Abstract
Lkb1 deficiency confers the Kras-mutant lung cancer with strong plasticity and the potential for adeno-to-squamous transdifferentiation (AST). However, it remains largely unknown how Lkb1 deficiency dynamically regulates AST. Using the classical AST mouse model (Kras LSL-G12D/+;Lkb1flox/flox, KL), we here comprehensively analyze the temporal transcriptomic dynamics of lung tumors at different stages by dynamic network biomarker (DNB) and identify the tipping point at which the Wnt signaling is abruptly suppressed by the excessive accumulation of reactive oxygen species (ROS) through its downstream effector FOXO3A. Bidirectional genetic perturbation of the Wnt pathway using two different Ctnnb1 conditional knockout mouse strains confirms its essential role in the negative regulation of AST. Importantly, pharmacological activation of the Wnt pathway before but not after the tipping point inhibits squamous transdifferentiation, highlighting the irreversibility of AST after crossing the tipping point. Through comparative transcriptomic analyses of mouse and human tumors, we find that the lineage-specific transcription factors (TFs) of adenocarcinoma and squamous cell carcinoma form a “Yin-Yang” counteracting network. Interestingly, inactivation of the Wnt pathway preferentially suppresses the adenomatous lineage TF network and thus disrupts the “Yin-Yang” homeostasis to lean towards the squamous lineage, whereas ectopic expression of NKX2-1, an adenomatous lineage TF, significantly dampens such phenotypic transition accelerated by the Wnt pathway inactivation. The negative correlation between the Wnt pathway and AST is further observed in a large cohort of human lung adenosquamous carcinoma. Collectively, our study identifies the tipping point of AST and highlights an essential role of the ROS-Wnt axis in dynamically orchestrating the homeostasis between adeno- and squamous-specific TF networks at the AST tipping point.
Similar content being viewed by others
Introduction
Lung cancer, the devastating disease with high mortality, is notorious for its extremely high heterogeneity and strong plasticity. Two main histological types of non-small-cell lung carcinoma (NSCLC, ~86% of lung cancer), adenocarcinoma (ADC) and squamous cell carcinoma (SCC), are characterized by their respective lineage transcription factors (TFs): NKX2-1 for ADC, and TP63 for SCC.1,2,3,4,5,6,7 Perturbation of those lineage-specific TFs may result in the destabilization and loss of the corresponding cellular identity.6,8,9 Interestingly, the existence of mixed adenosquamous (AdSCC) histology in clinic10,11 and the largely shared genetic alterations between these two histological portions12,13,14 indicate potential phenotypic transition, which is known as the adeno-to-squamous transdifferentiation (AST).15,16,17 Squamous transformation is recently observed in relapsed EGFR-mutant lung ADC patients after initially successful tyrosine kinase inhibitor (TKI) treatments.16,18 Moreover, a recent study shows that squamous transformation becomes prevalent in EGFR-mutant lung ADC patients relapsed from the treatment of Osimertinib, the third-generation TKI.18 Similar phenomena are also observed in 2/9 of lung ADC patients after the acquisition of KRASG12C inhibitor resistance.19 These studies have highlighted an important contribution of AST to the acquisition of molecular targeted therapy resistance in lung cancer clinics.
LKB1 (also named as serine-threonine kinase 11, STK11) is mutated in ~17% human lung ADC20 whereas its mutation rate is enriched in lung AdSCC, averaging at 39.66% (ranging from 22 to 66% in multiple studies).12,20,21,22,23 A recent study has analyzed LKB1 mutations in relapsed patients with potential adeno-to-squamous transdifferentiation, which shows a relatively low rate at 14.3% (1 out of 7).20 We reason that such low LKB1 mutation rate might be ascribed to the mutual exclusivity between LKB1 and EGFR mutations24,25,26 since most patients harbor EGFR genetic alterations, e.g., 4/7 with EGFR mutations and 1/7 with EGFR amplification.20 Excluding these EGFR-altered patients, the LKB1 mutation rate ranges from 33 to 50%, similar to previous reports.12,20,21,22,23 These clinical observations collectively indicate a potential role of LKB1 mutations in driving AST. Consistently, we have previously found that Lkb1 inactivation is able to drive lung cancer AST in the KrasG12D-based genetically engineered mouse models (GEMM).23,27,28,29 These pathologically transitioned mouse tumors also display therapeutic resistance to multiple inhibitors initially effective in lung ADC,29 similar to the clinical observations.
In the KL mouse model,30 ADC typically arises at 6 weeks post nasal inhalation of Adeno-Cre (Ad-Cre) and subsequently transdifferentiates into SCC at 8 weeks.28,29 LKB1-deficiency can trigger strong metabolic imbalance and excessive accumulation of reactive oxygen species (ROS).29 Such uncontrolled oxidative stress subsequently promotes the transdifferentiation from adenomatous pathology to squamous pathology.29 However, the underlying mechanisms remain largely unknown. Moreover, the transcriptomic and molecular dynamics of the Lkb1-loss-driven AST process, which should be crucial to uncover such mechanisms, have not yet been characterized.
From the system dynamics perspective, AST is among a category of nonlinear processes known as critical transition, which has been widely found in physiological systems, ecological systems, climate systems, and social systems.31,32,33,34,35 In general, such transition involves a drastic switch in the system state as it approaches the critical threshold referred to as the tipping point (TP).34 Before passing through the tipping point, the system state gradually changes but still holds its original state. As it gets close to the tipping point, a slight incremental change or even some stochastic noise could cause a catastrophic shift, which eventually brings the state to an alternative state and makes it difficult to return to the original state.36,37 In other words, the same perturbation on the system state before or after the tipping point might have different effects. It would therefore be important to detect the tipping point,36 which not only provides early-warning signals of disease progression but also identifies potentially important driving factors. Based on the non-linear dynamical theory, we have previously developed a network-based indicator of tipping point, i.e., dynamic network biomarker (DNB), which could identify the dominant genes controlling the TP based on differential covariance rather than differential gene expression commonly used in traditional bioinformatic analyses.36,38,39,40,41 The covariance includes both the correlations and deviations among genes. Notably, DNB has been recently successfully applied to multiple biological transitions including virus infection, disease onset, and cancer metastasis.36,38,39,40
Taking advantage of the KL mouse model for AST study, we here systematically characterize the temporal transcriptomic dynamics of lung tumors, delineate the non-linear dynamical process, and further identify the oxidative stress-triggered Wnt inactivation as the tipping-point regulator during the AST process. We propose a model for the AST process involving the Wnt inactivation-mediated disruption of the ADC- and SCC-lineage-specific transcriptional factors counteracting “Yin-Yang” network.
Results
Temporal transcriptomic characterization of AST
To uncover the temporal transcriptomic dynamics of lung tumors during the AST process, we took advantage of the KL mouse model, the well-established AST model.23,27,28 Consistent with the previous study,28 we found that lung ADC was evident at 6 weeks post Ad-Cre administration, whereas SCC began to emerge at 8 weeks. To monitor the temporal transcriptomic dynamics during the transdifferentiation process, we took a serial-sampling strategy post Ad-Cre administration for RNA-seq analyses. The mouse specimens were respectively atypical adenomatous hyperplasia (AAH) at 4 weeks (4 W), ADC at 6 and 7 weeks (6 W and 7 W), SCC at 8, 9, and 10 weeks (8 W, 9 W, 10 W) post Ad-Cre administration, which were validated by histopathological analyses and immunostaining of ADC markers including SFTPC and NKX2-1, and SCC markers including p63 and SOX2 (Fig. 1a). Normal lung (NL) tissues were also included as the control. The transcriptomic data showed a pattern of marker gene expression consistent with the histopathological analyses (Fig. 1b). After the dimension reduction with principal component analysis (PCA) and visualization of the samples on the first two principal components, we observed a clear trajectory of state transition, following three major phases consecutively: benign state (NL or AAH), to ADC, and eventually to SCC (Fig. 1c). Compared to the ADC at 6 W, the 7 W ADC showed a higher variability at transcriptomic level (Fig. 1c) as well as individual biomarker gene expression level, e.g., the ADC lineage markers Nkx2-1 and Napsa (Fig. 1b, d). This indicates that the 7 W ADC is at a relatively unstable state. This promoted us to treat the AST event as a non-linear process mediated by critical transition, using the theories and methods developed for dynamical systems.
