Abstract
Background
With continuous progress over the past few decades in understanding diagnosis, treatment, and genetics, much has been learned about the prostate cancer-diagnosed genome.
Methods
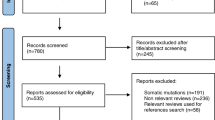
A comprehensive MEDLINE® and Google scholar literature search was conducted using keyword variations relating to the genetics of prostate cancer such as chromosomal alterations, androgen receptor, castration-resistant, inheritance, polymorphisms, oncogenes, metastasis, biomarkers, and immunotherapy.
Results
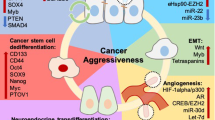
Traditionally, androgen receptors (AR) have been the focus of research. Recently, identification of recurrent chromosomal alterations that lead to either multiplication of regions (gain-of-function) or deletion of regions (loss-of-function) has opened the door to greater genetic accessibility. These chromosomal aberrations lead to variation in copy number and gene expression. Some of these chromosomal alterations are inherited, while others undergo somatic mutations during disease progression. Inherited gene mutations that make one susceptible to prostate cancer have been identified with familial-linked studies. Somatic genes that progress tumorigenesis have also been identified. Research on the molecular biology of prostate cancer has characterized these genes into tumor suppressor genes or oncogenes. Additionally, genome-wide assay studies have identified many high-risk single-nucleotide polymorphisms recurrent throughout the prostate cancer-diagnosed genome. Castration-resistant prostate cancer is the most aggressive form of prostate cancer, and its research has elucidated many types of mutations associated with AR itself, including enhanced expression and amplification, point mutations, and alternative splicing. Understanding the molecular biology of prostate cancer has permitted more accurate identification using advanced biomarkers and therapy for aggressive forms using immunotherapy.
Conclusions
An age-related disease, prostate cancer commands profound attention. With increasing life expectancy and the continuous pursuit of it, prostate cancer is a powerful obstacle best defeated using targeted therapies specifically designed for the unique molecular profile of the malignancy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 4 print issues and online access
$259.00 per year
only $64.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Packer JR, Maitland NJ. The molecular and cellular origin of human prostate cancer. Biochim Biophys Acta. 2016;1863:1238–60.
Mateo J, Carreira S, Sandhu S, Miranda S, Mossop H, Perez-Lopez R, et al. DNA-repair defects and olaparib in metastatic prostate cancer. N Engl J Med. 2015;373:1697–708.
Howlader NNA, Krapcho M, Miller D, Bishop K, Altekruse SF, Kosary CL, Yu M, Ruhl J, Tatalovich Z, Mariotto A, Lewis DR, Chen HS, Feuer EJ, Cronin KA, editors. SEER Cancer Statistics Review. Bethesda, MD: National Cancer Institute; 2016.
Hotte SJ, Saad F. Current management of castrate-resistant prostate cancer. Curr Oncol. 2010;17:S72–9.
Jakobsen NA, Hamdy FC, Bryant RJ. Novel biomarkers for the detection of prostate cancer. J Clin Urol. 2016;9:3–10.
Tomlins SA, Day JR, Lonigro RJ, Hovelson DH, Siddiqui J, Kunju LP, et al. Urine TMPRSS2:ERG plus PCA3 for individualized prostate cancer risk assessment. Eur Urol. 2016;70:45–53.
Ankerst DP, Hoefler J, Bock S, Goodman PJ, Vickers A, Hernandez J, et al. Prostate cancer prevention trial risk calculator 2.0 for the prediction of low- vs high-grade prostate cancer. Urology. 2014;83:1362–7.
Vedder MM, de Bekker-Grob EW, Lilja HG, Vickers AJ, van Leenders GJ, Steyerberg EW, et al. The added value of percentage of free to total prostate-specific antigen, PCA3, and a kallikrein panel to the ERSPC risk calculator for prostate cancer in prescreened men. Eur Urol. 2014;66:1109–15.
