Abstract
Primary immunodeficiency diseases (PIDs) caused by a single-gene defect generally are referred to as monogenic autoimmune disorders. For example, mutations in the transcription factor autoimmune regulator (AIRE) result in a condition called autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy; while mutations in forkhead box P3 lead to regulatory T cell (Treg)-deficiency-induced multiorgan inflammation, which in humans is called “immune dysregulation, polyendocrinopathy, enteropathy with X-linked inheritance” (or IPEX syndrome). Previous studies concluded that monogenic diseases are insensitive to commensal microbial regulation because they develop even in germ-free (GF) animals, a conclusion that has limited the number of studies determining the role of microbiota in monogenic PIDs. However, emerging evidence shows that although the onset of the disease is independent of the microbiota, several monogenic PIDs vary in severity in association with the microbiome. In this review, we focus on monogenic PIDs associated with Treg deficiency/dysfunction, summarizing the gut microbial dysbiosis that has been shown to be linked to these diseases. From limited studies, we have gleaned several mechanistic insights that may prove to be of therapeutic importance in the early stages of life.
Impact
-
This review paper serves to refute the concept that monogenic PIDs are not linked to the microbiome.
-
The onset of monogenic PIDs is independent of microbiota; single-gene mutations such as AIRE or Foxp3 that affect central or peripheral immune tolerance produce monogenic diseases even in a GF environment.
-
However, the severity and outcome of PIDs are markedly impacted by the microbial composition.
-
We suggest that future research for these conditions may focus on targeting the microbiome.
Similar content being viewed by others
Introduction
Autoimmune diseases (ADs) are a heterogeneous group of disorders in which the immune system responds to self-antigens, leading to damage or dysfunction of tissues.1 Relatively prevalent ADs such as rheumatoid arthritis (RA), systemic lupus erythematosus (SLE), type I diabetes (T1D), and inflammatory bowel disease (IBD) are polygenic autoimmune disorders, which means that the pathogenesis of the diseases may relate simultaneously to the changes of several genes. As an example, in a genome-wide association study (GWAS) including European and Asian subjects—29,880 patients with RA and 73,758 control subjects—377 candidate genes were identified in 100 non-major histocompatibility complex (MHC) RA risk loci. Among them, 98 genes were highly related to RA risk.2 GWAS has identified >50 robust loci associated with SLE susceptibility,3 and (similarly) >200 robust loci have been associated with IBD.4
The primary immunodeficiency diseases (PIDs) are genetic in origin and result in autoimmunity that affects different components of the innate and adaptive immune systems. Although PIDs can affect both adults and children, they are more common during childhood, inasmuch as newborns with PIDs generally do not survive into adolescence. The pathophysiological processes of both PIDs and ADs are associated with abnormal T cell development, immune tolerance, signaling, and inflammation, and they share some common mechanisms.5 However, there are many monogenic PIDs in which a single highly penetrant genetic alteration results in an autoimmune disorder. The role of a single mutated gene in the development of autoimmunity helps to elucidate the underlying mechanisms associated with autoimmunity. Understanding the mechanisms will benefit the development of effective treatments for these diseases. Currently, only bone marrow transplantation is commonly used to treat these diseases, but this procedure is limited by the procurement of a matched donor, is expensive, is associated with graft-versus-host disease, and is often lethal.6
The microbiome comprises the live microorganisms and genomes (bacteria, viruses, phage, fungi, and protozoa) that exist in a symbiosis within mammalian tissues. The microbiome has the ability to influence different physiological functions, such as immune defense, energy metabolism, and social behavior.7 The intestinal microbiota drives host immune homeostasis by regulating the differentiation and expansion of immune cells such as regulatory T cells (Tregs).8,9,10,11 Intestinal microbial dysbiosis (or imbalance) can develop as a consequence of an abnormal diet, severe or chronic infection, body habitus (lean or obese), or altered gut immunity.12,13,14,15 Many studies have provided evidence demonstrating the association between gut dysbiosis and polygenic ADs such as RA,16,17 IBD,4 type I diabetes,18,19 SLE,20 and multiple sclerosis (MS).21,22,23 As a result, the scientific community is developing optimized probiotics and fecal microbial transplantation (FMT) to modify gut microbiota and disease activity for many of these conditions. In addition, microbiota-associated biological metabolites, called postbiotics, may facilitate the treatment of ADs.24,25
However, there is limited information about monogenic PIDs caused by single-gene mutations and how they interact with the microbiome. Thus, this review will highlight our knowledge of the association of microbiome to monogenic PIDs, focusing on diseases related to Treg deficiency/dysregulation.
