Abstract
Microbial mercury (Hg) methylation in sediments can result in bioaccumulation of the neurotoxin methylmercury (MMHg) in aquatic food webs. Recently, the discovery of the gene hgcA, required for Hg methylation, revealed that the diversity of Hg methylators is much broader than previously thought. However, little is known about the identity of Hg-methylating microbial organisms and the environmental factors controlling their activity and distribution in lakes. Here, we combined high-throughput sequencing of 16S rRNA and hgcA genes with the chemical characterization of sediments impacted by a waste water treatment plant that releases significant amounts of organic matter and iron. Our results highlight that the ferruginous geochemical conditions prevailing at 1–2 cm depth are conducive to MMHg formation and that the Hg-methylating guild is composed of iron and sulfur-transforming bacteria, syntrophs, and methanogens. Deltaproteobacteria, notably Geobacteraceae, dominated the hgcA carrying communities, while sulfate reducers constituted only a minor component, despite being considered the main Hg methylators in many anoxic aquatic environments. Because iron is widely applied in waste water treatment, the importance of Geobacteraceae for Hg methylation and the complexity of Hg-methylating communities reported here are likely to occur worldwide in sediments impacted by waste water treatment plant discharges and in iron-rich sediments in general.
Similar content being viewed by others
Introduction
Methylmercury (MMHg) is a neurotoxin that is biomagnified in aquatic food webs with fish consumption as a primary route for human MMHg exposure [1]. In aquatic ecosystems, methylation of mercury (Hg) to MMHg is carried out by microorganisms that are usually present at oxic-anoxic boundaries of sediments [2], water columns [3] and settling particles [4]. Studies on sediments have implicated sulfate-reducing bacteria (SRB) as the main contributors to MMHg formation, as inhibition of SRB with molybdate often decreases MMHg production rates [5,6,7,8,9]. Accordingly, many studies have aimed to identify the SRB strains responsible for Hg methylation by testing their Hg-methylating potential in pure cultures [10, 11], or by determining correlations between Hg methylation and the presence of SRB markers in sediments [12, 13] or water column [5]. Few studies have however linked Hg methylation to iron-reducing bacteria (FeRB) [14, 15], or methanogenic archaea [16]. The identification of the genes (hgcAB) required for Hg methylation [17] led to the discovery that the diversity of Hg methylators was much broader and the process more complex than previously thought [18]. Notably, the ability to methylate Hg has been identified within new phyla (Firmicutes and Chloroflexi) featuring also fermentative and acetogenic metabolisms [17,18,19]. The discovery of genetic markers for Hg-methylation also opened the door to more efficient, standardized, and detailed monitoring of Hg methylators in the environment. Accordingly, microbial Hg-methylating communities have recently been described in wetlands and paddy soils and included methanogens, SRB, FeRB, and syntrophs [19,20,21,22]. Hg-methylating microbial communities in lakes, on the other hand, remain poorly investigated with regards to the types of microorganisms involved and the factors influencing their activities, except that MMHg formation is determined by the molecular composition of organic matter (OM) [23].
Waste production can have a strong impact on the chemical and biological characteristics of freshwaters [24, 25]. As an example, treated waste water from the City of Lausanne is discharged into Lake Geneva (Switzerland) via an outlet pipe located in Vidy Bay. The dephosphatation treatment, based on the addition of ferric chloride to the waste water, causes enhanced inputs of Fe into the bay [24, 26]. Moreover, the current volume of sewage exceeds the capacity of the waste water treatment plant (WWTP), requiring transient releases of partially untreated water, which contributes to the enrichment of the sediments with OM and various trace metals [27]. Among them Hg is of major concern because of its toxicity and the high ambient concentrations (e.g., refs. [2, 25]). Previous studies have shown that the areas affected by the WWTP discharge are hotspots for MMHg formation and concentration [2]. In particular, it was found that sediments enriched in OM with prevailing ferruginous conditions (i.e., high concentrations of dissolved FeII and no free sulfide in the porewater) had the highest Hg methylation rates found in the bay [2]. Furthermore, sediment amendment experiments revealed that inhibition of dissimilatory sulfate reduction with molybdate led to increased Fe-reduction rates and enhanced Hg methylation. It was thus concluded that microorganisms other than SRB were responsible for Hg methylation [2], but the identity of the Hg-methylating microorganisms and thus the pathways by which MMHg was formed were not elucidated.
Building on this earlier discovery and recognizing the potential global implications of enhanced Hg methylation rates in WWTP-influenced sediments, the aim of this study was to provide a mechanistic understanding of MMHg formation in Fe-rich freshwater sediments. For this purpose, we linked sediment geochemistry and Hg speciation with parallel analysis of the genes coding for 16S rRNA and hgcA. In addition, the functional potential of the microbial communities was assessed by qPCR quantification of specific genes involved in the biogeochemical cycling of carbon, sulfur and iron. This is the first study in WWTP-impacted sediments exploring the diversity of microorganisms involved in Hg methylation.
Material and methods
Study site and sampling
The present study was conducted in the Vidy Bay of Lake Geneva, one of the largest freshwater reservoirs in Europe. Samples were collected at a site close to the WWTP outlet pipe (CP, 46°30'52.89” N, 6°35'21.25” E) based on previous studies showing a strong impact of the WWTP on MMHg formation [2]. Three replicate sediment cores (A, B, and C) were collected in August 2010 as detailed in the Supplementary Information.