Pentyala S, Whyard T, Pentyala S, Muller J, Pfail J, Parmar S, et al. Prostate cancer markers: an update. Biomed Rep. 2016;4:263–8.
Platz EA, De Marzo AM. Epidemiology of inflammation and prostate cancer. J Urol. 2004;171:S36–40.
Walsh AL, Tuzova AV, Bolton EM, Lynch TH, Perry AS. Long noncoding RNAs and prostate carcinogenesis: the missing ‘linc’? Trends Mol Med. 2014;20:428–36.
Sun T, Ye H, Wu CL, Lee GS, Kantoff PW. Emerging players in prostate cancer: long non-coding RNAs. Am J Clin Exp Urol. 2014;2:294–9.
Crea F, Watahiki A, Quagliata L, Xue H, Pikor L, Parolia A, et al. Identification of a long non-coding RNA as a novel biomarker and potential therapeutic target for metastatic prostate cancer. Oncotarget. 2014;5:764–74.
Ferreira LB, Palumbo A, de Mello KD, Sternberg C, Caetano MS, de Oliveira FL, et al. PCA3 noncoding RNA is involved in the control of prostate-cancer cell survival and modulates androgen receptor signaling. BMC Cancer. 2012;12:507.
Hessels D, Schalken JA. Urinary biomarkers for prostate cancer: a review. Asian J Androl. 2013;15:333–9.
Prensner JR, Iyer MK, Sahu A, Asangani IA, Cao Q, Patel L, et al. The long noncoding RNA SChLAP1 promotes aggressive prostate cancer and antagonizes the SWI/SNF complex. Nat Genet. 2013;45:1392–8.
Mehra R, Udager AM, Ahearn TU, Cao X, Feng FY, Loda M, et al. Overexpression of the long non-coding RNA SChLAP1 independently predicts lethal prostate cancer. Eur Urol. 2016;70:549–52.
Filella X, Foj L. miRNAs as novel biomarkers in the management of prostate cancer. Clin Chem Lab Med. 2017;55:715–36.
Folini M, Gandellini P, Longoni N, Profumo V, Callari M, Pennati M, et al. miR-21: an oncomir on strike in prostate cancer. Mol Cancer. 2010;9:12.
Luu HN, Lin HY, Sorensen KD, Ogunwobi OO, Kumar N, Chornokur G, et al. miRNAs associated with prostate cancer risk and progression. BMC Urol. 2017;17:18.
Yang X, Yang Y, Gan R, Zhao L, Li W, Zhou H, et al. Down-regulation of mir-221 and mir-222 restrain prostate cancer cell proliferation and migration that is partly mediated by activation of SIRT1. PLoS ONE. 2014;9:e98833.
Song J, Ouyang Y, Che J, Li X, Zhao Y, Yang K, et al. Potential value of miR-221/222 as diagnostic, prognostic, and therapeutic biomarkers for diseases. Front Immunol. 2017;8:56.
Schulz WA, Burchardt M, Cronauer MV. Molecular biology of prostate cancer. Mol Hum Reprod. 2003;9:437–48.
Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun XW, et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science. 2005;310:644–8.
Hermans KG, van Marion R, van Dekken H, Jenster G, van Weerden WM, Trapman J. TMPRSS2:ERG fusion by translocation or interstitial deletion is highly relevant in androgen-dependent prostate cancer, but is bypassed in late-stage androgen receptor-negative prostate cancer. Cancer Res. 2006;66:10658–63.
Hermans KG, Boormans JL, Gasi D, van Leenders GJ, Jenster G, Verhagen PC, et al. Overexpression of prostate-specific TMPRSS2(exon 0)-ERG fusion transcripts corresponds with favorable prognosis of prostate cancer. Clin Cancer Res. 2009;15:6398–403.