Temporal transcriptomic characterization of AST in KL mouse model. a Representative H&E staining and immunohistochemistry (IHC) photos of KL mouse lungs at a series of time points after Ad-Cre nasal inhalation, ranging from 4 weeks (4 W) to 10 weeks (10 W). 4W-7W were ADC and 8W-10W were SCC. Normal lung (NL) were used as control. Scale bar: 50 µm. b RNA-seq analyses of KL lung lesions at a series of time points. Relative expression of canonical ADC and SCC marker genes were depicted. c Principal component analyses (PCA) of RNA-seq data revealing the trajectory of phenotypic transitions from benign tissue to ADC to SCC. Each node represented one sample, with the number indicating the time points (weeks) of sampling. 0 indicated NL. d Transcriptional dynamics of ADC markers (Nkx2-1, Napsa) and SCC markers (Trp63, Sox2) on the same PCA embedding as c. Node colors reflected the relative expression of each marker gene
Dynamic network biomarker analyses reveal the tipping point of AST
According to the bifurcation theory of dynamical systems, the tipping point indicates a state before the imminent critical transition of the system, accompanied by a drastic decline of system resilience, triggering abrupt and irreversible state shift.31,34,35,42,43 We then applied this theoretical framework to analyze the AST process (Fig. 2a). The lung tumor system begins with a gradual (reversible) state change in the ADC phase, approaching the tipping point (TP) after which it undergoes a critical (irreversible) state shift, and abruptly switches to the SCC state, which is then further stabilized. This whole process is accompanied by an earlier loss and a later re-gain of system resilience. To quantitatively delineate when and how the critical transition of AST occurred, we employed a previously developed method, dynamic network biomarker (DNB),36,38,39,40 to characterize the critical molecular network of the tipping point. The DNB method designs a composite index (CI) to measure system criticality, and the CI would reach its maximum as the system approaches the tipping point. The DNB analyses revealed a sharp peak at 7 W, indicating a candidate tipping point (Fig. 2b). From the dynamical perspective, the DNB member genes were composed of a strongly fluctuated and highly correlated sub-network specifically at 7 W (Fig. 2c), a time point consistent with our previous analyses (Fig. 1b–d). This was further confirmed by another method with the application of sliding window in silico to artificially increase the number of samples at each time point (Supplementary Fig. 1a). Our data collectively identified the 7 W was the TP in the AST process.
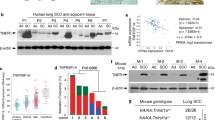
Dynamic network biomarker analyses reveal the tipping point of AST and its regulatory signaling. a Schematic diagram of a theoretical model for system state transition during AST. As lung tumors phenotypically switched from adenocarcinoma (ADC) to squamous carcinoma (SCC), three general phases could be observed: ADC phase, critical phase, SCC phase. The tumor state first gradually changes during the ADC phase, then reaches the tipping point (TP) before the drastic changes, and finally reaches a second slow-changing SCC state. b A composite index (CI) for quantifying the tipping point of system state. The peak of CI at 7 W indicates the tipping point. c. Network dynamics along the sampling time points during the AST process. A key module, called the dynamic network biomarker (DNB), is detected as the critical state at the 7 W (tipping point). Edges were colored by normalized absolute Pearson’s correlation coefficient (PCC). Nodes were colored by individual CI scores. d KEGG pathway enrichment for the DNB genes. Top 5 most significant pathways were shown. e TF enrichment in the upstream of genes repressed across the tipping point. Top 20 TFs are ranked with −log10(P value) and the ADC lineage-specific TFs and Wnt signaling related TFs were highlighted. f Representative photos for immunohistochemical staining of 8-oxo-dGuo and β-catenin in KL mouse lung tumors at 7 W post Ad-Cre treatment. Scale bar: 50 µm. g Correlation of immunohistochemical staining of 8-oxo-dGuo and β-catenin in KL mouse lung tumors at 7 W post Ad-Cre administration. Pearson correlation coefficent (PCC) and P value were shown. h The KL lung cancer cells were treated with indicated ROS inducers (PHEN at 500 μM, PL at 12.5 μM, PEITC at 15 μM) for 6 h and the relative expression of the Wnt pathway-related genes were assessed by real-time PCR. *P < 0.05. **P < 0.01. ***P < 0.001, ****P < 0.0001
Wnt signaling is inactivated by ROS at the AST tipping point
We next sought to identify those specifically dysregulated genes across the tipping point (7 W). We found that the DNB molecules were significantly enriched for the Wnt signaling pathway (Fig. 2d). Moreover, this pathway was also enriched in the differential genes before and after 7 W (Supplementary Fig. 1b). Interestingly, we observed a clear fluctuation of the Wnt signaling activity across the tipping point, after which its activity endured a sudden decline (Supplementary Fig. 1c, d). From these TP-induced genes, we found a significant enrichment of known squamous lineage TFs such as TP63 and SOX2 (Supplementary Fig. 1e). From the TP-repressed genes, known adenomatous lineage TFs such as NKX2-1 and FOXA2 were significantly enriched (Fig. 2e). Such attenuation of adenomatous lineage TFs and simultaneous activation of squamous lineage TFs well corresponded to the critical transition of cellular fates across the tipping point. Moreover, we noticed that three TFs of the Wnt signaling pathway were also among the top dysregulated TFs (Fig. 2e). These data together indicate that the Wnt signaling might be involved in the TP regulation and lineage transition during AST.
We then investigated the potential mechanisms of the Wnt pathway dysregulation. Given previous reports that excessive accumulation of ROS is considered as an important factor to promote AST29 and oxidative stress is also known to inhibit the Wnt pathway,44,45,46 we tested the hypothesis that the abrupt inactivation of the Wnt signaling at 7 W might be due to high ROS level. As expected, ROS accumulated and peaked at 7 W after Ad-cre administration in KL model (Supplementary Fig. 1f). Through immunostaining of 7 W ADC from KL mouse model, we found that β-catenin positivity was negatively correlated with the level of 8-oxo-2’-deoxyguanosine (8-oxo-dGuo), the most common oxidative lesion observed in duplex DNA (Fig. 2f, g). Moreover, treatments with phenformin (PHEN), piperlongumine (PL), and phenethyl isothiocyanate (PEITC) in KrasG12D; Lkb1-/- (KL) mouse lung ADC cells consistently increased the ROS level (Supplementary Fig. 1g) and subsequently inhibited the Wnt signaling (Fig. 2h). Consistent with previous studies,44,47 these ROS inducer treatments significantly increased the FoxO3a gene expression (Supplementary Fig. 2a). We found that FOXO3A expression was positively correlated with 8-oxo-dGuo level at 7 W, indicating the upregulation of FOXO3A in response to oxidative stress (Supplementary Fig. 2b). Moreover, β-catenin level was negatively correlated with FOXO3A expression (Supplementary Fig. 2c). Consistently, we found that FoxO3a knockout activated the Wnt pathway as indicated by the upregulation of Ctnnb1 and Axin2 (Supplementary Fig. 2d–f). Moreover, FoxO3a knockout also significantly relieved the inhibition of the Wnt pathway triggered by multiple ROS inducers (Supplementary Fig. 2g). Consistent with a previous study about the de-regulation of fatty acid oxidation in KL SCC,29 we found that the ROS inducers promoted the accumulation of lipid-related ROS (Supplementary Fig. 2h).48 These data together demonstrate that the Wnt pathway is inactivated by upregulated FOXO3A under the condition of excessive accumulation of ROS.