Chemical analysis
Total carbon, organic carbon (Corg), inorganic carbon (Cinorg; samples exposed twice to 5 % H3PO4 overnight) and nitrogen (Ntot) in sediments were measured with a CHN elemental analyzer (Perkin Elmer 2400 series II). OM content was determined by loss on ignition (LOI) [28].
Metals were extracted from freeze-dried sediment with 2 N HNO3 at 100 °C for 12 h. Total Fe and S (Fetot; Stot) were measured by ICP-AOS (iCAP-6300 Duo Thermo Scientific). A detailed description of mercury speciation and quantification can be found elsewhere [2]. Briefly, sediment total Hg (Hgtot) was quantified using an Advanced Mercury Analyser (254 Altec). MMHg in sediments was extracted by HNO3 leaching/CH2Cl2 extraction and measured by ethylation onto Tenex traps followed by GC separation [29]. Hgtot in porewater was measured by CV-AFS [30]. MMHg in porewater was measured by species-specific isotope dilution and GC-ICPMS [31].
Porewater concentrations of SO4 2− and NO3 − were analyzed by ion-chromatography (Dionex ICS-3000) with an Ion-Pac_AS19 column. Poorly crystalline FeIII -oxides in the sediment were extracted with 0.5 M HCl, reduced to FeII with 1 M hydroxylamine hydrochloride and subsequently quantified with the ferrozine method [32]. Elemental sulfur (S0) was extracted from wet sediment with methanol and subsequently analyzed by RP-HPLC and UV detection at 265 nm [33].
DNA extraction and bacterial community composition: 16S rRNA gene
DNA was extracted in triplicate from freeze-dried sediments with the Power Soil DNA Extraction kit (Mo-Bio) as previously described [34]. PCR primers 341 F (5′-CCTACGGGNGGCWGCAG-3′) and 805 R (5′-GACTACHVGGGTATCTAATCC-3′) were used for amplification of the 16S rRNA gene from most bacteria [35, 36]. The resulting PCR products were used as template in an additional 10-cycle amplification with sample-specific barcoded primers according to Sinclair et al. [37]. See Supplementary Table 1 and Supplementary Information for details.
Amplicon sequencing data (Illumina MiSeq pair-end 300 bp) were preprocessed using FastQC [38] and the illumitag pipeline as described in Sinclair et al. [37]. Chimeric sequences were removed and reads were grouped into Operational Taxonomic Units (OTU) clustering at 97 % sequence identity. The taxonomical annotation of the OTUs was subsequently performed by CREST using the SILVA database [37]. Data were not rarefied to avoid overlooking rare members of the communities that may be more sensitive to the environmental gradients of interest [39]. Sequence data have been deposited in the European Nucleotide Archive (http://www.ebi.ac.uk/ena) under the accession number PRJEB20838.
Hg methylator community composition: hgcA
Primers targeting hgcA sequences were adopted from Schaefer et al. [22] and modified to include secondary priming-sequences for a second stage PCR where sample-specific barcodes and Illumina sequencing adaptors were added. For this purpose, the forward primer hgcA_261 F (5′-CGGCATCAAYGTCTGGTGYGC-3′) and the reverse primer hgcA_912 R (5′-GGTGTAGGGGGTGCAGCCSGTRWARKT-3′) with barcode adaptors, were first used to amplify the hgcA gene from each sample. For the second PCR, each sample was individually barcoded with unique combinations of forward and reverse primers (Supplementary Table 2). For details on the amplification strategy, see Supplementary Fig. 1. Second PCR amplicons were then purified using Agencourt AMPure XP (Beckman Coulter, USA), quantified using the PicoGreen kit (Invitrogen) and subsequently pooled in equal proportions. Amplicons were sequenced using the Illumina MiSeq instrument and pair-end 300 bp mode. Due to the length of the PCR product, only the forward read sequence was used for downstream data-analysis. Bad quality reads were trimmed with SICKLE [40], and left-over adapters and primers were removed with CUTADAPT [41]. Reads were truncated, dereplicated and singletons removed with USEARCH version 8.0 [42]. The obtained set of reads was then clustered using cd-hit-est with a 60 % similarity threshold [43]. The original quality-controlled reads were finally mapped to the representative sequences of the obtained clusters to generate a count table using the USEARCH software. The database used for annotation of the sequences is based on the sequences used by Podar et al. [19], additionally a Hidden Markov Model (HMM) based on these sequences was made using HMMER [44] and subsequently used to mine δ-Proteobacteria from the IMG database of JGI. The Sequences where curated and the taxonomy homogenized. Analogous to the 16S rRNA gene diversity analysis, data were not rarefied. More detailed information on the construction of hgcA gene libraries and phylogenetic analyses is given in Supplementary Information. Sequence data can be found in the European Nucleotide Archive under the accession number PRJEB20838.
Quantitative PCR
Quantitative PCR (qPCR) of marker genes (16S rRNA, dsrA, GCS, mcrA, and merA) for key functional groups involved in carbon, sulfur and iron cycling was performed in 10 µL total reaction volumes with primer pairs listed in Table S3 using an Eco cycler (Illumina, San Diego, California) and the Kapa SYBR Fast qPCR kit (Kapa Biosystems, Wilmington, Massachusetts), as described in Bravo et al. [45]. All reactions were performed according to the supplier’s protocol with 40 amplification cycles and real-time data acquired during the annealing step of each cycle. Standard curves and no-template controls were included in triplicate for each reaction and the target copy number per sample was calculated. The quality of standard curve and melting curves were tested with qpcR package (www.dr-spiess.de/qpcR.html; [46, 47]) in the freely available software environment R version 3.1.0 (http://www.r-project.org/). See Supplementary Information for more details.