Soller MJ, Isaksson M, Elfving P, Soller W, Lundgren R, Panagopoulos I. Confirmation of the high frequency of the TMPRSS2/ERG fusion gene in prostate cancer. Genes Chromosomes Cancer. 2006;45:717–9.
Attard G, Parker C, Eeles RA, Schroder F, Tomlins SA, Tannock I, et al. Prostate cancer. Lancet. 2016;387:70–82.
Kim TS, Heinlein C, Hackman RC, Nelson PS. Phenotypic analysis of mice lacking the Tmprss2-encoded protease. Mol Cell Biol. 2006;26:965–75.
Adamo P, Ladomery MR. The oncogene ERG: a key factor in prostate cancer. Oncogene. 2016;35:403–14.
Liu CY, Yu T, Huang Y, Cui L, Hong W. ETS (E26 transformation-specific) up-regulation of the transcriptional co-activator TAZ promotes cell migration and metastasis in prostate cancer. J Biol Chem. 2017;292:9420–30.
Tomlins SA, Laxman B, Varambally S, Cao X, Yu J, Helgeson BE, et al. Role of the TMPRSS2-ERG gene fusion in prostate cancer. Neoplasia. 2008;10:177–88.
Deramaudt TB, Remy P, Stiegler P. Identification of interaction partners for two closely-related members of the ETS protein family, FLI and ERG. Gene. 2001;274:169–77.
Vanaja DK, Cheville JC, Iturria SJ, Young CY. Transcriptional silencing of zinc finger protein 185 identified by expression profiling is associated with prostate cancer progression. Cancer Res. 2003;63:3877–82.
Carver BS, Tran J, Chen Z, Carracedo-Perez A, Alimonti A, Nardella C, et al. ETS rearrangements and prostate cancer initiation. Nature. 2009;457:E1. discussion E2-3
Hagglof C, Hammarsten P, Stromvall K, Egevad L, Josefsson A, Stattin P, et al. TMPRSS2-ERG expression predicts prostate cancer survival and associates with stromal biomarkers. PloS ONE. 2014;9:e86824.
Gasi Tandefelt D, Boormans J, Hermans K, Trapman J. ETS fusion genes in prostate cancer. Endocr Relat Cancer. 2014;21:R143–52.
Kokontis JM, Liao S. Molecular action of androgen in the normal and neoplastic prostate. Vitam Horm. 1999;55:219–307.
Tan MH, Li J, Xu HE, Melcher K, Yong EL. Androgen receptor: structure, role in prostate cancer and drug discovery. Acta Pharmacol Sin. 2015;36:3–23.
Velcheti V, Karnik S, Bardot SF, Prakash O. Pathogenesis of prostate cancer: lessons from basic research. Ochsner J. 2008;8:213–8.
Nelson WG, De Marzo AM, Isaacs WB. Prostate cancer. N Engl J Med. 2003;349:366–81.
Sandhu SK, Omlin A, Hylands L, Miranda S, Barber LJ, Riisnaes R, et al. Poly (ADP-ribose) polymerase (PARP) inhibitors for the treatment of advanced germline BRCA2 mutant prostate cancer. Ann Oncol. 2013;24:1416–8.
Chang KH, Li R, Kuri B, Lotan Y, Roehrborn CG, Liu J, et al. A gain-of-function mutation in DHT synthesis in castration-resistant prostate cancer. Cell. 2013;154:1074–84.
Mitsiades N, Sung CC, Schultz N, Danila DC, He B, Eedunuri VK, et al. Distinct patterns of dysregulated expression of enzymes involved in androgen synthesis and metabolism in metastatic prostate cancer tumors. Cancer Res. 2012;72:6142–52.
Stanbrough M, Bubley GJ, Ross K, Golub TR, Rubin MA, Penning TM, et al. Increased expression of genes converting adrenal androgens to testosterone in androgen-independent prostate cancer. Cancer Res. 2006;66:2815–25.
