Buchnera has been surveyed in a diversity of aphids including several other aphid species within the subfamily Fordinae [20]. Of these, the closest relative to Geopemphigus is B. pistaciae. Based on its sequenced genome [19], Buchnera of B. pistaciae is unable to synthesize arginine, cysteine, folate, riboflavin, pantothenate, pyrimidines, and heme protein, whereas all of these compounds can be produced by most Buchnera for which genomes are currently available. Buchnera of B. pistaciae also lacks numerous genes involved in production of outer membrane structures. Thus, the Buchnera of Fordini species may have undergone relatively severe genome degradation and loss of genes useful to hosts, potentially leading to the acquisition of a novel symbiont and loss of Buchnera in the Geopemphigus lineage.
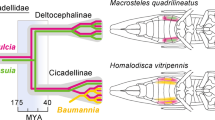
Several examples exist in which insect hosts establish and maintain two endosymbionts that appear to be co-obligate; these may represent intermediate stages of endosymbiont replacement. Gradual gene loss in an ancient symbiont may drive the eventual fixation of secondary symbionts to complement loss of function. Numerous cases have been documented for complementary roles of co-obligate symbionts in aphids [59, 60]. In contrast to these cases, the symbiont found in Geopemphigus aphids represents a case in which a new functional symbiont became established without persistence of the ancient partner. In adelgids, which are relatives of aphids feeding on sap of conifers, each host species contains two symbiont types, but a phylogenetic analysis shows a succession, in which a more ancient symbiont is replaced by a more recent one, which in turn is replaced by yet a more recent symbiont [10, 58, 61, 62]. Symbiont succession shows a somewhat different pattern in the sap-feeding Auchenorrhyncha (which includes cicadas, spittlebugs, leafhoppers, treehoppers, and planthoppers). Two ancestral bacterial symbionts have been retained in some auchenorrhynchan groups for 280 million years, but one of these (from Betaproteobacteria) is more subject to genomic degradation than is its partner (Sulcia muelleri from Bacteroidetes) and is more often replaced in particular host lineages [8, 10]. Similar succession of symbiont types has been documented for scale insects [63] and for weevils [62]. While these replacements appear to have happened more frequently in some groups (e.g., adelgids) than in others (Auchenorrhyncha and aphids), they are nonetheless rare events in all of these cases, happening on times scales of millions of years.
Our results show that Geopemphigus aphids have lost the ancient symbiont Buchnera and have replaced it with a new symbiont from the phylum Bacteroidetes. This novel symbiont, “Candidatus Skilesia alterna” has a reduced genome, but retains genes enabling it to perform the functional role previously fulfilled by Buchnera. This case of evolutionary replacement of an obligate symbiont provides an opportunity to further understand evolution and functional genomics of symbiosis.
References
Moran NA, McCutcheon JP, Nakabachi A. Genomics and evolution of heritable bacterial symbionts. Annu Rev Genet. 2008;42:165–90.
Douglas AE. Nutritional interactions in insect-microbial symbioses: aphids and their symbiotic bacteria Buchnera. Annu Rev Entomol. 1998;43:17–37.
Hansen AK, Moran NA. Aphid genome expression reveals host-symbiont cooperation in the production of amino acids. Proc Natl Acad Sci USA. 2011;108:2849–54.
Koga R, Meng X, Tsuchida T, Fukami T. Cellular mechanism for selective vertical transmission of an obligate insect symbiont at the bacteriocyte–embryo interface. Proc Natl Acad Sci USA. 2012;109:E1230–37.
Braendle C, Miura T, Bickel R, Shingleton AW, Kambhampati S, Stern DL. Developmental origin and evolution of bacteriocytes in the aphid-Buchnera symbiosis. PLoS Biol. 2003;1:70–76.
McCutcheon JP, Moran NA. Extreme genome reduction in symbiotic bacteria. Nat Rev Microbiol. 2012;10:13–26.
Moran NA. Accelerated evolution and Muller’s rachet in endosymbiotic bacteria. Proc Natl Acad Sci USA. 1996;93:2873–78.
Bennett GM, Moran NA. Heritable symbiosis: The advantages and perils of an evolutionary rabbit hole. Proc Natl Acad Sci USA. 2015;112:10169–76.