Statistical analysis
Pearson correlations were calculated between chemical compounds using centered and reduced values in Excel (Microsoft, Redmond, WA, USA). Rarefaction curves (Supplementary Fig. 2) were plotted using R. Barplots showing relative abundances of major taxa were built with the RAM package (http://cran.r-project.org/package = RAM, [48]).
Results
Chemical analyses
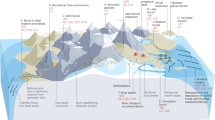
The geochemical composition of the 18 sediment samples features gradual changes with depth, but little variation among the 3 replicate sediment cores (Fig. 1; Supplementary Table 4). The decrease of SO4 2− together with increasing FeII and S0 concentrations with depth indicates that H2S produced by SRB is titrated out from the porewater by FeIII (Fig. 1, Supplementary Table 4). As a consequence, porewater sulfide concentrations remain low throughout the whole sediment core [2]. Carbon, OM and nitrogen varied only little with depth, with Corg representing about 40 % of Ctot, OM ranging from 5.5 to 7.2 %, and Corg/N-ratio ranging from 5.5 to 11.3, indicative of fresh OM of predominantly microbial and algal origin. Generally, the geochemical conditions are very similar to an earlier study conducted at the same site [2].
Map of the study area and chemical characterization of solid phases and porewater in three sediment cores collected at site CP near the outlet pipe of the sewage treatment plant (STP). Complete data are given in Supplementary Table 4.
Hgtot increased with depth from 0.13–1.67 µg g−1 d.w. (range of the three core replicates) to 1.73–5.58 µg g−1 d.w. at the deepest layer (Fig. 1). These values are 4- to 50-times higher than the natural background of Hgtot in Lake Geneva (0.03 mg kg−1) [49]. MMHg concentrations were generally higher in the surface layer (0–1 cm: 1.1–2.3 ng g−1) and in deeper layer (6–8 cm: 2.4–5.2 ng g−1) than at intermediate depth (Fig. 1, Supplementary Table 4). The proportion of MMHg to Hgtot was the highest at 1–2 cm (0.9–1.1 %) and decreased with depth to values between 0.28 % and 0.38 %. These Hgtot values and MMHg percentages are in line with analyses performed previously, but fall within the lower range of Hg concentrations reported for Vidy Bay [49, 50]. Pearson's correlation analysis revealed that MMHg concentrations were positively correlated to Corg, Stot, FeII and Hgtot (Figure S3). The proportion of Hgtot as MMHg was also correlated with porewater SO4 2− concentration, and inversely to the sedimentary S0 content (Supplementary Fig. 3). Overall, our results confirm the important role of S, Fe and Hg for the geochemistry in sediments of Vidy Bay [2].
Bacterial community composition: 16S rRNA gene
The composition and diversity of the combined bacterial community was investigated by sequencing of 16S rRNA amplicons. Close to 524’000 sequences (between 13’133 and 59’205 per sample) were clustered into OTUs at three different identity levels; 80 % roughly corresponding to the phylum level, 95 % corresponding to genus-level, and 97 % to species-level, the latter with 8044 OTUs identified. The overall coverage of the sediment community is reflected in the combined richness detected for random subsets of analyzed samples, where the logarithmic shape indicated that most of the species richness occurring in the sediments was covered in the combined dataset (Fig. 2). Overall the bacterial community composition was quite uniform across replicates and samples (Fig. 3, Supplementary Fig. 4), and showed only a slight change in diversity with depth at the OTU level (Supplementary Fig. 5).
Phylogenetic distribution of bacterial 16S rRNA gene sequences in the combined Vidy Bay sediment dataset: A. Relative abundance of major phyla and B classes within the δ-Proteobacteria. Presented data are average percentages (±SE) of reads from the 18 collected samples (6 depths in 3 cores). Categories representing >1 % are shown.
Phylogenetic analysis of 16S rRNA gene sequences identified Proteobacteria as the dominant phylum in terms of richness and relative abundance, represented by 1’605 OTUs or 28.6 ± 1.6 % of the reads, followed by Bacteroidetes (629 OTUs, 16.5 ± 2.4 %), and Chloroflexi (615 OTUs, 11 ± 2.7 %; Fig. 3). Within the Proteobacteria, δ-Proteobacteria (869 OTUs) were abundant and accounted for 31 % of the Proteobacteria reads. Firmicutes, mostly represented by Clostridiales, contributed 1.9 % of the total reads, but had representatives among the 50 most abundant OTUs in the total data set (Supplementary Table 5). Members of the Syntrophobacterales, including, e.g., Syntrophus, Syntrophorhabdus, and Syntrophobacteraceae, were also detected. Moreover, several lineages involved in Fe-transformation (e.g., Acidithiobacillus, Geobacter, Geothermobacter, and Shewanella; [51]) and sulfur cycling (e.g., Desulfobulbaceae, Desulfobacteraceae, etc.) were identified. Among the genera known to host representatives that carry hgcAB genes [18, 19], we identified Geobacter, Syntrophus, Syntrophorhabdus, Desulfobulbus, Desulfomonile, Desulfovibrio, Desulfomicrobium, unknown Ruminococcaceae genus, and Syntrophomonadaceae. In particular, Geobacter, Syntrophus, and Syntrophorhabdus represented 0.15, 0.29, and 0.21 % of total 16S rRNA reads, respectively.