Genetic perturbation validates an important role of the Wnt signaling in AST
To test the potential contribution of the Wnt pathway in AST, we performed bidirectional genetic perturbation in mouse models and analyzed their impacts upon the AST process. We first crossed the Ctnnb1flox/flox mice, in which the conditional deletion of the exons 2–6 inactivates Ctnnb1 and thus the Wnt pathway,49,50 with the KL mice to generate the KrasLSL-G12D/+; Lkb1flox/flox; Ctnnb1flox/flox (KLC) cohort. At 8 weeks post Ad-Cre administration, we found a striking increase of SCC development in the KLC mice (Fig. 3a–c). As expected, a negative staining of β-catenin was observed in KLC mouse tumors (Fig. 3d). All KLC mice (100%, 12/12) developed SCC, in contrast to only 17% (4 of 24) incidence of squamous transdifferentiation in the KL mice (Fig. 3e). The Ctnnb1 deletion clearly shifted the tumor transdifferentiation into SCC, with a notable decrease of ADC number (Fig. 3f). These data support an important role of the Wnt inactivation in promoting squamous transdifferentiation.
Inactivation of the Wnt pathway promotes the AST process in KL mouse model. a The Kras LSL-G12D/+; Lkb1flox/flox (KL, n = 24) and Kras LSL-G12D/+; Lkb1flox/flox; Ctnnb1flox/flox (KLC, n = 12) mice were given Ad-Cre via nasal inhalation and histologically analyzed at 8 weeks afterwards. b Representative H&E staining on KL and KLC mouse lung tumors. SCC lesions were indicated. Scale bar: 1 mm. c Representative H&E staining and p63 immunohistochemical staining on KL and KLC mouse lung tumors. SCC lesions were indicated. Scale bar: 500 µm. d Representative β-catenin immunohistochemical staining on KL and KLC mouse lung tumors. Scale bar: 50 µm. e SCC incidence of KL and KLC mice at 8 weeks post Ad-Cre treatment. Tumor No. = Tumor number, w SCC = with SCC, w/o SCC = without SCC. f. Average ADC and SCC numbers of KL and KLC mice at 8 weeks post Ad-Cre treatment. ****P < 0.0001
We further investigated whether constitutive activation of the Wnt pathway would inhibit the AST process. For this, we crossed the Ctnnb1(E3)flox/flox mice, in which the exon3 of β-catenin can be conditionally deleted through Ad-Cre administration for persistent β-catenin nuclear retention and thus the Wnt signaling activation,51,52,53,54 with the KL mice to generate the KrasLSL-G12D/+; Lkb1 flox/flox; Ctnnb1(E3)flox/flox (KLE) cohort for subsequent analyses (Fig. 4a). Histopathological analyses showed that constitutive Wnt activation resulted in a significant decrease of SCC number and incidence in the KLE mice (Fig. 4b–e). With the decrease of SCC number, we observed a slight increase of ADC number (Fig. 4f). Taken together, these data support a critical inhibitory role of the Wnt pathway in orchestrating the AST process.
Constitutive activation of the Wnt pathway inhibits the AST process in KL mouse model. a The Kras LSL-G12D/+; Lkb1flox/flox (KL) and Kras LSL-G12D/+; Lkb1flox/flox; Ctnnb1(E3)flox/flox (KLE) mice (n = 12 for each group) were given Ad-Cre via nasal inhalation and histologically analyzed at 10 weeks afterwards. b Representative H&E staining on KL and KLE mouse lung tumors. ADC and SCC lesions were indicated. Scale bar: 1 mm. c Representative H&E and p63 immunohistochemical staining on KL and KLE mouse lung tumors. ADC and SCC lesions were as indicated. Scale bar: 500 µm. d Representative β-catenin immunohistochemical staining on KL and KLE mouse lung tumors. Scale bar: 50 µm. e SCC incidence of KL and KLE mice at 10 weeks post Ad-Cre treatment. Tumor No. = Tumor number, w SCC = with SCC, w/o SCC = without SCC. f Average ADC and SCC numbers of KL and KLE mice at 10 weeks post Ad-Cre treatment. ***P < 0.001, ****P < 0.0001
Pharmacological activation of Wnt signaling inhibits AST only before the tipping point
According to the property of the tipping point before a critical transition and the hypothesis justified above, an early blockade of the Wnt signaling attenuation before the tipping point (at a reversible state) would effectively inhibit the AST, whereas a late perturbation after the tipping point (at an irreversible state) might not have such a significant effect. To test this hypothesis, we took advantage of lithium chloride (LiCl), a known β-catenin agonist that activates the Wnt signaling through GSK3 inhibition, to treat KL mice at 6 W (before TP) and 10 W (after TP) post Ad-Cre administration (Fig. 5a and Supplementary Fig. 3a). Consistently, we found that LiCl treatment resulted in a significant increase of nuclear β-catenin level (Fig. 5b). Moreover, we found that the early treatment with LiCl (at 6 W) resulted in a notable decrease in both SCC incidence and number (Fig. 5c–e). In stark contrast, the late treatment of LiCl showed almost no impact on SCC incidence and number (Supplementary Fig. 3b, c). We observed a slight increase of average ADC number after late LiCl treatment, presumably due to the promotive function of Wnt signaling upon lung ADC.55,56,57,58 These data together demonstrate the importance of the Wnt signaling perturbation in regulating AST process and highlight the irreversibility of AST after crossing tipping point.
LiCl treatment before the tipping point significantly inhibits the AST process in KL mouse model. a A scheme of LiCl treatment before the tipping point in KL mice (Veh: control group, n = 14, LiCl: LiCl treatment group, n = 9). b Representative β-catenin immunohistochemical staining on KL mouse lung tumors with or without LiCl treatment. Scale bar: 50 µm. c Representative H&E staining and p63 immunohistochemical staining on KL mouse lung tumors with or without LiCl treatment. ADC and SCC lesions were indicated. Scale bar: 500 µm. d Quantification of SCC incidence of KL mice with or without LiCl treatment at 6 weeks post Ad-Cre treatment. Tumor No. = Tumor number, w SCC = with SCC, w/o SCC = without SCC. e Quantification of average tumor numbers in KL mice with or without LiCl treatment at 6 weeks post Ad-Cre treatment. ***P < 0.001, ****P < 0.0001
Mutual suppressive lineage-specific transcription factor network between ADC and SCC
To decipher the mechanisms involved in TP-related Wnt signaling in the context of AST process, we explored the potential transcriptional modulation involved in this lineage transition. We first computationally analyzed the key transcriptional factor networks in contribution to ADC and SCC development using murine transcriptomic dataset categorized by ADC or SCC phenotype. We found that the ADC transcription factor network was composed of those known ADC lineage factors Nkx2-1, Foxa2, etc. (Fig. 6a). In contrast, the SCC transcriptional factor network was featured with key TFs such as p63, Sox2, etc. (Fig. 6a). Notably, two lineage-maintenance regulatory programs existed: the adenomatous program underlying the ADC differentiation, and the squamous program underlying the SCC differentiation. We found these two TF programs were counteracting and exerted their function in a mutual suppressive manner (Fig. 6a). The ADC-lineage TF program specifically drove the induction of genes that were actively tuned down by the SCC-lineage TF program (P = 1.28e−28), and vice versa (P = 4.24e−71) (Fig. 6a). Using the TCGA datasets, we further found that such counteracting pattern between ADC and SCC-lineage-specific TF programs also existed in human lung cancer (Supplementary Fig. 4a). Moreover, these ADC or SCC-lineage-specific TFs themselves also showed a direct mutual suppressive pattern at transcriptional levels (Supplementary Fig. 4b). These data together uncover a “Yin-Yang” counteracting pattern between mouse and human ADC and SCC TF networks which mutually suppress each other as well as their downstream target genes.