Monogenic PIDs related to Treg deficiency or dysfunction
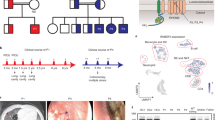
Monogenic PIDs are rare, but >350 unique PIDs have been identified so far. The children typically present with autoimmune phenomena and increased susceptibility to infection, eventually developing multiorgan inflammation.26 The main monogenic PIDs related to Treg deficiency/dysfunction are due to a single-gene defect, and the mechanisms that are implicated are listed in Table 1. The representative clinical phenotypes of some diseases listed in Table 1 are cutaneous or onchopathic (fingernail/toenail) manifestations (Fig. 1). Treg cells maintain immune homeostasis and play a pivotal role in immune tolerance. Forkhead box P3 (Foxp3) is a major transcription factor associated with Treg cell development and function.27 Mutations or deletions of the Foxp3 gene result in immune dysregulation, polyendocrinopathy, and enteropathy with X-linked inheritance (IPEX) syndrome in humans. IPEX syndrome is characteristically associated with eczema (Fig. 1a, b), severe enteropathy, type I diabetes, thyroiditis, hemolytic anemia, and thrombocytopenia.28,29,30 Several other genetic defects that affect the function of Tregs can give rise to IPEX-like syndromes. IPEX-like syndromes include loss of function mutations in the α-chain of the interleukin-2 (IL-2) receptor (CD25), itchy E3 ubiquitin protein ligase (ITCH), signal transducer and activator of transcription 5B (STAT5B)/BTB domain, and CNC homolog 2 (BACH2). Recently, there are mutations reported in other genes, including LPS responsive beige-like anchor protein (LRBA), cytotoxic T-lymphocyte associated protein 4 (CTLA4), dedicator of cytokinesis 8 (DOCK8), and mucosa-associated lymphoid tissue lymphoma translocation 1 (MALT1).31
Other monogenic PIDs are not classified as IPEX-like syndromes, but they are immunologically associated with Treg deficiency and/or dysfunction and linked to gastrointestinal inflammation. These include autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy (APECED) syndrome (Fig. 1c–f), Omenn syndrome (OS) (Fig. 1g), DiGeorge syndrome, chronic granulomatous disease (CGD), Wiskott–Aldrich syndrome (WAS) (Fig. 1h, i), and human phosphatidylinositol 3-kinase-gamma (PI3Kγ) deficiency (Table 1 and Fig. 1).31,32,33,34,35,36,37
Traditional concept of the innate-adaptive connection and microbial sensitivity at the onset of monogenic autoimmunity
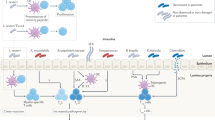
Mechanistically, common ADs require the innate-adaptive connection for their onset, during which process microbes contribute to the initiation of autoimmunity. Microbes and microbial peptides activate antigen-presenting cells (APCs) through the interaction with innate receptor signaling on APCs. Subsequently, APCs induce effector and memory T cell and B cells by antigen presentation and co-stimulation, and ultimately tissue target cells are destroyed by self-reactive T cells, antibodies, and/or cytokines.38,39 These ADs have been designated as group I diseases. They follow the rule of innate-adaptive immunity activation and require a microbial environment (Fig. 2).40
Group I diseases follow the rules of innate-adaptive immunity activation. They can be affected by microbial environment, and/or may result from genetic alterations to PRR sensing and signaling, co-stimulation, or cytokine production. Innate PRR expressed on APC recognize pathogenic antigens and presents to naive T cells, activating naive T cells by several pairs of ligand–receptor interactions to differentiate T cell subsets. This results in more inflammatory Teffs, and self-reactive T cells than Tregs, and damage to self-tissues ensues. Group II diseases are caused by genetic alterations in lymphocyte development or regulation. An example shown is a Foxp3 gene mutation in Foxp3+ Treg cells in the thymus, which results in defects in peripheral tolerance due to Treg deficiency/dysfunction that cannot inhibit inflammatory Teffs, yielding high levels of pro-inflammatory cytokines. Group II diseases are independent of innate-adaptive immune activation, and their onset is less affected by microbiota. Abbreviations: Ags antigens, PRR pattern recognition receptor, PAMPs pathogen-associated molecular patterns, APC antigen-presenting cell, nTreg natural regulatory T cell, Teff effector T cell, Tem effector/memory T cell.
Because alterations in microbial composition often correlate with the loss of immune tolerance, the human microbiome has been proposed to be a major player in autoimmunity. IBDs such as Crohn’s disease (CD) and ulcerative colitis (UC) represent an example of how the alteration of gut microbiome can induce disease. Many studies have shown that both CD and UC are associated with a reduced complexity of the commensal microbiota and are consistent shifts to a dysbiotic state, characterized by the outgrowth of “pathobionts,” such as the phyla proteobacteria, in particular the Enterobacteriaceae and Fusobacteriaceae families.41,42 Adherent-invasive E. coli, Yersinia, and Clostridium difficile are also much more common in patients with CD than in healthy individuals. These bacteria are also key contributors to IBD in animal models.43,44,45,46