Quantification of functional genes
Quantification of total bacterial abundance by qPCR detected between 1.4 × 107 and 6.4 × 108 16S rRNA gene copies g−1 of wet sediment (Supplementary Table 6). Abundance of 16S rRNA gene was highest in the uppermost sediment layers and decreased slightly with depth. Analyses of selected functional genes revealed wide occurrences of SRB (dsrA), Geobacteraceae (GCS) and methanogens (mcrA) (Supplementary Table 6). The abundance of SRB, Geobacteraceae and merA—encoding the mercuric reductase—slightly decreased with depth. Methanogens reached their highest representation in the deepest sediment layers, where the main electron acceptors FeIII and sulfate were depleted (Fig. 1).
A closer look at the Hg-methylating community through hgcA gene analysis
In addition to the phylogenetic community analysis, the diversity of Hg-methylating microorganisms was for the first time assessed by high-throughput sequencing of the hgcA gene. As for 16S rRNA, community rarefaction indicated that most of the Hg methylator diversity targeted by these primers was captured (Fig. 2). Analogous to 16S rRNA, Hg-methylating community composition was also quite uniform across replicates and samples with only minor changes with depth (Fig. 4, Supplementary Fig. 5). The hgcA gene data set consisted of altogether 741’890 reads ranging from 22’522 to 62’442 reads per sample. A total of 356 OTUs were identified at 88 % identity level, 325 of which were affiliated with Bacteria, 25 with Archaea, while the affiliation of 6 OTUs could not be resolved. Among the bacterial hgcA OTU’s, 264 were annotated as Proteobacteria, 41 as Firmicutes while 20 could not be linked to any specific phylum. In terms of their relative abundance, Bacteria represented 99 % of the reads, while Archaea represented 0.8 % of the reads.
Relative abundance of Hg-methylating families carrying hgcA in Vidy Bay sediments. “OTU_0032”, “OTU_0031”, “OTU_0014”, and “OTU_0630”, all abundant members of the Hg-methylating community, were annotated as unknown δ-Proteobacteria. Numbers in parentheses represent the sediment depth interval in centimeters.
All proteobacterial hgcA reads grouped with δ-Proteobacteria, with 27 % of the reads being Desulfuromonadales (153 OTUs), 71 % were from unknown δ-Proteobacteria (92 OTUs) and 0.1 % represented other δ-Proteobacteria, including Desulfovibrionales, Desulfobacterales and Syntrophobacterales (Figure S6). The 17 most abundant hgcA OTUs contributed 92 % of all reads and were all affiliated with δ-Proteobacteria. In particular, the three most abundant OTUs in terms of read counts (“OTU_0032”, “OTU_0031” and “OTU_0014”) were annotated as unknown δ-Proteobacteria and represented 22.8, 14.9, and 15.1 % of the reads, respectively (Fig. 4). The fourth most abundant OTU was affiliated with Geobacter spp. and contributed 14.7 % of reads. In addition, an unknown δ-Proteobacteria “OTU_0630” represented 4.4 % of the reads (Fig. 4). At higher taxonomic resolution, 99 % of the Desulfuromonadales reads were affiliated with Geobacteraceae (147 OTUs, 27 % of all hgcA reads). The two most abundant archaeal hgcA OTUs were affiliated with Methanomassiliicoccus spp. and each represented about 0.3 % of total hgcA reads. These OTUs were 20–112 times more abundant in the 4–6 cm and 6–8 cm of sediment than in 0–1 cm and 1–2 cm layers (Fig. 4). Hg-methylating Firmicutes were also detected (0.2 % of reads). In particular, a Dethiobacter sp. (Syntrophomonadaceae) represented 0.1 % of reads, whereas an Ethanoligenens sp. (Ruminococcaceae) accounted for 0.02 % of reads.
Phylogenetic analysis suggested that the most abundant Hg-methylating OTUs (“OTU_0032”, “OTU_0031” and “OTU_0014”) of Vidy Bay sediments were phylogenetically related to Geobacter species (Fig. 5). Therefore, when summing the reads of “OTU_0032”, “OTU_0031”, “OTU_0014”, and “OTU_0630” with those of identified Geobacteraceae, our data suggested that Desulfuromonadales could contribute to more than 85 % of the reads encountered (Fig. 4, Supplementary Fig. 6). Hence unknown Hg-methylating δ-Proteobacteria and unknown Hg-methylating bacteria combined, would account for less than 15 % of the reads (Supplementary Fig. 6). One OTU (“OTU_0121”), representing 0.9 % of reads and initially annotated as δ-Proteobacteria, was more closely related to Clostridium spp. according to the phylogenetic analysis.