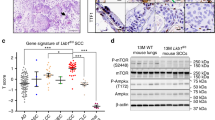
The Wnt-NKX2-1 axis regulates the AST process at the tipping point. a Mutually suppressive model of ADC and SCC lineage-specific TF networks based on computation analyses. The significance of overlapped target genes was evaluated with Fisher’s exact test. b ChIP-seq data showing the β-catenin binding peak located upstream of human NKX2-1 allele. β-cat: β-catenin. NCBI SRA accessions: SRX1036445 and SRX833403. c A total of 93 human lung adenosquamous carcinoma were used for RNA-seq analyses and aligned individually from the ADC-like state to the SCC-like state based on the AST score. The AST score was defined by the difference between SCC signature and ADC signature based on the TCGA dataset. The correlation between AST score and indicated gene signature was calculated. d Mouse KrasG12D; Lkb1-/- lung cancer cells were treated with LiCl (10 mM) for 24 h and the relative gene expression of Ctnnb1 and Nkx2-1 was assessed by real-time PCR. ***P < 0.001. e H&E staining of mouse lung tumors from KLC mice at 13 weeks post Lenti-Cre-Nkx2-1 nasal inhalation. Scale bar: 1 mm. Ctrl: KLC mice (Ctrl: n = 16, Nkx2-1: n = 12). f Representative photos of H&E staining, NKX2-1 and p63 immunohistochemical staining on mouse lung tumors from KLC mice post 13 weeks of Lenti-Cre or Lenti-Cre-Nkx2-1 nasal inhalation. Scale bar: 50 µm. Ctrl: KLC mice. g SCC incidence and average tumor numbers in KLC mice post 13 weeks of Lenti-Cre or Lenti-Nkx2-1 nasal inhalation. ****P < 0.0001, n.s. not significant. Ctrl: KLC mice. Tumor No. = Tumor number, w SCC = with SCC, w/o SCC = without SCC. h Mechanistic model of AST. In this model, the dynamic homeostasis between ADC and SCC lineage-specific transcriptional factors is finely tuned by the Wnt signaling, which maintains the ADC state. At the AST tipping point, excessive ROS accumulation abruptly suppresses the Wnt signaling as mediated by FOXO3A, and leads to the destabilization of adenomatous TF program and eventually push towards the activation of squamous TF program
A mechanistic model for AST orchestrated by the Wnt signaling
We reasoned that the adenomatous program should be switched to the antagonistic squamous program during the AST process and hypothesized that the Wnt signaling might help maintain the adenomatous state. To test this, we first analyzed the ChIP-seq data of β-catenin. Clearly, we can detect the enrichment of the β-catenin binding motif in the gene promoter of the adenomatous lineage factor NKX2-1 (Fig. 6b). To further solidify this conclusion, we collected a cohort of 93 Chinese lung adenosquamous carcinomas and performed transcriptomic analyses. Although these samples were pathologically confirmed to contain both adenomatous and squamous lesions, some specimens might contain more ADC components whereas others contain more SCC components due to the bias of sample collection. To overcome this issue, we defined an AST score by quantifying the proximity towards the squamous state. Using this score, we could align these samples individually from the ADC-like state to the SCC-like state. Clearly, our analyses showed that the ADC lineage TFs such as NKX2-1 and FOXA2 were significantly negatively correlated with the AST score whereas the TP63 and SOX2, two SCC lineage TFs, were opposite (Fig. 6c). Importantly, we found that the signatures for the Wnt signaling were negatively correlated with the AST score (Fig. 6c), similar to the NKX2-1 signature. These data indicate the potential regulation of the Wnt pathway on NKX2-1 gene transcription.
To further solidify this, we treated mouse KL lung ADC cells with LiCl. We found that the activation of the Wnt pathway indeed promoted the Nkx2-1 gene expression (Fig. 6d and Supplementary Fig. 5a). Treatment with ICG-001, a β-catenin inhibitor,59,60,61,62,63 significantly down-regulated Nkx2-1 expression (Supplementary Fig. 5b, c). Moreover, the inactivation of the Wnt pathway using Ctnnb1 knockout significantly decreased Nkx2-1 expression (Supplementary Fig. 5d–i). To investigate whether Nkx2-1 indeed regulates AST in context with the Wnt signaling, we delivered the lentivirus carrying the expression of Nkx2-1 and Cre (Lenti-Nkx2-1-Cre) into the KLC mice (Supplementary Fig. 4c). We found that, in contrast to control mice, ectopic NKX2-1 expression obviously inhibited the squamous transdifferentiation (Fig. 6e, f). Only about 33% (4/12) mice showed SCC in comparison to 100% SCC incidence in control mice (Fig. 6g). Moreover, the majority of lesions from the Nkx2-1 group were ADC (Fig. 6e–g). Consistently, the SCC number was also reduced in the Nkx2-1 group (Fig. 6g). These data demonstrate that ectopic Nkx2-1 expression inhibits the AST process accelerated by the Wnt pathway inactivation.
Collectively, we propose a mechanistic model for AST in KL mouse model (Fig. 6h). The ADC and SCC differentiation states are respectively controlled by the adenomatous TF program (e.g., NKX2-1) and the squamous TF program (e.g., TP63). The Wnt signaling functions as a stabilizer of the adenomatous TF program and favors the adenomatous differentiation through the activation of NKX2-1. In the KL mice, excessive accumulation of ROS elicits a FOXO3A-mediated switch-off of the Wnt/β-catenin signaling at the tipping point (7 W), leading to the destabilization of the adenomatous TF program which in turn activates the squamous TF program. After a short fluctuation period near the tipping point, the squamous TF program wins out and the system shifts to the squamous differentiation state.
Discussion
Lung cancer is a highly malignant cancer with strong plasticity. Several clinically important phenomena, such as the occurrence of lung AdSCC and relapse-associated AST, have been consistently detected but remain poorly understood. Using the well-established AST mouse model, we here extensively characterize the temporal transcriptomic dynamics of lung tumors during the AST process from a network perspective and uncover the tipping point. We further identify the Wnt signaling as the critical regulator at the tipping point and propose an ADC and SCC-lineage specific TFs mutual suppressive model. Our previous study has shown that excessive ROS accumulation serves as an important upstream trigger for AST.29 However, it remains unknown how this strong ROS level drives AST. Our data demonstrate that excessive ROS indeed results in an abrupt inactivation of the Wnt signaling, which subsequently disrupts the mutually suppressive homeostasis between ADC and SCC-lineage specific TF network via tuning down NKX2-1 level and eventually promotes the squamous transdifferentiation.
Critical transitions and identification of the tipping point, as exemplified in physiological systems, ecological systems, climate systems, and social systems, have become a useful framework in system state monitoring and perturbation.31,33,34,35,37,64 According to the theory of dynamical systems, the system is very difficult to shift back to its original state once it goes across the tipping point. However, when approaching the tipping point, a very small force might be sufficient to push the system forward. Theoretically, early perturbations before the TP are considered to be important. Of course, a better mechanistic understanding of the tipping point would be largely desired. In biological systems, a network-based DNB has been shown to be effective in the identification of tipping points based on high-dimensional data.36,38,39,40,41 In this study, we have applied the DNB method in a series of temporal transcriptomic data and uncovered the critical tipping point as the 7th week in KL AST mouse model. Our LiCl treatment experiments also prove the irreversibility of the tipping point during the AST process. Interestingly, the critical behavior at the tipping point between the adenomatous and the squamous states is like “Yin” and “Yang”, a pair of key categories borrowed from the classical Chinese literature that vividly signifies counteractions and inter-transitions. Indeed, we find that ADC and SCC are characterized by two mutually suppressive TF network and this homeostasis state is disrupted towards the tipping point of AST. Restoring this subtle perturbation before the tipping point could largely prevent the AST process. Besides the KL mouse model, other models have also been established for investigating the AST event. For example, the proportion of mice with DNp63+ tumor cells increase over time, with 33% at 4 weeks post induction and 100% at 12 weeks in the Rosa26LSL-Sox2-IRES-GFP;Nkx2-1flox/flox; Lkb1flox/flox (SNL) mouse model.65 They point out that NKX2-1 loss accelerates adeno-to-squamous transdifferentiation. Similarly, sequential Kras activation and Nkx2-1/Foxa1/2 deletion resulted in squamocolumnar-junction-like tumors as well as discrete SCC lesions in mice.66 Their data show that KRASG12D activated SPC-positive cells have the potential to undergo full squamous transdifferentiation (CK7-negative/CK5-positive) and become well-differentiated keratinizing SCCs. In our study, the positive regulation of Wnt on Nkx2-1 is essential to stabilize the ADC state. It would be interesting to investigate whether Wnt is the regulation of Nkx2-1in above models. Our analyses of 93 human lung adenosquamous carcinomas showed that the signatures for the Wnt signaling were negatively correlated with the AST score, similar to the NKX2-1 signature. Based on this, we believe that the consistent patterns of the positive regulation of Wnt on NKX2-1 would be applied to other mouse models. Future studies will be interesting to check this link between Wnt and AST in other emerging models. Another important extension is in the clinical setting of drug-resistance-related AST,15,16,18 which is theoretically and practically important for better understanding the AST process and provides ways for early intervention. Nonetheless, we believe this work would be an important early step in elucidating the tipping point and related mechanism during the AST process, and might provide useful clues for future pharmacological intervention.