The separation of monogenic from polygenic autoimmune disorders is based on evidence in germ-free (GF) mice that the innate-adaptive connection is not critical for the onset of monogenic disorders. For example, re-deriving AIRE−/− mice, a mouse model of human APECED, into a GF condition does not alter the disease phenotype. The implication of these studies was that the loss of central tolerance in the thymus alone can lead to autoimmunity that overrides peripheral tolerance mechanisms.47 In addition, Foxp3 mutant scurfy (SF) mice, as mentioned a mouse model of human IPEX syndrome, exhibit an autoimmune phenotype in the GF condition, due to loss of control of inflammatory effector T cells by Tregs.48 These observations were interpreted to prove that monogenic diseases related to Treg deficiency, for example, APECED and IPEX are independent of commensal microbial regulation. Authors suggested that autoimmune T cells are activated in the absence of the innate-adaptive connection,38 leading to the designation of group II diseases, which resulted from genetic alterations in lymphocyte development and not from microbial factors (Fig. 2).40
Microbial dysbiosis and Treg-associated monogenic autoimmunity
Chinen et al.48 proposed that monogenic autoimmunity is not associated with the gut microbiome, because innate-adaptive connection is not critical for disease onset. However, in a Treg-depleted model (Foxp3-DTR), inflammation in the small intestine of SPF mice was more severe than in GF mice, as shown by significantly increased gut lymphocyte infiltration, decreased body weight, and increased percentage of interferon-γ-producing T helper cells. That study suggested that Treg deficiency-induced inflammation is indeed modified by gut microbiota.48
Other experiments provided evidence that the severity of single-gene mutation-associated ADs is modified by the gut microbiota. We observed that development of autoimmunity was accompanied by gut microbial dysbiosis during the first 22 days of life in SF mice,49 at which time these mice had reduced bacterial diversity and composition. Altered gut microbiota has also been found in immunodeficient mice, in models lacking B cells (Ighm−/−), T cells (Cd3e−/−), or both B and T cells (Rag1−/−). Furthermore, the administration of Foxp3+ Treg cells to T cell-deficient mice restored bacterial diversity.50 A study of APECED patients also concluded that the composition of their intestinal flora was abnormal;51 and consequently the patients developed early and sustained immunological responses to gut microbial antigens. In addition, abnormal immune recognition of gut commensals is linked to gut-associated Tregs, indicating that AIRE is an important regulator of intestinal homeostasis.52 A detailed study comparing Rag1+/+ and Rag1−/− mice of the same genetic background (with the appropriate elimination of cage effect) demonstrated that Rag1 status was a source of variation in gut microbiota community structure.53
A related study examining a mouse model of Omenn syndrome (OS) with hypomorphic mutations in Rag genes reported that impaired Treg function played a role in development of intestinal inflammation, while mucosal B cell deficiency caused enhanced bacterial translocation and altered microbiota composition. Reducing bacterial load by broad-spectrum antibiotics ameliorated intestinal and systemic autoimmunity by diminishing the frequency of mucosal and circulating gut-tropic CCR9+ Th1 and Th17 cells and normalizing very high levels of serum IgE, a hallmark of the disease. These observations indicate that intestinal microbes play a critical role in the distinctive immune aberrations seen in OS.54
Furthermore, studies of children with WAS found aberrant microbial community richness and structure in WAS children. WAS children had reduced microbial community richness and diversity. The relative abundance of Bacteroidetes and Verrucomicrobia in WAS children was significantly reduced, whereas Proteobacteria were at markedly higher levels.55 WASp-knockout mice modeling human WAS harbored a significantly decreased relative abundance of Firmicutes.55 In humans, up to 50% of WAS patients develop IBD, often early in onset and severe in phenotype.56 Fecal microbial dysbiosis caused by WASp deficiency was quite similar to that observed for polygenic IBD,41,55,57 indicating that WASp may play a crucial function in microbial homoeostasis and/or that microbial dysbiosis may contribute to WAS-related IBD.
Interestingly, a recent study investigating PIK3CG-deficient mice tried to mimic a monogenic condition in humans, PI3Kγ deficiency, found a lack of spontaneous inflammation in the lung and gut of PIK3CG-deficient mice when the mice were housed in specific pathogen-free environments. However, “dirty” PIK3CG-deficient mice that were co-housed with “pet-store mice” shared many of the features of disease witnessed in patients with human PI3Kγdeficiency. Specifically, they exhibited elevated IL-12 release from macrophages, defective immunoglobulin production, increased inflammatory T cell infiltration in the intestine, and a reduced frequency of circulating Tregs.37 This natural pathogen-dependent mouse model of immunodeficiency and immunopathology indicates a crucial role of microbes to contribute to the clinical features of human monogenic PI3Kγ deficiency.
In addition to gut microbial dysbiosis in gastrointestinal disorders, children with severe atopic dermatitis were found to harbor altered fungal skin microbiomes. Notably, in monogenic PIDs such as STAT3, WASp, and DOCK8 deficiency, skin mycobiome has been found to differ from that of healthy controls.58 PID skin also displayed altered microbial population structures not observed in controls, including colonization with Clostridium species and Serratia marcescens, as well as elevated fungal diversity and increased representation of opportunistic fungi such as Candida and Aspergillus. Clinical parameters, including markers of disease severity, correlated positively with prevalence of Staphylococcus and Corynebacterium. Further investigation is needed to determine if an altered landscape of the human skin microbiome contributes to disease in patients with these Treg-related monogenic PIDs.