Discussion
WWTP effluents foster specific geochemical conditions that favor Hg methylation processes
Effluents of the WWTP with Fe as dephosphatation agent are a known source of Hg [49] and create geochemical conditions favoring Hg methylation [2]. Here the highest proportion of Hgtot as MMHg was observed at 1–2 cm depth (Fig. 1, Table S4). The ferruginous conditions of Vidy Bay sediments, i.e. high porewater Fe2+ concentrations but virtually no dissolved sulfide available to bind Hg (Fig. 1), most likely keep Hg bound to OM [52] and thus available for methylation. Also, degradation of specific organic compounds, or reductive dissolution of FeIII(oxy)hydroxides under anoxic conditions, likely trigger the release of Hg, as indicated by porewater Hgtot peaking at this specific depth and accordingly enhance net MMHg formation in this layer. Considering that the abundance of functional genes (Supplementary Table 6), bacterial communities (16S rRNA, Figure S4) and the Hg-methylating communities (hgcA, Fig. 4, Supplementary Fig. 6) showed only minor variation over depth across all studied sediment profiles, our results highlight that geochemical conditions prevailing at 1–2 cm depth control MMHg formation. Thus the effluents from the WWTP might not only exert a strong influence on the redox states of Fe and S but also on Hg speciation and cycling.
Geobacteraceae: dominant members of the Hg-methylating guild in ferruginous sediments
By combined analysis of 16S rRNA and hgcA sequences, we identified microorganisms potentially involved in MMHg production. Here the observed richness and abundance imply that FeRB but also fermenters and other hitherto unknown Hg methylators are members of the Hg-methylating microbial community in sediments affected by OM- and Fe-rich WWTP effluents. However, both the 16S rRNA gene and hgcA gene analyses showed that δ-Proteobacteria, and in particular Desulfuromonadales, were abundant. Recent studies of hgcA diversity in wetlands and rice paddies have concluded that δ-Proteobacteria dominate Hg-methylating microbial communities [20,21,22] and this seems to hold also for the freshwater sediments studied here. Although the primers used in these studies were designed based on genomic DNA from a small subset of the known Hg-methylating microorganisms, most of them δ-Proteobacteria, our results demonstrate that hgcA sequencing was also successful in detecting some Archaea (Fig. 4) and members of other bacteria phyla such as Firmicutes. Among δ-Proteobacteria, the richness and the abundance of Geobacter spp. in the hgcA data set pointed to their importance for Hg methylation in the studied sediments. Only a few previous studies have demonstrated that Hg methylation in sediments can occur under Fe-reducing conditions with FeRB mediating the process [2, 14, 15]. Instead, SRB have historically been identified as the principal Hg methylators in aquatic systems [9, 12, 13, 53, 54]. Although we identified several SRB by 16S rRNA gene sequencing—with quantification of drsA gene confirming that SRB are widely present in these sediments—the abundance of identified SRB (e.g., Desulfovibrio spp.) in the hgcA diversity analyses was significantly smaller than for fermenters (e.g., Ruminococcaceae and Syntrophomonadaceae) and FeRB (Geobacteraceae). The latter observation is in line with previous cultivation-based estimates suggesting FeRB to be three orders of magnitude more abundant than SRB in these sediments and that FeIII reduction rates was positively correlated to Hg methylation rates [2].
A geomicrobiological model for coupled Fe, S cycling and MMHg formation in sediments affected by WWTP discharges
We propose that FeRB play a central role in both Hg methylation and anaerobic OM degradation in these sediments. Indeed, Geobacter spp., known to inhabit sediments and sludge [55], can reduce FeIII or S0 but also perform interspecies electron transfer to S0-reducers or methanogens [56]. Syntrophic bacteria, however, have been recently identified as putatively important Hg methylators [21], and have also been detected in sediments adjacent to the WWTP discharge pipe outlet [57]. Accordingly, one known syntrophic lineage, Smithella spp. (Syntrophaceae, 17 reads) was identified in the current hgcA analyses. Some lineages detected in Vidy Bay sediments, such as Syntrophobacteraceae (3574 reads) and Syntrophaceae (5805 reads), might degrade OM in syntrophic association with H2 consuming organisms, such as FeRB, SRB or methanogens [58, 59]. In this context, indirect effects of non-Hg-methylating microorganisms on MMHg formation should also be considered. For example methanogens could maintain the activity of syntrophic Hg methylators, especially in deeper sediment layers where FeIII and SO4 2− are depleted [60]. In short, it is likely that Geobacter sp get enriched by the continuous import of FeIII to the sediment surface and continuous reoxidation of FeII at its surface through mixing. In deeper sediment layers, these microbes likely survive through alternative metabolic processes such as S0 reduction or syntrophic oxidation of OM [60]. Considering our results here and previous research carried out in Vidy Bay [2], we propose a geomicrobiological model (Figure S7) where Fe reacts with sulfide (either produced by SRB through sulfate reduction or liberated from biomass through anaerobic degradation of amino acids) according to: 2FeOOH + HS− + 5 H+ ⇒ 2Fe2+ + S0 + 4H2O. As a result, anoxic conditions prevail in these sediments while sulfide is low and Fe2+ and S0 are high. Elemental sulfur in these sediments might then be recycled by S0-reduction performed by e.g., Geobacter or Dethiobacter [61], which are both identified Hg methylators [19]. Hence, FeIII might act as a direct electron acceptor for Geobacter, but also as an sulfide “scrubber” according to the stoichiometry above and as previously suggested for freshwater sediments (Haveman et al., 2008; [62]). These results provide novel understanding on the role of Geobacter in Hg methylation processes linked to Fe and S cycling in a low-sulfate freshwater environment.