Our study has also revealed an unprecedented role for the Wnt signaling in regulating cancer plasticity at the tipping point of AST. The Wnt signaling is well known as a developmental regulatory pathway, where it modulates cellular fates in many organs including the lung.67,68,69 The Wnt signaling has also been linked to diseases including multiple types of cancers, such as colon cancer,70 liver cancer,71 prostate cancer72 and lung cancer.73 It has been long proposed that the Wnt signaling maintains cancer stemness.74,75,76 In lung ADC mouse models, the Wnt signaling is often beneficial to tumor growth. Intriguingly, we find that the Wnt signaling can actually specify various cellular fates in the context of lung ADC through stabilizing the adenomatous differentiation state. This provides a very primitive clue that embryonic cellular fate decisions and cancer cell identity might actually be controlled by shared molecular mechanisms. Moreover, the mutually repressive model orchestrated by the Wnt signaling constitutes a transcriptional circuit, which is also highly comparable to classical developmental circuits such as the pluripotency network in stem cells.77,78 Wnt signaling is the most significantly enriched pathway in the DNB module whereas other enriched pathways such as Hedgehog pathway79 and amino acid metabolism pathways80 might also regulate the tipping point. Consistent with our previous studies,28,29,44,47 excessive ROS accumulation, e.g., lipid-related ROS, remains as an upstream trigger for such phenotypic transition mediated by the abrupt inactivation of the Wnt signaling and ADC lineage TF dysfunction. Consistent with previous studies,44,47 FOXO3A appears to be a major factor involved in the ROS-regulated Wnt pathway dynamics.
It’s worth noting that a recent study on squamous transitioned lung cancer indicates a potential role of Wnt signaling in promoting AST,20 e.g., Wnt pathway is upregulated in transitioned lung cancer after Osimertinib resistance. Since this study mainly compares the Wnt pathway between two time points, before and after TKI resistance, it might not be able to uncover the dynamic changes of Wnt signaling. In contrast, we take advantage of the RNA-Seq data from mouse lung tumors at a series of time points during AST process for the pathway analyses. This allows us to detect the dynamic and delicate fluctuations of defined pathway. Our study reaches the consensus that the Wnt signaling is transiently disrupted at the tipping point of AST and such delicate fluctuation is further proven to be important for the AST accomplishment in GEMM. Taken together, our current study has uncovered the signaling importantly for orchestrating the tipping point before AST. Future efforts are necessary to elucidate the whole picture of transcriptional and epigenetic pathways dynamically involved in the AST process.
The epigenetic regulatory mechanism underlying the AST process is another important yet less explored topic. We and collaborators have previously revealed the contribution of Polycomb Repressive Complex 2 (PRC2) in regulating the AST process in KL mouse model.23 The chromatin analysis reveals the loss of H3K27me3 and gain of H3K27ac and H3K4me3 at squamous lineage genes, including Sox2, ΔNp63 and Ngfr. Our current work shows an important role of the Wnt signaling in regulating the expression of the adenomatous lineage gene Nkx2-1. Moreover, LKB1 inactivation not only promotes AST but also contributes to lung cancer metastasis, which is like two faces of LKB1. LKB1-deficient lung tumors have a SOX17-mediated epigenetic reprogramming to substantiate metastasis.81 Such metastatic regulation involves epigenetic mechanisms through modulating chromatin accessibility and enhancing SOX17-targeted transcription. This study has implications for the epigenetic modulation of metastasis in LKB1-deficient lung tumors. Whether such epigenetic modulation contributes to AST would require future studies.
Materials and methods
Ethical approval
The research performed in the present study complies with all ethical regulations. Clinical studies were approved by the institutional review board of Shanghai Cancer Hospital, Fudan University. Written informed consent was obtained from all patients (ethical approval no. IRB 2008223-9). Mice were housed in a specific-pathogen-free environment at the Shanghai Institute of Biochemistry and Cell Biology, and treated in accordance with protocols conforming to the guidelines and approved by the Institutional Animal Care and Use Committee of the Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences (ethical approval no. IBCB0011).
Mouse cohorts
Kras LSL-G12D/+, Lkb1flox/flox, Ctnnb1flox/flox and Ctnnb1(E3)flox/flox mice were originally generously provided by Drs. Tyler Jacks, Ronald A. DePinho, and Lijian Hui, respectively. After Ad-Cre administration, the Ctnnb1flox/flox mice had the deletion of exon 2 to 6 and thus the inactivation of the Wnt signaling.49,50 Similarly, the Ctnnb1 (E3)flox/flox mice had exon 3 deletion after Ad-Cre administration, which resulted in persistent nuclear retention of β-catenin and thus the constitutive activation of the Wnt signaling.51,52,53,54 All mice were housed in a pathogen-free environment at Shanghai Institute of Biochemistry and Cell Biology and treated in strict accordance with protocols approved by the Institutional Animal Care and Use Committee of the Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences. Mice were treated with adenovirus or lentivirus carrying Cre recombinase (2 × 106 pfu) via nasal inhalation as previously described.27 For temporal transcriptomic analyses, the Kras LSL-G12D/+; Lkb1flox/flox (KL) mice were given Ad-Cre through nasal inhalation and sacrificed at a series of time points (0 W, 4 W, 6 W, 7 W, 8 W, 9 W, 10 W) for tumor analyses. The Kras LSL-G12D/+; Lkb1flox/flox; Ctnnb1flox/flox (KLC) mice and control mice were analyzed at 8 weeks post Ad-Cre nasal inhalation. The Kras LSL-G12D/+; Lkb1 flox/flox; Ctnnb1(E3)flox/flox (KLE) mice and control mice were analyzed at 10 weeks post Ad-Cre nasal inhalation. The KLC mice were also given Lenti-Nkx2-1-Cre and analyzed at 13 weeks afterwards. For pharmacological treatment, the KL mice were treated with LiCl (500 mg/kg) via intraperitoneal injection every two days for 4 weeks at 6 weeks or 10 weeks post Ad-Cre administration.
Lentivirus production and infection
The ORFs of Nkx2-1 and N-terminal truncated β-catenin were amplified from mouse tumor cDNAs, and ligated into expression vectors pCDH-EF1-Puro (Systems Biosciences) or pCDH-EF1-Cre (adapted vector for Nkx2-1 expression in vivo). The production of lentivirus supernatant and cell infection were described previously.27
Histology and immunohistochemistry
Mouse lungs were inflated with 1 ml 4% paraformaldehyde (PFA), fixed overnight and dehydrated in ethanol, embedded in paraffin, sectioned at 5 μm followed by staining with haematoxylin and eosin (H&E). Photos were taken using Olympus BX53 and Axio Scan.Z1 (Carl Zeiss IMT Co. Ltd.) and tumor numbers were measured using Image J software. The numbers of ADC and.SCC were counted as previously described.82 Immunohistochemistry (IHC) was performed as previously described.27 The IHC scores of FOXO3A, β-catenin and 8-oxo-dGuo expression were measured as previously described.83 Briefly, staining intensities were assigned into 4 grades (intensity score) as follows: 0, negative; 1, low; 2, medium; and 3, high. The IHC score was the sum of intensity score multiplied by percentage. The following antibodies were used: β-catenin (8480, Cell Signaling Technology, 1:500), p63 (ab124762, abcam, 1:1000), SOX2 (ab92494, abcam, 1:500), NKX2-1 (ab133638, abcam, 1:500), SFTPC (AB3786, Millipore, 1:1000), 8-oxo-dGuo (ab48508, abcam, 1:250), FOXO3A (12829, Cell Signaling Technology, 1:250).