Increasingly, children with IBD are found to have monogenic gene deficiencies (Table 1) that can lead to very early-onset IBD (VEO-IBD), as defined by disease development before the age of 6 years. VEO-IBDs are phenotypically and genetically distinct from older-onset IBD,59 even though their discovery indicated a polygenic component.60 It is important to remember that the gut microbiota colonizes and develops primarily between birth and 3 years of life,61 at a time that coincides with the age of onset of VEO-IBD. Advances in next-generation sequencing methods improve the identification of known molecular defects in VEO-IBD caused by single-gene mutations.62 As mentioned, it is noteworthy that CGD caused by the CYBB mutation results in deficiency of NADPH oxidase 2 that impairs the production of reactive oxygen species (ROS) in the colon. Once patients with CGD are infected with encapsulated bacteria, these patients often develop VEO-IBD, with a distal colitis and Treg deficiency.36,63 Recently, rare variants in Nox1 have been found in patients with VEO-IBD, showing elevated level of ROS in patient-derived colonic organoid cultures, and constitutively generating a high level of ROS in the colonic crypt lumen.64
A recent report compared fecal microbiota composition of patients with VEO-IBD and three types of PIDs: CGD (11 samples), X-linked inhibitor of apoptosis (XIAP) deficiency (7 samples), and partial tetratricopeptide repeat domain 7A (TTC7A) deficiency (7 samples), comparing their 16S sequences with those patients with non-VEO-IBD (18 samples) and healthy subjects (23 samples).65 Abnormal stool microbiota composition and diversity in these PIDs were characterized by disease-specific changes, with a dramatic increase in Proteobacteria from the Enterobacteriaceae family in the TTC7A group; an increased proportion of bacteria from the Bacteroidetes phylum in CGD; and increased Clostridiaceae family in XIAP. Patients with CGD also exhibited an increased abundance of Ruminococcus gnavus, an organism that has also been associated with ileal CD in children.65 The dysbiosis induced by the host genetic defect might play a role in the PID phenotype, such as perpetuating inflammation and intestinal involvement, but it seems unlikely that dysbiosis is the cause of IBD. In support of this concept, treatment of the primary immunodeficiency may “cure the dysbiosis.” SCID-X1 gene therapy for children with severe-combined immune deficiency (SCID-X1) caused by mutation of the IL2RG gene has been shown to reconstitute T cell repertoire and modulate antibiotic resistance gene levels. Interestingly, SCID-X1 gene therapy enabled not only the expression of therapeutic IL2RG gene but also cleared microbial pathogens (viruses) and normalized the gut microbiome.66
Microbial modification as a potentially therapeutic strategy
Microbial dysbiosis associated with Treg-related monogenic autoimmunity may be altered by targeting the enteric microbiota with the aim of modifying the course of these monogenic disorders. The early assembly of a healthy microbiome is heavily influenced by the mode of delivery (vaginal or C-section) and feeding type (breast milk or formula).61 Recently, Aagaard et al.67 reported that the placenta contains a unique microbiota, which can affect the formation of a child’s microbiome. Mothers are key determinants in establishing the microbiota in early stages of life. Maternal influences in early life, including gestation, birth history, breastfeeding, and toddler exposures, are extremely important in microbiota formation, making this period a promising target for bio-therapeutics that may prevent diseases later in life.68 Factors such as diet, antibiotic use, and enteric diseases will also impact the composition of the microbiota, but the community generally will subsequently recover.69
Current strategies for gut microbiota modulation include prebiotics, probiotics, postbiotics, and FMT.68 Prebiotics are foods (containing high dietary fiber and other substrates such as polysaccharides and casein-derived amino acids) used by probiotics or commensal bacteria for growth. Recent studies showed that a lack of dietary fiber induces a substantial loss of microbial community diversity and influences the ability of gut bacteria to be transferred from parents to their offspring.70
Probiotics have been defined as live, natural microorganisms that are given orally to confer health benefits to the host.71 Bifidobacterium and Lactobacillus strains are the most widely available commercialized probiotics, although there are numerous other species with proposed or demonstrated health benefits. The beneficial effects of probiotics in treating diseases are strain-specific, which means that different probiotic strains are not equally potent. Probiotics interact with the host within a very complex microbiota ecosystem. Therefore, it has been challenging to define the crosstalk between individual bacterial strains and the host. However, studies suggest that probiotics modulate immune responses, ensure homeostasis of the healthy mucosal microbiota, and may ameliorate immune‐mediated diseases. Their mechanisms of action include enhanced production of antimicrobial peptides; maintenance of the gastrointestinal epithelial barrier; and facilitation of adequate interactions between the gut microbes, intestinal epithelial cells, and mucosal immune cells. Finally, probiotics or their associated microbiota produce biologically functional metabolites that modulate the immune system and may act as neurotransmitters to modulate brain functions.72
Probiotics have been studied for ADs in humans and animal models, mostly in polygenic autoimmune disorders including T1D, MS, RA, SLE, and IBD.73 Stirsciuglio et al.74 found that a commercial probiotic (Tribif), composed of three strains of Bifidobacterium (B. longum, B. breve, and B. infantis) improved antigen sampling and processing by dendritic cells, ameliorated the impairment of intestinal innate immunity, and reduced uncontrolled microbial expansion in the intestine of children with CD, but not in those with UC. The study evaluated how probiotics may be used as potential preventive therapy for chronic pediatric inflammatory diseases (CIDs) including celiac disease and IBD through altering the balance of Tregs and inflammatory T effector cells (Teffs) in the diseases.75 However, large high-quality studies on the effect of probiotics in pediatric IBD are limited.75,76
For monogenic PIDs, as exemplified by the IPEX syndrome, we have demonstrated that the gut microbial dysbiosis due to Foxp3+ Treg deficiency in SF mice, a mouse model of IPEX syndrome, could be reprogramed by probiotic Lactobacillus reuteri DSM 17938 (recently renamed as Limosilactobacillus reuteri, L. reuteri), a human-derived probiotic, which has been used to treat infantile colic and acute infectious diarrhea.77,78,79 In several studies, we found that L. reuteri prolongs the survival and reduces hepatic and lung autoimmunity in SF mice. The L. reuteri mechanism of action involved modulating gut microbiota to produce functional metabolites. One key metabolite, adenosine-derived inosine, we found exerted anti-inflammatory effects that were mediated via the adenosine 2A (A2A) receptors. The A2A receptors expressed on T cells once bound with A2A agonists result in inhibition of inflammatory Th1 and Th2 differentiation and reducing Th1- and Th2- associated cytokines (Fig. 3). It is now well established that one of the ways by which Tregs control inflammatory Teffs (Th1/Th2/Th17) is by producing adenosine, generated from ATP/AMP by CD39/CD73 signaling expressed on Treg cells (Fig. 3a).80,81 However, this adenosine-mediated control Treg/Teff balance was broken in the setting of Treg deficiency (Fig. 3b).