Gosalbes MJ, Lamelas A, Moya A, Latorre A. The striking case of tryptophan provision in the cedar aphid Cinara cedri. J Bacteriol. 2008;190:6026–29.
Koga R, Bennett GM, Cryan JR, Moran NA. Evolutionary replacement of obligate symbionts in an ancient and diverse insect lineage. Environ Microbiol. 2013;15:2073–81.
McCutcheon JP, McDonald BR, Moran NA. Convergent evolution of metabolic roles in bacterial co-symbionts of insects. Proc Natl Acad Sci USA. 2009;106:15394–99.
Sudakaran S, Kost C, Kaltenpoth M. Symbiont acquisition and replacement as a source of ecological innovation. Trends Microbiol. 2017;25:375–90.
Moran NA, Munson MA, Baumann P, Ishikawa H. A molecular clock in endosymbiotic bacteria is calibrated using the insect host. Proc Biol Sci. 1993;253:167–71.
Jiang Z, Jones DH, Shuir S, Tsinoremas N, Wyss T, Jander G, et al. Comparative analysis of genome sequences from four strains of the Buchnera aphidicola Mp endosymbion of the green peach aphid, Myzus persicae. BMC Genomics. 2013;14:1–11.
Moran NA, Bennett GM. The tiniest tiny genomes. Annu Rev Microbiol. 2014;68:195–15.
Perez-Brocal V, Gil R, Ramos S, Lamelas A, Postigo M, Michelena JM, et al. A small microbial genome: the end of a long symbiotic relationship? Science. 2006;314:312–13.
International Aphid Genomics Consortium. Genome sequence of the pea aphid Acyrthosiphon pisum. PLoS Biol. 2010;8:e1000313.
Shigenobu S, Watanabe H, Hattori M, Sakaki Y, Ishikawa H. Genome sequence of the endocellular bacterial symbiont of aphids Buchnera sp APS. Nature. 2000;407:81–6.
van Ham RCHJ, Kamerbeek J, Palacios C, Rausell C, Abascal F, Bastolla U, et al. Reductive genome evolution in Buchnera aphidicola. Proc Natl Acad Sci USA. 2003;100:581–86.
Nováková E, Hypsa V, Klein J, Foottit RG, von Dohlen CD, Moran NA. Reconstructing the phylogeny of aphids (Hemiptera: Aphididae) using DNA of the obligate symbiont Buchnera aphidicola. Mol Phylogenet Evol. 2013;68:42–54.
Fukatsu T, Aoki S, Kurosu U, Ishikawa H. Phylogeny of Cerataphidini aphids revealed by their symbiotic microorganisms and basic structure of their galls: implications for host-symbiont coevolution and evolution of sterile soldier castes. Zool Sci. 1994;11:613–23.
Fukatsu T, Ishikawa H. Phylogenetic position of yeast-like symbiont of Hamiltonaphis styraci (Homoptera, Aphididae) based on 18S rDNA sequence. Insect Biochem Molec Biol. 1996;26:383–88.
Vogel KJ, Moran NA. Sources of variation in dietary requirements in an obligate nutritional symbiosis. Proc Biol Sci. 2011;278:115–21.
Kozhoridze G, Orlovsky N, Orlovsky L, Blumberg DG, Golan-Goldhirsh A. Geographic distribution and migration pathways of Pistacia- present, past and future. Ecography. 2015;38:1141–54.
Muñoz-Viveros AL, Remaudiere G. Les espèces de Geopemphigus gallicoles sur Pistacia mexicana [Hemiptera, Aphididae, Eriosomatinae]. Rev Fr d’entomologie. 1999;21:35–48.
Hajibabaei M, Janzen D, Burns J, Hallwachs W, Hebert PDN. DNA barcodes distinguish species of tropical Lepidoptera. Proc Natl Acad Sci USA. 2006;103:968–71.
Lowe TM, Chan PP. tRNAscan-SE On-line: Search and contextual analysis of transfer RNA genes. Nucl. Acids Res. 2016;44: W54-57
Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28:27–30.
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016;44:D457–62.
Keseler IM, Mackie A, Peralta-Gil M, Santos-Zavaleta A, Gama-Castro S, Bonavides-Martinez C, et al. EcoCyc: fusing model organism databases with systems biology. Nucleic Acids Res. 2013;41:D605–12. (Database issue)
Moran NA, Tran P, Gerardo NM. Symbiosis and insect diversification: an ancient symbiont of sap-feeding insects from the bacterial phylum Bacteroidetes. Appl Environ Microbiol. 2005;71:8802–10.