Cell lines and drug treatment
The KrasG12D;Lkb1-/- (KL) lung cancer cell line was established as previously described29 and cultured in DMEM (Hyclone) supplemented with 10% FBS (Biochrom). For LiCl treatment, cells were exposed to LiCl (10 mM) for 24 h and total RNA was extracted for real-time PCR analyses. For ICG-001 treatment, cells were exposed to ICG-001 (1.5 μM) for 48 h and total RNA was extracted for real-time PCR analyses. ICG-001 was purchased from MedChemExpress (HY-14428) and diluted in Dimethyl Sulfoxide (DMSO). For ROS inducer treatment, cells were exposed to PHEN (500 μM) or PL (12.5 μM) or PEITC (15 μM) for 6 h and the total RNA was extracted for real-time PCR analyses. Based on 3-(4, 5-Dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) results, we used the indicated concentration.
RNA extraction and real-time PCR analyses
Mouse lung tumors were freshly dissected at indicated time points post Ad-Cre treatment. Total RNA and genomic DNA from mouse lung tumors and cell lines were prepared as previously described.27 Briefly, total RNA was extracted from cells using Trizol reagent (Invitrogen) and reverse transcribed with the PrimeScript RT Reagent Kit (Takara). The cDNAs were then used for real-time PCR on a LightCycler®480/96 Real-Time PCR System (Roche) using SYBR-Green Master PCR mix (Toyobo). β-actin served as internal control. The primers used for real time PCR were:
Mouse β-Actin forward: 5′-TGAGCGCAAGTACTCTGTGTGGAT-3′
Mouse β-Actin reverse: 5′-ACTCATCGTACTCCTGCTTGCTGA-3′
Mouse Ctnnb1 forward: 5′-ATGGAGCCGGACAGAAAAGC-3′
Mouse Ctnnb1 reverse: 5′-TGGGAGGTGTCAACATCTTCTT-3′
Mouse Nkx2-1 forward: 5′-CAGCGCTTCGGGCCCCGGAT-3′
Mouse Nkx2-1 reverse: 5′-AGCGAGCCCAGGCCGCCCAT-3′
Mouse Axin2 forward: 5′-ATGAGTAGCGCCGTGTTAGTG-3′
Mouse Axin2 reverse: 5′-GGGCATAGGTTTGGTGGACT-3′
Mouse Ccnd1 forward: 5′-GCGTACCCTGACACCAATCTC-3′
Mouse Ccnd1 reverse: 5′-ACTTGAAGTAAGATACGGAGGGC-3′
Mouse c-Myc forward: 5′-ATGCCCCTCAACGTGAACTTC-3′
Mouse c-Myc reverse: 5′-GTCGCAGATGAAATAGGGCTG-3′
Mouse FoxO3a forward: 5′-AACGGCTCACTTTGTCCCA-3′
Mouse FoxO3a reverse: 5′-TTGATGATCCACCAAGAGC-3′.
Human lung cancer specimen collection
A total of 93 human lung adenosquamous carcinomas were collected from 2007 to 2016 with the approval of the institutional review board of Shanghai Cancer Hospital, Fudan University. All patients gave written informed consent. Samples were snap-frozen in liquid nitrogen at the time of resection and stored at −80 °C until the mRNA extraction.
RNA-seq analyses for mouse and human tumor samples
RNA extraction, reverse transcription, library preparation, and quality control were performed following the Illumina standard protocol. Sequencing of the libraries was performed on an Illumina HiSeq sequencing machine, generating paired-end reads (150 bases x 2). Clean reads were obtained after trimming adapters and filtering low-quality reads. The clean reads were mapped to reference genomes (mm10 and hg38) with the STAR aligner and subsequently quantified with the htseq-count tool. A normalization factor was estimated using the RLE method as implemented in edgeR. After normalizing the effective library size, the expression data in reads per million (RPM) was further log2 transformed for downstream analyses.84
Temporal transcriptomic analyses
The temporal transcriptomic data were dimensionally reduced with the principal component analyses (PCA) method. The components were ranked with proportions of variances explained and the first two principal components were used for visualization of the temporal trajectory. Relative expression of selected marker genes was mapped onto the two-dimensional PCA map to examine their respective fluctuations along the trajectory.
Dynamic network biomarker and tipping point analyses
The dynamic network biomarker (DNB) framework was used to delineate the network dynamics for the tipping point analyses. Briefly, we unraveled the network dynamics within a subspace constrained by the differential genes (n = 3092) between the adenomatous and squamous lineages, which are computed from the TCGA cohorts by fitting genewise negative binomial models and the quasi-likelihood F-tests, as implemented in the edgeR package. For each time point with replicates, an estimate of Pearson’s correlation coefficient (PCC) was performed for every gene pair. Absolute values of PCCs were taken as edge strength in these temporal networks. Network communities were detected with the deterministic hierarchical clustering algorithm. As shown by the DNB method, when the system state approaches a tipping point, a group of genes or DNB (module) would emerge, which satisfies the following three conditions simultaneously: (1) the average absolute correlation of any two genes in the group (PCCi) increases; (2) the average absolute correlation between genes in and outside of this group (PCCo) decreases; (3) the average standard deviation (SDi) highly increases in the group. The composite index (CI) of criticality was thus a combination of these three criteria:36,40,85
The module with maximum CI score was chosen as the dominant group of genes, with its CI score depicting the dynamic changes of system criticality, and the time point of maximum CI considered as a reasonable approximation of the tipping point. This theoretical result means that the appearance of a group of collectively fluctuating genes among the observed high-dimensional data signals an imminent critical transition, or briefly, strongly collective fluctuation implies an imminent state transition from one state to another. For a full derivation, simulation and subsequent validations of the DNB method for critical analyses, please refer to the previous publications.36,38,39,40,41,86
Analyses of the Wnt signaling and transcriptional signatures
The Wnt signaling pathway and β-catenin signatures were obtained from MSigDB (v6.0), which are the upregulated signatures trained from two experimental datasets (“WNT UP.V1”, “FEVR CTNNB1 TARGETS”). The signature scores are computed using the z-score method implemented in GSVA (v1.26.0).
Transcriptional targets for TP63, SOX2, NKX2-1 and FOXA2 in three mammalian species (human, mouse and rat) were compiled from IPA (Ingenuity Pathway Analysis, Qiagen). For each TF, the expression of the target genes was used to compute the signature score using the z-score method from GSVA (v1.26.0).
The AST signature was based on top differential genes for ADC and SCC computed using the quasi-likelihood method from edgeR v3.20.9 (fold change > 2 and adjusted P value < 0.001). An ADC score was calculated as the average of standardized expression for genes upregulated in ADC. An SCC score is similarly calculated using the genes upregulated in SCC. The final AST score is defined as the difference between the two scores in each sample (AST score = SCC score−ADC score).
Statistical analyses
Differences were compared using the two-tailed Student’s t test, with P value < 0.05 considered statistically significant. All analyses were performed with Graphpad Prism 7.
Data availability
The mouse KL lung cancer and human AdSCC datasets have been deposited in the NODE database (project accessions OEP002019 and OEP001032). The materials used in this study are available from the corresponding authors upon reasonable request.
References
Bass, A. J. et al. SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nat. Genet. 41, 1238–1242 (2009).
Dotto, G. P. & Rustgi, A. K. Squamous cell cancers: a unified perspective on biology and genetics. Cancer Cell. 29, 622–637 (2016).
Ferone, G. et al. SOX2 is the determining oncogenic switch in promoting lung squamous cell carcinoma from different cells of origin. Cancer Cell. 30, 519–532 (2016).
Li, C. M.-C. et al. Foxa2 and Cdx2 cooperate with Nkx2-1 to inhibit lung adenocarcinoma metastasis. Genes Dev. 29, 1850–1862 (2015).
Massion, P. P. et al. Significance of p63 amplification and overexpression in lung cancer development and prognosis. Cancer Res. 63, 7113–7121 (2003).
Snyder, E. L. et al. Nkx2-1 represses a latent gastric differentiation program in lung adenocarcinoma. Mol. Cell. 50, 185–199 (2013).