Conclusions
Previous efforts to identify Hg methylators have mainly been limited to functional characterization of cultured isolates and inferences made about their close relatives based on phylogenetic markers such as the 16S rRNA gene. A more recent alternative strategy has been to probe directly hgcA genes using target amplification, cloning and sequencing. Here, combination of high-throughput sequencing approach with molecular barcoding and amplification protocols identified Hg-methylating microbial communities in freshwater sediments. Our results reveal the coexistence of a wide abundance of fermenters, secondary fermenters (e.g. syntrophs), SRB, FeRB, and methanogens in this Hg-contaminated freshwater lake ecosystem and point to the significant contribution of syntrophs, fermenters, Fe and S reducers to the diversity and abundance of the Hg-methylating community in the sediments rich in OM and Fe. Moreover, known SRB Hg methylators constituted relatively minor groups compared to the total Hg-methylating and SRB community, supporting the idea that sediments affected by WWTP effluents are in this regard different from most other anoxic aquatic settings previously studied. Our results suggest that geochemical conditions rather than the composition of the resident microbiota dictate net MMHg formation. Since Fe is widely used in waste water treatment across the globe, the importance of Geobacteraceae for Hg methylation and the geochemical conditions reported here are likely relevant for WWTP recipients worldwide.
References
Mason RP, Choi AL, Fitzgerald WF, Hammerschmidt CR, Lamborg CH, Soerensen AL, et al. Mercury biogeochemical cycling in the ocean and policy implications. Environ Res. 2012;119:101–117.
Bravo AG, Bouchet S, Guédron S, Amouroux D, Dominik J, Zopfi J. High methylmercury production under ferruginous conditions in sediments impacted by sewage treatment plant discharges. Water Res. 2015;80:245–255.
Eckley CS, Watras CJ, Hintelmann H, Morrison K, Kent AD, Regnell O. Mercury methylation in the hypolimnetic waters of lakes with and without connection to wetlands in northern Wisconsin. Can J Fish Aquat Sci. 2005;411:400–411.
Gascon Diez E, Loizeau J-L, Cosio C, Bouchet S, Adatte T, Amouroux D, et al. Role of settling particles on mercury methylation in the oxic water column of freshwater systems. Environ Sci Technol. 2016;50:11672–11679.
Achá D, Pabón CA, Hintelmann H. Mercury methylation and hydrogen sulfide production among unexpected strains isolated from periphyton of two macrophytes of the Amazon. FEMS Microbiol Ecol. 2012;80:637–45.
Gilmour CC, Henry EA, Mitchell R. Sulfate stimulation of mercury methylation in freshwater sediments. Environ Sci Technol. 1992;26:2281–2287.
Hines ME, Poitras EN, Covelli S, Faganeli J, Emili A, Žižek S, et al. Mercury methylation and demethylation in Hg-contaminated lagoon sediments (Marano and Grado Lagoon, Italy). Estuar Coast Shelf Sci. 2012;113:85–95.
Avramescu M-L, Yumvihoze E, Hintelmann H, Ridal J, Fortin D, Lean DRS. Biogeochemical factors influencing net mercury methylation in contaminated freshwater sediments from the St. Lawrence River in Cornwall, Ontario, Canada. Sci Total Environ. 2011;409:968–978.
Pak K, Bartha R. Mercury methylation and demethylation in anoxic lake sediments and by strictly anaerobic bacteria. Appl Environ Microbiol. 1998;64:1–6.
Bridou R, Monperrus M, Gonzalez PR, Guyoneaud R, Amouroux D. Simultaneous determination of mercury methylation and demethylation capacities of various sulfate-reducing bacteria using species-specific isotopic tracers. Environ Toxicol Chem. 2011;30:337–344.
Ranchou-Peyruse M, Monperrus M, Bridou R, Duran R, Amouroux D, Salvado JC, et al. Overview of mercury methylation capacities among anaerobic bacteria including representatives of the sulphate-reducers: implications for environmental studies. Geomicrobiol J. 2009;26:1–8.
Devereux R, Winfrey MR, Winfrey J, Stahl DA. Depth profile of sulfate-reducing bacterial ribosomal RNA and mercury methylation in an estuarine sediment. FEMS Microbiol Ecol. 1996;20:23–31.
Shao D, Kang Y, Wu S, Wong MH. Effects of sulfate reducing bacteria and sulfate concentrations on mercury methylation in freshwater sediments. Sci Total Environ. 2012;424:331–6.
Fleming EJ, Mack EE, Green PG, Nelson DC. Mercury methylation from unexpected sources: molybdate-inhibited freshwater sediments and an iron-reducing bacterium. Appl Environ Microbiol. 2006;72:457–464.
Yu R-Q, Flanders JR, Mack EE, Turner R, Mirza MB, Barkay T. Contribution of coexisting sulfate and iron reducing bacteria to methylmercury production in freshwater river sediments. Environ Sci Technol. 2012;46:2684–2691.
Hamelin S, Amyot M, Barkay T, Wang Y, Planas D. Methanogens: principal methylators of mercury in lake periphyton. Environ Sci Technol. 2011;45:7693–7700.
Parks JM, Johs A, Podar M, Bridou R, Hurt RA, Smith SD, et al. The genetic basis for bacterial mercury methylation. Science. 2013;339:1332–1335.
Gilmour CC, Podar M, Bullock AL, Graham AM, Brown SD, Somenahally AC, et al. Mercury methylation by novel microorganisms from new environments. Environ Sci Technol. 2013;47:11810–11820.
Podar M, Gilmour CC, Brandt CC, Soren A, Brown SD, Crable BR, et al. Global prevalence and distribution of genes and microorganisms involved in mercury methylation. Sci Adv. 2015;1:e1500675.