Bonci D, Coppola V, Patrizii M, Addario A, Cannistraci A, Francescangeli F, et al. A microRNA code for prostate cancer metastasis. Oncogene. 2016;35:1180–92.
Carter BS, Beaty TH, Steinberg GD, Childs B, Walsh PC. Mendelian inheritance of familial prostate cancer. Proc Natl Acad Sci USA. 1992;89:3367–71.
Edwards SM, Eeles RA. Unravelling the genetics of prostate cancer. Am J Med Genet C Semin Med Genet. 2004;129c:65–73.
Nwosu V, Carpten J, Trent JM, Sheridan R. Heterogeneity of genetic alterations in prostate cancer: evidence of the complex nature of the disease. Hum Mol Genet. 2001;10:2313–8.
Bratt O. Hereditary prostate cancer: clinical aspects. J Urol. 2002;168:906–13.
Simard J, Dumont M, Labuda D, Sinnett D, Meloche C, El-Alfy M, et al. Prostate cancer susceptibility genes: lessons learned and challenges posed. Endocr Relat Cancer. 2003;10:225–59.
Schulz WA, Burchardt M, Cronauer MV. Molecular biology of prostate cancer. MHR: Basic Sci Reprod Med. 2003;9:437–48.
Hope Q, Bullock S, Evans C, Meitz J, Hamel N, Edwards SM, et al. Macrophage scavenger receptor 1 999C>T (R293X) mutation and risk of prostate cancer. Cancer Epidemiol Biomark Prev. 2005;14:397–402.
Yu X, Guo C, Fisher PB, Subjeck JR, Wang XY. Scavenger Receptors: Emerging Roles in Cancer Biology and Immunology. Adv Cancer Res. 2015;128:309–64.
Li XL, Xu JH. MTHFR polymorphism and the risk of prostate cancer: a meta-analysis of case-control studies. Prostate Cancer Prostatic Dis. 2012;15:244–9.
Castro E, Eeles R. The role of BRCA1 and BRCA2 in prostate cancer. Asian J Androl. 2012;14:409–14.
Martignano F, Gurioli G, Salvi S, Calistri D, Costantini M, Gunelli R, et al. GSTP1 Methylation and protein expression in prostate cancer: diagnostic implications. Dis Markers. 2016;2016:4358292.
Ronen S, Abbott DW, Kravtsov O, Abdelkader A, Xu Y, Banerjee A et al. PTEN loss and p27 loss differ among morphologic patterns of prostate cancer, including cribriform. Hum Pathol. 2017;65:85-91.
Nickerson ML, Im KM, Misner KJ, Tan W, Lou H, Gold B, et al. Somatic alterations contributing to metastasis of a castration-resistant prostate cancer. Hum Mutat. 2013;34:1231–41.
Nickerson ML, Das S, Im KM, Turan S, Berndt SI, Li H, et al. TET2 binds the androgen receptor and loss is associated with prostate cancer. Oncogene. 2017;36:2172–83.
Classon M, Harlow E. The retinoblastoma tumour suppressor in development and cancer. Nat Rev Cancer. 2002;2:910–7.
Mohler JL, Morris TL, Ford OH 3rd, Alvey RF, Sakamoto C, Gregory CW. Identification of differentially expressed genes associated with androgen-independent growth of prostate cancer. Prostate. 2002;51:247–55.
Davies MA, Kim SJ, Parikh NU, Dong Z, Bucana CD, Gallick GE. Adenoviral-mediated expression of MMAC/PTEN inhibits proliferation and metastasis of human prostate cancer cells. Clin Cancer Res. 2002;8:1904–14.
Chappell WH, Lehmann BD, Terrian DM, Abrams SL, Steelman LS, McCubrey JA. p53 expression controls prostate cancer sensitivity to chemotherapy and the MDM2 inhibitor Nutlin-3. Cell Cycle. 2012;11:4579–88.