Weir, B. A. et al. Characterizing the cancer genome in lung adenocarcinoma. Nature 450, 893 (2007).
Maeda, Y., Davé, V. & Whitsett, J. A. Transcriptional control of lung morphogenesis. Physiol. Rev. 87, 219–244 (2007).
Tata, P. R. et al. Developmental history provides a roadmap for the emergence of tumor plasticity. Dev. Cell. 44, 679–693.e675 (2018).
Melek, H. et al. P2.16-33 adenosquamous carcinoma of the lung: comparison of surgical outcomes with squamous cell and adenocarcinoma. J. Thorac. Oncol. 13, S844–S845 (2018).
Maeda, H. et al. Adenosquamous carcinoma of the lung: surgical results as compared with squamous cell and adenocarcinoma cases. Eur. J. Cardiothorac. Surg. 41, 357–361 (2012).
Krause, A. et al. Deciphering the clonal relationship between glandular and squamous components in adenosquamous carcinoma of the lung using whole exome sequencing. Lung Cancer 150, 132–138 (2020).
Lin, G. et al. Genomic origin and EGFR-TKI treatments of pulmonary adenosquamous carcinoma. Ann. Oncol. 31, 517–524 (2020).
Wang, R. et al. Analysis of major known driver mutations and prognosis in resected adenosquamous lung carcinomas. J. Thorac. Oncol. 9, 760–768 (2014).
Hou, S. et al. Evidence, mechanism, and clinical relevance of the transdifferentiation from lung adenocarcinoma to squamous cell carcinoma. Am. J. Pathol. 187, 954–962 (2017).
Hou, S., Han, X. & Ji, H. Squamous transition of lung adenocarcinoma and drug resistance. Trends Cancer 2, 463–466 (2016).
Jiang, Y. et al. Adenocarcinoma transformed into squamous cell carcinoma in non-small cell lung cancer. J. Genet Genomics. 48, 656–659 (2021).
Schoenfeld, A. J. et al. Tumor analyses reveal squamous transformation and off-target alterations as early resistance mechanisms to first-line osimertinib in EGFR-mutant lung cancer. Clin. Cancer Res. 26, 2654–2663 (2020).
Awad, M. M. et al. Acquired resistance to KRAS(G12C) inhibition in cancer. N. Engl. J. Med. 384, 2382–2393 (2021).
Quintanal-Villalonga, A. et al. Comprehensive molecular characterization of lung tumors implicates AKT and MYC signaling in adenocarcinoma to squamous cell transdifferentiation. J. Hematol. Oncol. 14, 170 (2021).
Koivunen, J. P. et al. Mutations in the LKB1 tumour suppressor are frequently detected in tumours from Caucasian but not Asian lung cancer patients. Br. J. Cancer 99, 245–252 (2008).
Matsumoto, S. et al. Prevalence and specificity of LKB1 genetic alterations in lung cancers. Oncogene 26, 5911–5918 (2007).
Zhang, H. et al. Lkb1 inactivation drives lung cancer lineage switching governed by Polycomb Repressive Complex 2. Nat. Commun. 8, 14922 (2017).
Mahoney, C. L. et al. LKB1/KRAS mutant lung cancers constitute a genetic subset of NSCLC with increased sensitivity to MAPK and mTOR signalling inhibition. Br. J. Cancer 100, 370–375 (2009).
Sanchez-Cespedes, M. et al. Inactivation of LKB1/STK11 is a common event in adenocarcinomas of the lung. Cancer Res. 62, 3659–3662 (2002).
Ab Mutalib, N. S. et al. Molecular characterization of serous ovarian carcinoma using a multigene next generation sequencing cancer panel approach. BMC Res. Notes 7, 805 (2014).
Gao, Y. et al. YAP inhibits squamous transdifferentiation of Lkb1-deficient lung adenocarcinoma through ZEB2-dependent DNp63 repression. Nat. Commun. 5, 4629 (2014).
Han, X. et al. Transdifferentiation of lung adenocarcinoma in mice with Lkb1 deficiency to squamous cell carcinoma. Nat. Commun. 5, 3261 (2014).
Li, F. et al. LKB1 inactivation elicits a redox imbalance to modulate non-small cell lung cancer plasticity and therapeutic response. Cancer Cell. 27, 698–711 (2015).
Ji, H. et al. LKB1 modulates lung cancer differentiation and metastasis. Nature 448, 807–810 (2007).
Barnosky, A. D. et al. Approaching a state shift in Earth’s biosphere. Nature 486, 52–58 (2012).
Dakos, V. et al. Slowing down as an early warning signal for abrupt climate change. Proc. Natl Acad. Sci. USA 105, 14308–14312 (2008).
McSharry, P. E., Smith, L. A. & Tarassenko, L. Prediction of epileptic seizures: are nonlinear methods relevant? Nat. Med. 9, 241–242 (2003).
Scheffer, M. et al. Early-warning signals for critical transitions. Nature 461, 53–59 (2009).
Scheffer, M. et al. Anticipating critical transitions. Science 338, 344–348 (2012).
Chen, L. et al. Detecting early-warning signals for sudden deterioration of complex diseases by dynamical network biomarkers. Sci. Rep. 2, 342 (2012).
Scheffer, M. et al. Catastrophic shifts in ecosystems. Nature 413, 591–596 (2001).
Li, M., Zeng, T., Liu, R. & Chen, L. Detecting tissue-specific early warning signals for complex diseases based on dynamical network biomarkers: study of type 2 diabetes by cross-tissue analysis. Brief. Bioinforma. 15, 229–243 (2014).
Shi, J., Li, T. & Chen, L. Towards a critical transition theory under different temporal scales and noise strengths. Phys. Rev. E 93, 032137 (2016).
Yang, B. et al. Dynamic network biomarker indicates pulmonary metastasis at the tipping point of hepatocellular carcinoma. Nat. Commun. 9, 1–14 (2018).
Liu, R. et al. Identifying critical transitions of complex diseases based on a single sample. Bioinformatics 30, 1579–1586 (2014).
Steffen, W. et al. Planetary boundaries: guiding human development on a changing planet. Science 347, 1259855 (2015).
Walker, B., Holling, C. S., Carpenter, S. & Kinzig, A. Resilience, adaptability and transformability in social-ecological systems. Ecol. Soc. 9, 5 (2004).
Almeida, M. et al. Glucocorticoids and tumor necrosis factor α increase oxidative stress and suppress Wnt protein signaling in osteoblasts. J. Biol. Chem. 286, 44326–44335 (2011).
Ebrahimi, K. B. et al. Oxidative stress induces an interactive decline in Wnt and Nrf2 signaling in degenerating retinal pigment epithelium. Antioxid. Redox Signal. 29, 389–407 (2017).
Zhang, X. et al. Notum is required for neural and head induction via Wnt deacylation, oxidation, and inactivation. Dev. Cell. 32, 719–730 (2015).
Bartell, S. M. et al. FoxO proteins restrain osteoclastogenesis and bone resorption by attenuating H2O2 accumulation. Nat. Commun. 5, 3773 (2014).
Wang, X. et al. Stem cell factor SOX2 confers ferroptosis resistance in lung cancer via upregulation of SLC7A11. Cancer Res. 81, 5217–5229 (2021).
Brault, V. et al. Inactivation of the (β)-catenin gene by Wnt1-Cre-mediated deletion results in dramatic brain malformation and failure of craniofacial development. Development 128, 1253–1264 (2001).
Grigoryan, T., Wend, P., Klaus, A. & Birchmeier, W. Deciphering the function of canonical Wnt signals in development and disease: conditional loss- and gain-of-function mutations of β-catenin in mice. Genes Dev. 22, 2308–2341 (2008).
Harada, N. et al. Intestinal polyposis in mice with a dominant stable mutation of the β-catenin gene. EMBO J. 18, 5931–5942 (1999).
Okubo, T., Knoepfler, P. S., Eisenman, R. N. & Hogan, B. L. M. Nmyc plays an essential role during lung development as a dosage-sensitive regulator of progenitor cell proliferation and differentiation. Development 132, 1363–1374 (2005).