a In WT mice, Tregs generate adenosine from ATP/AMP by CD39/CD73 signaling expressed on Treg cells. Adenosine interacts with A2A expressed on inflammatory Teffs to control Teffs and reduce inflammation. b In SF mice, inflammatory Teffs lose their controlling Tregs and also lose the adenosine mechanism, resulting in severe inflammation and autoimmunity. c Gavage feeding of L. reuteri to SF mice modulates gut microbiota, generates adenosine and its metabolite inosine, an A2A agonist, which interacts with A2A to inhibit Teff cell differentiation, and reduce multiorgan inflammation. d The therapeutic effect of L. reuteri and inosine was blocked by genetically knocking out receptor A2A in SF mice, indicating that A2A plays a key role in L. reuteri protection. Abbreviations: ATP adenosine triphosphate, AMP adenosine monophosphate, A2A adenosine receptor 2A, TCR T cell receptor, SF mice scurfy mice, Th1 T helper cell, Treg regulatory T cell, Teff effector T cell.
In our studies, the probiotic L. reuteri increased production of adenosine73,82 and its metabolite inosine,49 which directly interacted with A2A expressed on Teffs (in the absence of Tregs) to control inflammation/autoimmunity (Fig. 3c).49,83 We proved the importance of this adenosine-A2A receptor mechanism by (a) blocking A2A receptors by a pharmacological blocker (Fig. 3c) and (b) genetic knockout of A2A receptors in SF mice (Fig. 3d). Either manipulation prevented any beneficial effect of L. reuteri in ameliorating clinical symptoms.49,84 The exact mechanism of how L. reuteri produces adenosine and inosine, either individually and/or by L. reuteri-modulated commensal bacteria, requires further characterization. Lactobacillus reuteri may additionally have a therapeutic effect in other monogenic autoimmune disorders that involve reduced numbers and/or dysfunction of Tregs, for example, in IPEX-like syndromes as listed in Table 1. Currently, there are no reports on FMT as a treatment for monogenic PID.
In summary, the metabolic byproducts from probiotics with beneficial biological activity in the host, for example, adenosine/inosine, short-chain fatty acids (acetate, butyrate, and propionate), and tryptophan have been termed postbiotics.68 The engineering of genetically modified probiotics that produce this type of postbiotic, or the choice and dose of postbiotic, to benefit host health is a formidable challenge for scientists dedicated to improving the outlook of people with ADs.
Conclusion
The onset of monogenic PIDs related to Treg deficiency/dysfunction is not linked to the composition of the intestinal microbiome. However, current evidence shows that the severity of Treg-associated monogenic autoimmune disorders is clearly associated with gut microbial dysbiosis. The therapeutic effect of probiotics and the probiotic-derived inosine in a mouse model of human IPEX syndrome provides a good example of how modulating gut microbiota can be of therapeutic value in treating monogenic PID. The translatability of microbiome-directed therapies for the diseases listed in Table 1 will undoubtedly be hindered by the fact that these conditions are rare, and clinical trials will initially lack statistical power. To aid in discovery, mouse models with a humanized immune system such as mice with humanized Foxp3 gene-edited mutations85,86 and/or with a humanized microbiome may become very useful tools. Thus, translational research will help to assess the impact of prebiotics, probiotics, synbiotics, postbiotics, and FMT in benefitting children with these disorders.
References
Anaya, J. M., Shoenfeld, Y., Rojas-Villarrage, A. & Cervera R. Autoimmunity. From Bench to Bedside (Rosario University Press, 2013).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Deng, Y. & Tsao, B. P. Advances in lupus genetics and epigenetics. Curr. Opin. Rheumatol. 26, 482–492 (2014).
Momozawa, Y. et al. IBD risk loci are enriched in multigenic regulatory modules encompassing putative causative genes. Nat. Commun. 9, 2427 (2018).
Amaya-Uribe, L., Rojas, M., Azizi, G., Anaya, J. M. & Gershwin, M. E. Primary immunodeficiency and autoimmunity: a comprehensive review. J. Autoimmun. 99, 52–72 (2019).
Castagnoli, R., Delmonte, O. M., Calzoni, E. & Notarangelo, L. D. Hematopoietic stem cell transplantation in primary immunodeficiency diseases: current status and future perspectives. Front. Pediatr. 7, 295 (2019).
Sonnenburg, E. D. & Sonnenburg, J. L. The ancestral and industrialized gut microbiota and implications for human health. Nat. Rev. Microbiol. 17, 383–390 (2019).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Round, J. L. & Mazmanian, S. K. Inducible Foxp3+ regulatory T-cell development by a commensal bacterium of the intestinal microbiota. Proc. Natl Acad. Sci. USA 107, 12204–12209 (2010).