Fournier PG, Juarez P, Jiang G, Clines GA, Niewolna M, Kim HS, et al. The TGF-beta signaling regulator PMEPA1 suppresses prostate cancer metastases to bone. Cancer Cell. 2015;27:809–21.
Lee C, Zhang Q, Zi X, Dash A, Soares MB, Rahmatpanah F, et al. TGF-beta mediated DNA methylation in prostate cancer. Transl Androl Urol. 2012;1:78–88.
Liu X, Gomez-Pinillos A, Loder C, Carrillo-de Santa Pau E, Qiao R, Unger PD, et al. KLF6 loss of function in human prostate cancer progression is implicated in resistance to androgen deprivation. Am J Pathol. 2012;181:1007–16.
Wang CF, Zhou XY, Zhang TM, Sun MH, Shi DR. Detection of germline mutations of hMLH1 and hMSH2 based on cDNA sequencing in China. World J Gastroenterol. 2005;11:6620–3.
Chang L, Chang M, Chang HM, Chang F. Expending role of microsatellite instability in diagnosis and treatment of colorectal cancers. J Gastrointest Cancer. 2017;48(4):305–313.
Karayi MK, Markham AF. Molecular biology of prostate cancer. Prostate Cancer Prostatic Dis. 2004;7:6–20.
Grindedal EM, Moller P, Eeles R, Stormorken AT, Bowitz-Lothe IM, Landro SM, et al. Germ-line mutations in mismatch repair genes associated with prostate cancer. Cancer Epidemiol Biomark Prev. 2009;18:2460–7.
Reyes GX, Schmidt TT, Kolodner RD, Hombauer H. New insights into the mechanism of DNA mismatch repair. Chromosoma. 2015;124:443–62.
Chen Y, Wang J, Fraig MM, Metcalf J, Turner WR, Bissada NK, et al. Defects of DNA mismatch repair in human prostate cancer. Cancer Res. 2001;61:4112–21.
Pritchard CC, Morrissey C, Kumar A, Zhang X, Smith C, Coleman I, et al. Complex MSH2 and MSH6 mutations in hypermutated microsatellite unstable advanced prostate cancer. Nat Commun. 2014;5:4988.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7.
Itkonen HM, Kantelinen J, Vaara M, Parkkinen S, Schlott B, Grosse F, et al. Human DNA polymerase alpha interacts with mismatch repair proteins MSH2 and MSH6. FEBS Lett. 2016;590:4233–41.
Bhardwaj A, Srivastava SK, Khan MA, Prajapati VK, Singh S, Carter JE, et al. Racial disparities in prostate cancer: a molecular perspective. Front Biosci (Landmark Ed). 2017;22:772–82.
Bensen JT, Xu Z, Smith GJ, Mohler JL, Fontham ET, Taylor JA. Genetic polymorphism and prostate cancer aggressiveness: a case-only study of 1,536 GWAS and candidate SNPs in African-Americans and European-Americans. Prostate. 2013;73:11–22.
Zhang L, Wang J, Wang Y, Zhang Y, Castro P, Shao L, et al. MNX1 Is Oncogenically Upregulated in African-American Prostate Cancer. Cancer Res. 2016;76:6290–8.
Wang Y, Wang J, Zhang L, Karatas OF, Shao L, Zhang Y, et al. RGS12 is a novel tumor-suppressor gene in African American prostate cancer that represses AKT and MNX1 expression. Cancer Res. 2017;77:4247–57.
Zeigler-Johnson C, Friebel T, Walker AH, Wang Y, Spangler E, Panossian S, et al. CYP3A4, CYP3A5, and CYP3A43 genotypes and haplotypes in the etiology and severity of prostate cancer. Cancer Res. 2004;64:8461–7.
Kittles RA, Baffoe-Bonnie AB, Moses TY, Robbins CM, Ahaghotu C, Huusko P, et al. A common nonsense mutation in EphB2 is associated with prostate cancer risk in African American men with a positive family history. J Med Genet. 2006;43:507–11.