Liu Y-R, Yu R-Q, Zheng Y-M, He J-Z. Analysis of the microbial community structure by monitoring an Hg methylation gene (hgcA) in paddy soils along an Hg gradient. Appl Environ Microbiol. 2014;80:2874–2879.
Bae HS, Dierberg FE, Ogram A. Syntrophs dominate sequences associated with the mercury methylation-related gene hgcA in the water conservation areas of the Florida Everglades. Appl Environ Microbiol. 2014;80:6517–6526.
Schaefer JK, Kronberg R-M, Morel FMM, Skyllberg U. Detection of a key Hg methylation gene, hgcA, in wetland soils. Environ Microbiol Rep. 2014;6:441–447.
Bravo AG, Bouchet S, Tolu J, Björn E, Mateos-Rivera A, Bertilsson S. Molecular composition of organic matter controls methylmercury formation in boreal lakes. Nat Commun. 2017;8:14255.
Gibbs-Eggar Z, Jude B, Dominik J, Loizeau J, Oldfield F. Possible evidence for dissimilatory bacterial magnetite dominating the magnetic properties of recent lake sediments. Earth Planet Sci Lett. 1999;168:1–6.
Poté J, Haller L, Loizeau J-L, Garcia Bravo A, Sastre V, Wildi W. Effects of a sewage treatment plant outlet pipe extension on the distribution of contaminants in the sediments of the Bay of Vidy, Lake Geneva, Switzerland. Bioresour Technol. 2008;99:7122–7131.
Percak-Dennett EM, Loizeau J-L, Beard BL, Johnson CM, Roden EE. Iron isotope geochemistry of biogenic magnetite-bearing sediments from the Bay of Vidy, Lake Geneva. Chem Geol. 2013;360–361:32–40.
Loizeau J-L, Pardos M, Monna F, Peytremann C, Haller L, Dominik J. The impact of a sewage treatment plant’s effluent on sediment quality in a small bay in Lake Geneva (Switzerland-France). Part 2: Temporal evolution of heavy metals. Lakes Reserv Res Manag. 2004;9:53–63.
Dean WE. Determination of carbonate and organic matter in calcareous sediments and sedimentary rocks by loss on ignition: comparison with other methods. J Sediment Res. 1974;44:242–248.
Liu B, Yan H, Wang C, Li Q, Guédron S, Spangenberg JE, et al. Insights into low fish mercury bioaccumulation in a mercury-contaminated reservoir, Guizhou, China. Environ Pollut. 2012;160:109–117.
Bloom N, Fitzgerald WF. Determination of volatile mercury species at the picogram level by low-temperature gas chromatography with cold-vapour atomic fluorescence detection. Anal Chim Acta. 1988;208:151–161.
Rodriguez Martín-Doimeadios RC, Monperrus M, Krupp E, Amouroux D, Donard OFX. Using speciated isotope dilution with GC-inductively coupled plasma MS to determine and unravel the artificial formation of monomethylmercury in certified reference sediments. Anal Chem. 2003;75:3202–3211.
Thamdrup B, Fossing H, Jorgensen BB. Manganese, iron, and sulfur cycling in a coastal marine sediment, Aarhus Bay, Denmark. Geochim Cosmochim Acta. 1994;58:5115–5129.
Zopfi J, Böttcher ME, Jørgensen BB. Biogeochemistry of sulfur and iron in Thioploca-colonized surface sediments in the upwelling area off central Chile. Geochim Cosmochim Acta. 2008;72:827–843.
Regier N, Frey B, Converse B, Roden E, Grosse-Honebrink A, Bravo AG, et al. Effect of Elodea nuttallii roots on bacterial communities and MMHg proportion in a Hg polluted sediment. PLoS ONE. 2012;7:e45565.
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013;41:1–11.
Logares R, Lindström ES, Langenheder S, Logue JB, Paterson H, Laybourn-Parry J, et al. Biogeography of bacterial communities exposed to progressive long-term environmental change. ISME J. 2013;7:937–948.
Sinclair L, Osman OA, Bertilsson S, Eiler A. Microbial community composition and diversity via 16S rRNA gene amplicons: Evaluating the Illumina platform. PLoS ONE. 2015;10:e0116955.
Andrews S. (2012). FastQC: A quality control tool for high throughput sequence data. Available online at: http://www.bioinformatics.bab.
McMurdie PJ, Holmes S. Waste not, want not: Why rarefying microbiome data is inadmissible McHardy AC (ed). PLoS Comput Biol. 2014;10:e1003531.
Joshi NA, Fass JN. (2011). Sickle: A sliding-window, adaptive, quality-based trimming tool for FastQ files.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:10–12.
Edgar RC. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods. 2013;10:996–998.
Fu L, Niu B, Zhu Z, Wu S, Li W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics. 2012;28:3150–3152.
Eddy SR. Accelerated profile HMM searches. PLoS Comput Biol. 2011;7:e1002195.
Bravo AG, Loizeau JL, Dranguet P, Makri S, Björn E, Ungureanu VG et al. Persistent Hg contamination and occurrence of Hg-methylating transcript (hgcA) downstream of a chlor-alkali plant in the Olt River (Romania). Environ Sci Pollut Res. 2016;11:10529–10541.
Ritz C, Spiess AN. qpcR: An R package for sigmoidal model selection in quantitative real-time polymerase chain reaction analysis. Bioinformatics. 2008;24:1549–1551.