Nikoh N, Nakabachi A. Aphids acquired symbiotic genes via lateral gene transfer. BMC Biol. 2009;7:12.
Nikoh N, McCutcheon JP, Kudo T, Miyagishima SY, Moran NA, Nakabachi A. Bacterial genes in the aphid genome: absence of functional gene transfer from Buchnera to its host. PLoS Genet. 2010;6:e1000827.
Shigenobu S, Stern DL. Aphids evolved novel secreted proteins for symbiosis with bacterial endosymbiont. Proc Biol Sci. 2013;280:20121952.
Nicholson SJ, Nickerson ML, Dean M, Song Y, Hoyt PR, Rhee H, et al. The genome of Diuraphis noxia, a global aphid pest of small grains. BMC Genomics. 2015;16:429.
Ramsey JS, Wilson AC, de Vos M, Sun Q, Tamborindeguy C, Winfield A, et al. Genomic resources for Myzus persicae: EST sequencing, SNP identification, and microarray design. BMC Genomics. 2007;8:423.
Wenger JA, Cassone BJ, Legeai F, Johnston JS, Bansal R, Yates AD et al. Whole genome sequence of the soybean aphid, Aphis glycines. Insect Biochem Mol Biol. 2017. https://doi.org/10.1016/j.ibmb.2017.01.005.
Koga R, Tsuchida T, Fukatsu T. Quenching autofluorescence of insect tissues for in situ detection of endosymbionts. Appl Entomol Zool. 2009;44:281–91.
Park JT, Uehara T. How bacteria consume their own exoskeletons (turnover and recycling of cell wall peptidoglycan). Microbiol Mol Biol Rev. 2008;72:211–27.
Rosas-Perez T, Rosenblueth M, Rincon-Rosales R, Mora J, Martinez-Romero E. Genome sequence of “Candidatus Walczuchella monophlebidarum” the flavobacterial endosymbiont of Llaveia axin axin (Hemiptera: Coccoidea: Monophlebidae). Genome Biol Evol. 2014;6:714–26.
Sabree ZL, Huang CY, Okusu A, Moran NA, Normark BB. The nutrient supplying capabilities of Uzinura, an endosymbiont of armoured scale insects. Environ Microbiol. 2013;15:1988–99.
Bennett GM, Moran NA. Small, smaller, smallest: the origins and evolution of ancient dual symbioses in a phloem-feeding insect. Genome Biol Evol. 2013;5:1675–88.
Muñoz-Viveros AL, Martinez J-JI, Molist P, González-Sierra S, Julián PG, Álvarez R, et al. Microscopic study of galls induced by three species of Geopemphigus (Hemiptera: Aphididae: Eriosomatinae) on Pistacia mexicana. Arthropod Plant Interact. 2014;8:531–38.
Al-Saghir MG. Evolutionary history of the genus Pistacia (Anacardiaceae). Int J Bot. 2009;5:255–57.
Inbar M. Systematics of Pistacia: Insights from specialist parasitic aphids. TAXON. 2008;57:238–42.
Yi T, Wen J, Golan-Goldhirsh A, Parfitt DE. Phylogenetics and reticulate evolution in Pistacia (Anacardiaceae). Am J Bot. 2008;95:241–51.
McCutcheon JP. The bacterial essence of tiny symbiont genomes. Curr Opin Microbiol. 2010;13:73–78.
Wernegreen JJ. Genome evolution in bacterial endosymbionts of insects. Nat Rev Genet. 2002;3:850–61.
Snyder AK, McMillen CM, Wallenhorst P, Rio RV. The phylogeny of Sodalis-like symbionts as reconstructed using surface-encoding loci. FEMS Microbiol Lett. 2011;317:143–51.
Clayton AL, Oakeson KF, Gutin M, Pontes A, Dunn DM, von Niederhausern AC, et al. A novel human-infection-derived bacterium provides insights into the evolutionary origins of mutualistic insect-bacterial symbioses. PLoS Genet. 2012;8:e1002990.
Enomoto S, Chari A, Clayton AL, Dale C. Quorum sensing attenuates virulence in Sodalis praecaptivus. Cell Host Microbe. 2017;21:629–36 e625.