Peng L, Wang H, Dong Y, Ma J, Wen J, Wu J, et al. Increased expression of EphA1 protein in prostate cancers correlates with high Gleason score. Int J Clin Exp Pathol. 2013;6:1854–60.
Kwabi-Addo B, Wang S, Chung W, Jelinek J, Patierno SR, Wang BD, et al. Identification of differentially methylated genes in normal prostate tissues from African American and Caucasian men. Clin Cancer Res. 2010;16:3539–47.
Wang BD, Yang Q, Ceniccola K, Bianco F, Andrawis R, Jarrett T, et al. Androgen receptor-target genes in African American prostate cancer disparities. Prostate Cancer. 2013;2013:763569.
Nwaneri AC, McBeth L, Hinds TD Jr. Prostate cancer in African American men: the effect of androgens and microRNAs on epidermal growth factor signaling. Horm Cancer. 2016;7:296–304.
Higgins PB, Fernandez JR, Goran MI, Gower BA. Early ethnic difference in insulin-like growth factor-1 is associated with African genetic admixture. Pediatr Res. 2005;58:850–4.
De Marzo AM, Platz EA, Sutcliffe S, Xu J, Gronberg H, Drake CG, et al. Inflammation in prostate carcinogenesis. Nat Rev Cancer. 2007;7:256–69.
Liu TZ, Guo ZQ, Wang T, Cao Y, Huang D, Wang XH. Meta-analysis of the role of IL-6 rs1800795 polymorphism in the susceptibility to prostate cancer: evidence based on 17 studies. Medicine. 2017;96:e6126.
Gruenewald TL, Cohen S, Matthews KA, Tracy R, Seeman TE. Association of socioeconomic status with inflammation markers in black and white men and women in the coronary artery risk development in young adults (CARDIA) study. Soc Sci Med. 2009;69:451–9.
Fan L, Peng G, Sahgal N, Fazli L, Gleave M, Zhang Y, et al. Regulation of c-Myc expression by the histone demethylase JMJD1A is essential for prostate cancer cell growth and survival. Oncogene. 2016;35:2441–52.
Ma B, Wells A. The mitogen-activated protein (MAP) kinases p38 and extracellular signal-regulated kinase (ERK) are involved in hepatocyte-mediated phenotypic switching in prostate cancer cells. J Biol Chem. 2014;289:11153–61.
Catz SD, Johnson JL. BCL-2 in prostate cancer: a minireview. Apoptosis. 2003;8:29–37.
Chesire DR, Ewing CM, Gage WR, Isaacs WB. In vitro evidence for complex modes of nuclear beta-catenin signaling during prostate growth and tumorigenesis. Oncogene. 2002;21:2679–94.
Li LC, Carroll PR, Dahiya R. Epigenetic changes in prostate cancer: implication for diagnosis and treatment. J Natl Cancer Inst. 2005;97:103–15.
Wang B, Huang J, Zhou J, Hui K, Xu S, Fan J, et al. DAB2IP regulates EMT and metastasis of prostate cancer through targeting PROX1 transcription and destabilizing HIF1alpha protein. Cell Signal. 2016;28:1623–30.
Duggan D, Zheng SL, Knowlton M, Benitez D, Dimitrov L, Wiklund F, et al. Two genome-wide association studies of aggressive prostate cancer implicate putative prostate tumor suppressor gene DAB2IP. J Natl Cancer Inst. 2007;99:1836–44.
Siampanopoulou M, Galaktidou G, Dimasis N, Gotzamani-Psarrakou A. Profiling serum HER-2/NEU in prostate cancer. Hippokratia. 2013;17:108–12.
Hamada S, Horiguchi A, Kuroda K, Ito K, Asano T, Miyai K, et al. Erratum: increased fatty acid synthase expression in prostate biopsy cores predicts higher Gleason score in radical prostatectomy specimen. BMC Clin Pathol. 2015;15:7.