Ruohtula, T. et al. Maturation of gut microbiota and circulating regulatory T cells and development of IgE sensitization in early life. Front. Immunol. 10, 2494 (2019).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
Goodrich, J. K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799 (2014).
Lukens, J. R. et al. Dietary modulation of the microbiome affects autoinflammatory disease. Nature 516, 246–249 (2014).
Lupp, C. et al. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe 2, 119–129 (2007).
Kishikawa, T. et al. Metagenome-wide association study of gut microbiome revealed novel aetiology of rheumatoid arthritis in the Japanese population. Ann. Rheum. Dis. 79, 103–111 (2020).
Maeda, Y. & Takeda, K. Host-microbiota interactions in rheumatoid arthritis. Exp. Mol. Med. 51, 1–6 (2019).
Durazzo, M., Ferro, A. & Gruden, G. Gastrointestinal microbiota and type 1 diabetes mellitus: the state of art. J. Clin. Med. 8, 1843 (2019).
Thomas, R. M. & Jobin, C. Microbiota in pancreatic health and disease: the next frontier in microbiome research. Nat. Rev. Gastroenterol. Hepatol. 17, 53–64 (2020).
Kim, J. W., Kwok, S. K., Choe, J. Y. & Park, S. H. Recent advances in our understanding of the link between the intestinal microbiota and systemic lupus erythematosus. Int. J. Mol. Sci. 20, 4871 (2019).
Brown, J., Quattrochi B., Everett, C., Hong, B. Y. & Cervantes, J. Gut commensals, dysbiosis, and immune response imbalance in the pathogenesis of multiple sclerosis. Mult. Scler. https://doi.org/10.1177/352458520928301 (2020).
He, B. et al. Lactobacillus reuteri reduces the severity of experimental autoimmune encephalomyelitis in mice by modulating gut microbiota. Front Immunol. 10, 385 (2019).
Kadowaki, A. & Quintana, F. J. The gut-CNS axis in multiple sclerosis. Trends Neurosci. 43, 622–634 (2020).
Balakrishnan, B. & Taneja, V. Microbial modulation of the gut microbiome for treating autoimmune diseases. Expert. Rev. Gastroenterol. Hepatol. 12, 985–996 (2018).
Vangoitsenhoven, R. & Cresci, G. A. M. Role of microbiome and antibiotics in autoimmune diseases. Nutr. Clin. Pract. 35, 406–416 (2020).
Schmidt, R. E., Grimbacher, B. & Witte, T. Autoimmunity and primary immunodeficiency: two sides of the same coin? Nat. Rev. Rheumatol. 14, 7–18 (2017).
Ouyang, W. et al. Foxo proteins cooperatively control the differentiation of Foxp3+ regulatory T cells. Nat. Immunol. 11, 618–627 (2010).
Bennett, C. L. et al. The immune dysregulation, polyendocrinopathy, enteropathy, X-linked syndrome (IPEX) is caused by mutations of FOXP3. Nat. Genet. 27, 20–21 (2001).
Bennett, C. L. & Ochs, H. D. IPEX is a unique X-linked syndrome characterized by immune dysfunction, polyendocrinopathy, enteropathy, and a variety of autoimmune phenomena. Curr. Opin. Pediatr. 13, 533–538 (2001).
Halabi-Tawil, M. et al. Cutaneous manifestations of immune dysregulation, polyendocrinopathy, enteropathy, X-linked (IPEX) syndrome. Br. J. Dermatol. 160, 645–651 (2009).
Azizi, G. et al. Monogenic polyautoimmunity in primary immunodeficiency diseases. Autoimmun. Rev. 17, 1028–1039 (2018).
Albert, M. H. & Freeman, A. F. Wiskott-Aldrich syndrome (WAS) and dedicator of cytokinesis 8- (DOCK8) deficiency. Front. Pediatr. 7, 451 (2019).
Candotti, F. Clinical manifestations and pathophysiological mechanisms of the Wiskott-Aldrich syndrome. J. Clin. Immunol. 38, 13–27 (2018).
El-Darouti, M. A. & Al-Ali, F. M. Alopecia, nail dystrophy, vitiligo, and hypoparathyroidism. Challenging Cases Dermatol. 2, 199–206 (2019).
Hsu, C., Lee, J. Y. Y. & Chao, S. C. Omenn syndrome: a case report and review of literature. Dermatol. Sin. 29, 50–54 (2011).
Roos, D. et al. Mutations in the X-linked and autosomal recessive forms of chronic granulomatous disease. Blood 87, 1663–1681 (1996).
Takeda, A. J. et al. Human PI3Kgamma deficiency and its microbiota-dependent mouse model reveal immunodeficiency and tissue immunopathology. Nat. Commun. 10, 4364 (2019).
Chervonsky, A. V. Microbiota and autoimmunity. Cold Spring Harb. Perspect. Biol. 5, a007294 (2013).
Mogensen, T. H. Pathogen recognition and inflammatory signaling in innate immune defenses. Clin. Microbiol. Rev. 22, 240–273 (2009). Table.
Chervonsky, A. V. Influence of microbial environment on autoimmunity. Nat. Immunol. 11, 28–35 (2010).
Frank, D. N. et al. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl Acad. Sci. USA 104, 13780–13785 (2007).
Gevers, D. et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe 15, 382–392 (2014).
Chassaing, B. et al. Crohn disease-associated adherent-invasive E. coli bacteria target mouse and human Peyer’s patches via long polar fimbriae. J. Clin. Invest. 121, 966–975 (2011).