Reynolds, S. D. et al. Conditional stabilization of β-catenin expands the pool of lung stem cells. Stem Cells 26, 1337–1346 (2008).
Hashimoto, S. et al. β-Catenin–SOX2 signaling regulates the fate of developing airway epithelium. J. Cell Sci. 125, 932–942 (2012).
Tammela, T. et al. A Wnt-producing niche drives proliferative potential and progression in lung adenocarcinoma. Nature 545, 355–359 (2017).
Xu, Q. et al. Loss of TET reprograms Wnt signaling through impaired demethylation to promote lung cancer development. Proc. Natl Acad. Sci. USA. 119, e2107599119 (2022).
Juan, J. et al. Diminished WNT -> β-catenin -> c-MYC signaling is a barrier for malignant progression of BRAFV600E-induced lung tumors. Genes Dev. 28, 561–575 (2014).
Pacheco-Pinedo, E. C. et al. Wnt/β-catenin signaling accelerates mouse lung tumorigenesis by imposing an embryonic distal progenitor phenotype on lung epithelium. J. Clin. Invest. 121, 1935–1945 (2011).
Ye, Y. C. et al. NOTCH signaling via WNT regulates the proliferation of alternative, CCR2-independent tumor-associated macrophages in hepatocellular carcinoma. Cancer Res. 79, 4160–4172 (2019).
Yang, Y. et al. Crosstalk between hepatic tumor cells and macrophages via Wnt/β-catenin signaling promotes M2-like macrophage polarization and reinforces tumor malignant behaviors. Cell Death Dis. 9, 793 (2018).
Liu, L. et al. MTNR1B loss promotes chordoma recurrence by abrogating melatonin-mediated β-catenin signaling repression. J. Pineal Res. 67, e12588 (2019).
Liu, Y. et al. ICG-001 suppresses growth of gastric cancer cells and reduces chemoresistance of cancer stem cell-like population. J. Exp. Clin. Cancer Res. 36, 125 (2017).
Wang, L. et al. Oncolytic herpes simplex virus and PI3K inhibitor BKM120 synergize to promote killing of prostate cancer stem-like cells. Mol. Ther. Oncolytics 13, 58–66 (2019).
Gunderson, L. H. Ecological resilience—in theory and application. Annu Rev. Ecol. Syst. 31, 425–439 (2000).
Mollaoglu, G. et al. The lineage-defining transcription factors SOX2 and NKX2-1 determine lung cancer cell fate and shape the tumor immune microenvironment. Immunity 49, 764–779.e769 (2018).
Camolotto, S. A. et al. FoxA1 and FoxA2 drive gastric differentiation and suppress squamous identity in NKX2-1-negative lung cancer. eLife 7, e38579 (2018).
Li, C. et al. Stabilized β-catenin in lung epithelial cells changes cell fate and leads to tracheal and bronchial polyposis. Dev. Biol. 334, 97–108 (2009).
Moon, R. T., Kohn, A. D., Ferrari, G. V. D. & Kaykas, A. WNT and β-catenin signalling: diseases and therapies. Nat. Rev. Genet. 5, 691–701 (2004).
Nabhan, A. N. et al. Single-cell Wnt signaling niches maintain stemness of alveolar type 2 cells. Science 359, 1118–1123 (2018).
Clevers, H. Wnt/β-catenin signaling in development and disease. Cell 127, 469–480 (2006).
Perugorria, M. J. et al. Wnt–β-catenin signalling in liver development, health and disease. Nat. Rev. Gastroenterol. Hepatol. 16, 121–136 (2019).
Murillo-Garzón, V. & Kypta, R. WNT signalling in prostate cancer. Nat. Rev. Urol. 14, 683–696 (2017).
Rapp, J. et al. WNT signaling – lung cancer is no exception. Respir. Res. 18, 167 (2017).
Fodde, R. & Brabletz, T. Wnt/β-catenin signaling in cancer stemness and malignant behavior. Curr. Opin. Cell Biol. 19, 150–158 (2007).
Takebe, N. et al. Targeting Notch, Hedgehog, and Wnt pathways in cancer stem cells: clinical update. Nat. Rev. Clin. Oncol. 12, 445–464 (2015).
Zhan, T., Rindtorff, N. & Boutros, M. Wnt signaling in cancer. Oncogene 36, 1461–1473 (2017).
Kim, J. et al. An extended transcriptional network for pluripotency of embryonic stem cells. Cell 132, 1049–1061 (2008).
Ng, H.-H. & Surani, M. A. The transcriptional and signalling networks of pluripotency. Nat. Cell Biol. 13, 490–496 (2011).
Velcheti, V. & Govindan, R. Hedgehog signaling pathway and lung cancer. J. Thorac. Oncol. 2, 7–10 (2007).
Zhou, X. et al. Activated amino acid response pathway generates apatinib resistance by reprograming glutamine metabolism in non-small-cell lung cancer. Cell Death Dis. 13, 636 (2022).
Pierce, S. E. et al. LKB1 inactivation modulates chromatin accessibility to drive metastatic progression. Nat. Cell Biol. 23, 915–924 (2021).
Wang, X. et al. Targeting HSPA1A in ARID2-deficient lung adenocarcinoma. Natl Sci. Rev. 8, nwab014 (2021).
Zheng, Y. et al. Transmembrane protein 100 inhibits the progression of colorectal cancer by promoting the ubiquitin/proteasome degradation of HIF-1α. Front. Oncol. 12, 899385 (2022).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Zhang, C. et al. Landscape dynamic network biomarker analysis reveals the tipping point of transcriptome reprogramming to prevent skin photodamage. J. Mol. Cell Biol. 13, 822–833 (2021).
Fang, Z. & Chen, L. Personalized prediction of human diseases with single-sample dynamic network biomarkers. Biomark. Med. 14, 615–620 (2020).
Acknowledgements
We thank Drs. Tyler Jacks, Ronald A. DePinho, Kwok-kin Wong, and Lijian Hui for the generous gift of various mouse strains. We also thank Ruiqi Wang, Rui Liu, Pei Chen, Chao Zheng, and Jifan Shi for helpful discussion. This work was supported by the National Basic Research Program of China (Nos. 2017YFA0505500 to H.J. and L.C.; 2020YFA0803300 to H.J.); the National Natural Science Foundation of China (Nos. 91731314, 82030083, 31621003, 81872312, 82011540007 to H.J.; 12131020, 31930022, 12026608 to L.C.; 82273093 to Z.F.; 81871875, 82173340 to L.H., 81802279 to H.H., 81902326 to X.W., 81402371 to Y.J.); the Strategic Priority Research Program of the Chinese Academy of Sciences (Nos. XDB19020201 to H.J.; XDB38040400 to L.C.); Basic Frontier Scientific Research Program of Chinese Academy of Science (No. ZDBS-LY-SM006 to H.J.); International Cooperation Project of Chinese Academy of Sciences (No. 153D31KYSB20190035 to H.J.); the Science and Technology Commission of Shanghai Municipality (No. 21ZR1470300 to L.H.); the Youth Innovation Promotion Association CAS (No. Y919S31371 to X.W.); Special Fund for Science and Technology Innovation Strategy of Guangdong Province (Nos. 2021B0909050004, 2021B0909060002 to L.C.); Major Key Project of PCL (No.PCL2021A12 to L.C.); JST Moonshot R&D Project (No. JPMJMS2021 to L.C.).
Author information
Authors and Affiliations
Contributions
H.J. and L.C. designed and conceived the study. Z.F. performed computational analyses; X.H., Y.C., S.Y., and X.T. performed the experimental analyses; X.W., Y.J., L.H., L.L., and L.J.H. provided material and technical supports; Y.P., Y.S., and H.C. provided clinical samples and information. H.J., L.C., Y.C., and Z.F. wrote the manuscript. All authors have contributed to the manuscript. All authors have read and approved the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fang, Z., Han, X., Chen, Y. et al. Oxidative stress-triggered Wnt signaling perturbation characterizes the tipping point of lung adeno-to-squamous transdifferentiation. Sig Transduct Target Ther 8, 16 (2023). https://doi.org/10.1038/s41392-022-01227-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-022-01227-0