Huang M, Koizumi A, Narita S, Inoue T, Tsuchiya N, Nakanishi H, et al. Diet-induced alteration of fatty acid synthase in prostate cancer progression. Oncogenesis. 2016;5:e195.
Gudmundsson J, Sulem P, Manolescu A, Amundadottir LT, Gudbjartsson D, Helgason A, et al. Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat Genet. 2007;39:631–7.
Yeager M, Orr N, Hayes RB, Jacobs KB, Kraft P, Wacholder S, et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat Genet. 2007;39:645–9.
Gudmundsson J, Sulem P, Steinthorsdottir V, Bergthorsson JT, Thorleifsson G, Manolescu A, et al. Two variants on chromosome 17 confer prostate cancer risk, and the one in TCF2 protects against type 2 diabetes. Nat Genet. 2007;39:977–83.
Gudmundsson J, Sulem P, Rafnar T, Bergthorsson JT, Manolescu A, Gudbjartsson D, et al. Common sequence variants on 2p15 and Xp11.22 confer susceptibility to prostate cancer. Nat Genet. 2008;40:281–3.
Eeles RA, Kote-Jarai Z, Giles GG, Olama AA, Guy M, Jugurnauth SK, et al. Multiple newly identified loci associated with prostate cancer susceptibility. Nat Genet. 2008;40:316–21.
Thomas G, Jacobs KB, Yeager M, Kraft P, Wacholder S, Orr N, et al. Multiple loci identified in a genome-wide association study of prostate cancer. Nat Genet. 2008;40:310–5.
Gudmundsson J, Sulem P, Gudbjartsson DF, Blondal T, Gylfason A, Agnarsson BA, et al. Genome-wide association and replication studies identify four variants associated with prostate cancer susceptibility. Nat Genet. 2009;41:1122–6.
Eeles RA, Kote-Jarai Z, Al Olama AA, Giles GG, Guy M, Severi G, et al. Identification of seven new prostate cancer susceptibility loci through a genome-wide association study. Nat Genet. 2009;41:1116–21.
Yeager M, Chatterjee N, Ciampa J, Jacobs KB, Gonzalez-Bosquet J, Hayes RB, et al. Identification of a new prostate cancer susceptibility locus on chromosome 8q24. Nat Genet. 2009;41:1055–7.
Al Olama AA, Kote-Jarai Z, Giles GG, Guy M, Morrison J, Severi G, et al. Multiple loci on 8q24 associated with prostate cancer susceptibility. Nat Genet. 2009;41:1058–60.
Easton DF, Pooley KA, Dunning AM, Pharoah PD, Thompson D, Ballinger DG, et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature. 2007;447:1087–93.
Amundadottir LT, Sulem P, Gudmundsson J, Helgason A, Baker A, Agnarsson BA, et al. A common variant associated with prostate cancer in European and African populations. Nat Genet. 2006;38:652–8.
Haiman CA, Patterson N, Freedman ML, Myers SR, Pike MC, Waliszewska A, et al. Multiple regions within 8q24 independently affect risk for prostate cancer. Nat Genet. 2007;39:638–44.
Takata R, Akamatsu S, Kubo M, Takahashi A, Hosono N, Kawaguchi T, et al. Genome-wide association study identifies five new susceptibility loci for prostate cancer in the Japanese population. Nat Genet. 2010;42:751–4.
Boyd LK, Mao X, Lu YJ. The complexity of prostate cancer: genomic alterations and heterogeneity. Nat Rev Urol. 2012;9:652–64.
Zheng SL, Sun J, Wiklund F, Smith S, Stattin P, Li G, et al. Cumulative association of five genetic variants with prostate cancer. N Engl J Med. 2008;358:910–9.
Ewing CM, Ray AM, Lange EM, Zuhlke KA, Robbins CM, Tembe WD, et al. Germline mutations in HOXB13 and prostate-cancer risk. N Engl J Med. 2012;366:141–9.