Buchner P. Endosymbiosis of animals with plant microorganisms. New York: John Wiley & Sons; 1965.
Wegierek P, Michalik A, Wieczorek K, Kanturski M, Kobiałka M, Śliwa K et al. Buchnera aphidicola of the birch blister aphid, Hamamelistes betulinus (Horváth, 1896) (Insecta, Hemiptera, Aphididae: Hormaphidinae): molecular characterization, transmission between generations and its geographic significance. Acta Zoologica. 2017. https://doi.org/10.1111/azo.12186.
Vogel KJ, Moran NA. Functional and evolutionary analysis of the genome of an obligate fungal symbiont. Genome Biol Evol. 2013;5:891–904.
Burke G, Fiehn O, Moran N. Effects of facultative symbionts and heat stress on the metabolome of pea aphids. ISME J. 2010;4:242–52.
Dunbar HE, Wilson AC, Ferguson NR, Moran NA. Aphid thermal tolerance is governed by a point mutation in bacterial symbionts. PLoS Biol. 2007;5:1006–15.
Koga R, Tsuchida T, Fukatsu T. Changing partners in an obligate symbiosis: a facultative endosymbiont can compensate for loss of the essential endosymbiont Buchnera in an aphid. Proc Biol Sci. 2003;270:2543–50.
von Dohlen CD, Spaulding U, Patch KB, Weglarz KM, Foottit RG, Havill NP, et al. Dynamic acquisition and loss of dual-obligate symbionts in the plant-sap-feeding Adelgidae (Hemiptera: Sternorrhyncha: Aphidoidea). Front Microbiol. 2017;8:1037.
Manzano-Marin A, Szabo G, Simon JC, Horn M, Latorre A. Happens in the best of subfamilies: establishment and repeated replacements of co-obligate secondary endosymbionts within Lachninae aphids. Environ Microbiol. 2017;19:393–408.
Meseguer AS, Manzano-Marin A, Coeur d’Acier A, Clamens AL, Godefroid M, Jousselin E. Buchnera has changed flatmate but the repeated replacement of co-obligate symbionts is not associated with the ecological expansions of their aphid hosts. Mol Ecol. 2017;26:2363–78.
Toenshoff ER, Gruber D, Horn M. Co-evolution and symbiont replacement shaped the symbiosis between adelgids (Hemiptera: Adelgidae) and their bacterial symbionts. Environ Microbiol. 2012;14:1284–95.
Toju H, Tanabe AS, Notsu Y, Sota T, Fukatsu T. Diversification of endosymbiosis: replacements, co-speciation and promiscuity of bacteriocyte symbionts in weevils. ISME J. 2013;7:1378–90.
Rosenblueth M, Sayavedra L, Samano-Sanchez H, Roth A, Martinez-Romero E. Evolutionary relationships of flavobacterial and enterobacterial endosymbionts with their scale insect hosts (Hemiptera: Coccoidea). J Evol Biol. 2012;25:2357–68.
Acknowledgments
We acknowledge Ana Lilia Muñoz Viveros and Universidad Nacional Autónoma de México for samples. We also thank members of the N. Moran lab and the H. Ochman lab for comments and H. Ochman and S. Zhao for assistance with collections. Funding sources in support of this work were USDA 2016-67012-24676 to RAC and NSF DEB-1551092 to NAM.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Subject Categories: Microbe-microbe and microbe-host interactions.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Chong, R.A., Moran, N.A. Evolutionary loss and replacement of Buchnera, the obligate endosymbiont of aphids. ISME J 12, 898–908 (2018). https://doi.org/10.1038/s41396-017-0024-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-017-0024-6
This article is cited by
-
Host-specific co-evolution likely driven by diet in Buchnera aphidicola
BMC Genomics (2024)
-
Comparative transcriptomics of aphid species that diverged > 22 MYA reveals genes that are important for the maintenance of their symbiosis
Scientific Reports (2023)
-
Nutrient supplementation by genome-eroded Burkholderia symbionts of scale insects
The ISME Journal (2023)
-
Gene flow and introgression are pervasive forces shaping the evolution of bacterial species
Genome Biology (2022)
-
Aphid species specializing on milkweed harbor taxonomically similar bacterial communities that differ in richness and relative abundance of core symbionts
Scientific Reports (2022)