Lamps, L. W. et al. Pathogenic Yersinia DNA is detected in bowel and mesenteric lymph nodes from patients with Crohn’s disease. Am. J. Surg. Pathol. 27, 220–227 (2003).
Leu, S. B. et al. Pathogenic Yersinia DNA in intestinal specimens of pediatric patients with Crohn’s disease. Fetal Pediatr. Pathol. 32, 367–370 (2013).
Navaneethan, U., Venkatesh, P. G. & Shen, B. Clostridium difficile infection and inflammatory bowel disease: understanding the evolving relationship. World J. Gastroenterol. 16, 4892–4904 (2010).
Gray, D. H., Gavanescu, I., Benoist, C. & Mathis, D. Danger-free autoimmune disease in Aire-deficient mice. Proc. Natl Acad. Sci. USA 104, 18193–18198 (2007).
Chinen, T., Volchkov, P. Y., Chervonsky, A. V. & Rudensky, A. Y. A critical role for regulatory T cell-mediated control of inflammation in the absence of commensal microbiota. J. Exp. Med. 207, 2323–2330 (2010).
He, B. et al. Resetting microbiota by Lactobacillus reuteri inhibits T reg deficiency-induced autoimmunity via adenosine A2A receptors. J. Exp. Med. 214, 107–123 (2017).
Kawamoto, S. et al. Foxp3(+) T cells regulate immunoglobulin a selection and facilitate diversification of bacterial species responsible for immune homeostasis. Immunity 41, 152–165 (2014).
Dobes, J. et al. Gastrointestinal autoimmunity associated with loss of central tolerance to enteric alpha-defensins. Gastroenterology 149, 139–150 (2015).
Hetemaki, I. et al. Anticommensal responses are associated with regulatory T cell defect in autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy patients. J. Immunol. 196, 2955–2964 (2016).
Zhang, H., Sparks, J. B., Karyala, S. V., Settlage, R. & Luo, X. M. Host adaptive immunity alters gut microbiota. ISME J. 9, 770–781 (2015).
Rigoni, R. et al. Intestinal microbiota sustains inflammation and autoimmunity induced by hypomorphic RAG defects. J. Exp. Med. 213, 355–375 (2016).
Zhang, L., Li, Y. Y., Tang, X. & Zhao, X. Faecal microbial dysbiosis in children with Wiskott-Aldrich syndrome. Scand. J. Immunol. 91, e12805 (2020).
Ohya, T. et al. Childhood-onset inflammatory bowel diseases associated with mutation of Wiskott-Aldrich syndrome protein gene. World J. Gastroenterol. 23, 8544–8552 (2017).
Packey, C. D. & Sartor, R. B. Commensal bacteria, traditional and opportunistic pathogens, dysbiosis and bacterial killing in inflammatory bowel diseases. Curr. Opin. Infect. Dis. 22, 292–301 (2009).
Oh, J. et al. The altered landscape of the human skin microbiome in patients with primary immunodeficiencies. Genome Res. 23, 2103–2114 (2013).
Kelsen, J. R., Baldassano, R. N., Artis, D. & Sonnenberg, G. F. Maintaining intestinal health: the genetics and immunology of very early onset inflammatory bowel disease. Cell Mol. Gastroenterol. Hepatol. 1, 462–476 (2015).
Serra, E. G. et al. Somatic mosaicism and common genetic variation contribute to the risk of very-early-onset inflammatory bowel disease. Nat. Commun. 11, 995 (2020).
Dominguez-Bello, M. G. et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc. Natl Acad. Sci. USA 107, 11971–11975 (2010).
Charbit-Henrion, F. et al. Diagnostic yield of next-generation sequencing in very early-onset inflammatory bowel diseases: a multicentre study. J. Crohns Colitis 12, 1104–1112 (2018).
Kraaij, M. D. et al. Induction of regulatory T cells by macrophages is dependent on production of reactive oxygen species. Proc. Natl Acad. Sci. USA 107, 17686–17691 (2010).
Schwerd, T. et al. NOX1 loss-of-function genetic variants in patients with inflammatory bowel disease. Mucosal Immunol. 11, 562–574 (2018).
Sokol, H. et al. Intestinal dysbiosis in inflammatory bowel disease associated with primary immunodeficiency. J. Allergy Clin. Immunol. 143, 775–778 (2019).
Clarke, E. L. et al. T cell dynamics and response of the microbiota after gene therapy to treat X-linked severe combined immunodeficiency. Genome Med. 10, 70 (2018).
Aagaard, K. et al. The placenta harbors a unique microbiome. Sci. Transl. Med. 6, 237ra65 (2014).
Vieira, A. T., Fukumori, C. & Ferreira, C. M. New insights into therapeutic strategies for gut microbiota modulation in inflammatory diseases. Clin. Transl. Immunol. 5, e87 (2016).
Lozupone, C. A., Stombaugh, J. I., Gordon, J. I., Jansson, J. K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230 (2012).
Walker, A. W. et al. Dominant and diet-responsive groups of bacteria within the human colonic microbiota. ISME J. 5, 220–230 (2011).
Hill, C. et al. Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 11, 506–514 (2014).
Liu, Y., Tran, D. Q. & Rhoads, J. M. Probiotics in disease prevention and treatment. J. Clin. Pharm. 58(Suppl. 10), S164–S179 (2018).