Tichopad A, Dilger M, Schwarz G, Pfaffl M. Standardized determination of real-time PCR efficiency from a single reaction set-up. Nucleic Acids Res. 2003;31:122–122.
Chen W, Simpson J, Levesque C. (2016). RAM: R for amplicon-sequencing-based microbial-ecology. R package version 1.2.1.3.
Bravo AG, Bouchet S, Amouroux D, Poté J, Dominik J. Distribution of mercury and organic matter in particle-size classes in sediments contaminated by a waste water treatment plant: Vidy Bay, Lake Geneva, Switzerland. J Environ Monit. 2011;13:974–982.
Gascon Diez E, Bravo AG, Porta N, Masson M, Graham ND, Stoll S, et al. Influence of a wastewater treatment plant on mercury contamination and sediment characteristics in Vidy Bay (Lake Geneva, Switzerland). Aquat Sci. 2014;76:21–32.
Weber KA, Achenbach LA, Coates JD. Microorganisms pumping iron: anaerobic microbial iron oxidation and reduction. Nat Rev Microbiol. 2006;4:752–764.
Liem-Nguyen V, Skyllberg U, Björn E. Thermodynamic modelling of the solubility and chemical speciation of mercury and methylmercury driven by organic thiols and micromolar sulfide concentrations in boreal wetlands. Environ Sci Technol. 2017;51:3678–3686.
Bravo AG, Cosio C, Amouroux D, Zopfi J, Chevalley P-A, Spangenberg JE, et al. Extremely elevated methyl mercury levels in water, sediment and organisms in a Romanian reservoir affected by release of mercury from a chlor-alkali plant. Water Res. 2014;49:391–405.
Compeau GC, Bartha R. Sulfate-Reducing Bacteria: principal methylators of mercury in anoxic estuarine sediment. Appl Environ Microbiol. 1985; 50(2):498-502.
McInerney MJ, Struchtemeyer CG, Sieber J, Mouttaki H, Stams AJM, Schink B, et al. Physiology, ecology, phylogeny, and genomics of microorganisms capable of syntrophic metabolism. Ann NY Acad Sci 2008;1125:58–72.
Lovley DR, Ueki T, Zhang T, Malvankar NS, Shrestha PM, Flanagan KA et al. Geobacter: The microbe electric’s physiology, ecology, and practical applications.Adv Microb Physiol. 2011;59:1-100.
Haller L, Tonolla M, Zopfi J, Peduzzi R, Wildi W, Poté J. Composition of bacterial and archaeal communities in freshwater sediments with different contamination levels (Lake Geneva, Switzerland). Water Res. 2011;45:1213–1228.
Chen S, Liu X, Dong X. Syntrophobacter sulfatireducens sp. nov., a novel syntrophic, propionate-oxidizing bacterium isolated from UASB reactors. Int J Syst Evol Microbiol. 2005;55:1319–1324.
Bok FAM, Stams AJM, Dijkema C, Boone DR. Pathway of propionate oxidation by a syntrophic culture of Smithella propionica and Methanospirillum hungatei. Appl Environ Microbiol. 2001;67:1800–1804.
Holmes DE, Shrestha PM, Walker DJF, Dang Y, Nevin KP, Woodard TL et al. Metatranscriptomic evidence for direct interspecies electron transfer between Geobacter and Methanothrix species in methanogenic rice paddy soils. Appl Environ Microbiol. 2017; 83:e9 e00223-17.
Poser A, Lohmayer R, Vogt C, Knoeller K, Planer-Friedrich B, Sorokin D, et al. Disproportionation of elemental sulfur by haloalkaliphilic bacteria from soda lakes. Extremophiles. 2013;17:1003–1012.
Hansel CM, Lentini CJ, Tang Y, Johnston DT, Wankel SD, Jardine PM. Dominance of sulfur-fueled iron oxide reduction in low-sulfate freshwater sediments. ISME J. 2015;9:2400–2412.
Acknowledgements
We thank Philippe Arpagaus and Alexander Grosse-Honebrink for their help during field-work. Camille Thomas and Daniel Ariztegui are acknowledged for helping with the CHN analyses. This work was supported by the State Secretariat for Education and Research (SER) through grants to C.C. (COST-FA0906 contract no C11.0117), the Swiss National Science Foundation through grants to C.C. (project no 205321_138254 and 200020_157173) and the Swedish Research Council (VR) through grants to A.G.B. (project no 2011-7192) and S.B. (project no 2012-3892 and 2013-6978).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bravo, A.G., Zopfi, J., Buck, M. et al. Geobacteraceae are important members of mercury-methylating microbial communities of sediments impacted by waste water releases. ISME J 12, 802–812 (2018). https://doi.org/10.1038/s41396-017-0007-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-017-0007-7
This article is cited by
-
Recent advance of microbial mercury methylation in the environment
Applied Microbiology and Biotechnology (2024)
-
Soil Geobacteraceae are the key predictors of neurotoxic methylmercury bioaccumulation in rice
Nature Food (2024)
-
Mining-impacted rice paddies select for Archaeal methylators and reveal a putative (Archaeal) regulator of mercury methylation
ISME Communications (2023)
-
Metabolically diverse microorganisms mediate methylmercury formation under nitrate-reducing conditions in a dynamic hydroelectric reservoir
The ISME Journal (2023)
-
Effect of exogenous and endogenous sulfide on the production and the export of methylmercury by sulfate-reducing bacteria
Environmental Science and Pollution Research (2023)