Park JJ, Jin YB, Lee YJ, Lee JS, Lee YS, Ko YG, et al. KAI1 suppresses HIF-1alpha and VEGF expression by blocking CDCP1-enhanced Src activation in prostate cancer. BMC Cancer. 2012;12:81.
Mashimo T, Watabe M, Cuthbert AP, Newbold RF, Rinker-Schaeffer CW, Helfer E, et al. Human chromosome 16 suppresses metastasis but not tumorigenesis in rat prostatic tumor cells. Cancer Res. 1998;58:4572–6.
Pritchard CC, Mateo J, Walsh MF, De Sarkar N, Abida W, Beltran H, et al. Inherited DNA-repair gene mutations in men with metastatic prostate cancer. N Engl J Med. 2016;375:443–53.
Noguchi M, Koga N, Igawa T, Itoh K. Clinical development of immunotherapy for prostate cancer. Int J Urol. 2017;24(9):675–680.
Grozescu T, Popa F. Immunotherapy and gene therapy in prostate cancer treatment. J Med Life. 2017;10:54–55.
Singh P, Pal SK, Alex A, Agarwal N. Development of PROSTVAC immunotherapy in prostate cancer. Future Oncol. 2015;11:2137–48.
Mandl SJ, Rountree RB, Dela Cruz TB, Foy SP, Cote JJ, Gordon EJ, et al. Elucidating immunologic mechanisms of PROSTVAC cancer immunotherapy. J Immunother Cancer. 2014;2:34.
Slovin SF, Higano CS, Hamid O, Tejwani S, Harzstark A, Alumkal JJ, et al. Ipilimumab alone or in combination with radiotherapy in metastatic castration-resistant prostate cancer: results from an open-label, multicenter phase I/II study. Ann Oncol. 2013;24:1813–21.
Kershaw MH, Westwood JA, Darcy PK. Gene-engineered T cells for cancer therapy. Nat Rev Cancer. 2013;13:525–41.
Wong YNS, Joshi K, Pule M, Peggs KS, Swanton C, Quezada SA, et al. Evolving adoptive cellular therapies in urological malignancies. Lancet Oncol. 2017;18:e341–e53.
Hillerdal V, Essand M. Chimeric antigen receptor-engineered T cells for the treatment of metastatic prostate cancer. BioDrugs. 2015;29:75–89.
Silvestri I, Cattarino S, Giantulli S, Nazzari C, Collalti G, Sciarra A. A perspective of immunotherapy for prostate cancer. Cancers. 2016;8(7),64; doi:10.3390/cancers8070064.
Acknowledgements
The authors are thankful to Drs. Kelly Warren, Inefta Reid, Todd Miller, and Peter Brink for departmental support, as well as Mrs. Wendy Isser and Ms. Grace Garey for literature retrieval.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Gandhi, J., Afridi, A., Vatsia, S. et al. The molecular biology of prostate cancer: current understanding and clinical implications. Prostate Cancer Prostatic Dis 21, 22–36 (2018). https://doi.org/10.1038/s41391-017-0023-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41391-017-0023-8
This article is cited by
-
SNAP25 is a potential prognostic biomarker for prostate cancer
Cancer Cell International (2022)
-
SLC4A4 promotes prostate cancer progression in vivo and in vitro via AKT-mediated signalling pathway
Cancer Cell International (2022)
-
Prostate cancer histopathology using label-free multispectral deep-UV microscopy quantifies phenotypes of tumor aggressiveness and enables multiple diagnostic virtual stains
Scientific Reports (2022)
-
The role of the p90 ribosomal S6 kinase family in prostate cancer progression and therapy resistance
Oncogene (2021)
-
Targeting the Lnc-OPHN1-5/androgen receptor/hnRNPA1 complex increases Enzalutamide sensitivity to better suppress prostate cancer progression
Cell Death & Disease (2021)