Liu, Y., Alookaran, J. J. & Rhoads, J. M. Probiotics in autoimmune and inflammatory disorders. Nutrients 10, 1537 (2018).
Strisciuglio, C. et al. Bifidobacteria enhance antigen sampling and processing by dendritic cells in pediatric inflammatory bowel disease. Inflamm. Bowel. Dis. 21, 1491–1498 (2015).
Serena, G. & Fasano, A. Use of probiotics to prevent celiac disease and IBD in pediatrics. Adv. Exp. Med. Biol. 1125, 69–81 (2019).
Miele, E. et al. Nutrition in pediatric inflammatory bowel disease: A Position Paper on Behalf of the Porto Inflammatory Bowel Disease Group of the European Society of Pediatric Gastroenterology, Hepatology and Nutrition. J. Pediatr. Gastroenterol. Nutr. 66, 687–708 (2018).
Savino, F. et al. Lactobacillus reuteri DSM 17938 in infantile colic: a Randomized, Double-Blind, Placebo-Controlled Trial. Pediatrics 126, e526–e533 (2010).
Dinleyici, E. C. et al. Lactobacillus reuteri DSM 17938 shortens acute infectious diarrhea in a pediatric outpatient setting. J. Pediatr. 91, 392–396 (2015).
Fatheree, N. Y. et al. Lactobacillus reuteri for infants with colic: a Double-Blind, Placebo-Controlled, Randomized Clinical Trial. J. Pediatr. 191, 170–178 (2017).
Ohta, A. & Sitkovsky, M. Extracellular adenosine-mediated modulation of regulatory T cells. Front. Immunol. 5, 304 (2014).
Romio, M. et al. Extracellular purine metabolism and signaling of CD73-derived adenosine in murine Treg and Teff cells. Am. J. Physiol. Cell Physiol. 301, C530–C539 (2011).
Liu, Y. et al. Lactobacillus reuteri DSM 17938 feeding of healthy newborn mice regulates immune responses while modulating gut microbiota and boosting beneficial metabolites. Am. J. Physiol. Gastrointest. Liver Physiol. 317, G824–G838 (2019).
Antonioli, L., Pacher, P., Vizi, E. S. & Hasko, G. CD39 and CD73 in immunity and inflammation. Trends Mol. Med. 19, 355–367 (2013).
He, B., Hoang, T. K., Tran, D. Q., Rhoads, J. M. & Liu, Y. Adenosine A2A receptor deletion blocks the beneficial effects of Lactobacillus reuteri in regulatory T-deficient scurfy mice. Front. Immunol. 8, 1680 (2017).
Santoni de Sio, F. R. et al. Role of human forkhead box P3 in early thymic maturation and peripheral T-cell homeostasis. J. Allergy Clin. Immunol. 142, 1909–1921 (2018).
Goodwin, M. et al. CRISPR-based gene editing enables FOXP3 gene repair in IPEX patient cells. Sci. Adv. 6, eaaz0571 (2020).
Bacchetta, R. et al. Defective regulatory and effector T cell functions in patients with FOXP3 mutations. J. Clin. Invest. 116, 1713–1722 (2006).
Bacchetta, R., Barzaghi, F. & Roncarolo, M. G. From IPEX syndrome to FOXP3 mutation: a lesson on immune dysregulation. Ann. NY Acad. Sci. 1417, 5–22 (2018).
Blair, P. J. et al. CD4+CD8- T cells are the effector cells in disease pathogenesis in the scurfy (sf) mouse. J. Immunol. 153, 3764–3774 (1994).
Suscovich, T. J., Perdue, N. R. & Campbell, D. J. Type-1 immunity drives early lethality in scurfy mice. Eur. J. Immunol. 42, 2305–2310 (2012).
Caudy, A. A., Reddy, S. T., Chatila, T., Atkinson, J. P. & Verbsky, J. W. CD25 deficiency causes an immune dysregulation, polyendocrinopathy, enteropathy, X-linked-like syndrome, and defective IL-10 expression from CD4 lymphocytes. J. Allergy Clin. Immunol. 119, 482–487 (2007).
Garg, G. et al. Type 1 diabetes-associated IL2RA variation lowers IL-2 signaling and contributes to diminished CD4+CD25+ regulatory T cell function. J. Immunol. 188, 4644–4653 (2012).
Goudy, K. et al. Human IL2RA null mutation mediates immunodeficiency with lymphoproliferation and autoimmunity. Clin. Immunol. 146, 248–261 (2013).
d’Azzo, A., Bongiovanni, A. & Nastasi, T. E3 ubiquitin ligases as regulators of membrane protein trafficking and degradation. Traffic 6, 429–441 (2005).
Liu, Y. C. The E3 ubiquitin ligase Itch in T cell activation, differentiation, and tolerance. Semin. Immunol. 19, 197–205 (2007).
Roychoudhuri, R. et al. BACH2 represses effector programs to stabilize T(reg)-mediated immune homeostasis. Nature 498, 506–510 (2013).
Buchbinder, E. I. & Desai, A. CTLA-4 and PD-1 pathways: similarities, differences, and implications of their inhibition. Am. J. Clin. Oncol. 39, 98–106 (2016).
Charbonnier, L. M. et al. Regulatory T-cell deficiency and immune dysregulation, polyendocrinopathy, enteropathy, X-linked-like disorder caused by loss-of-function mutations in LRBA. J. Allergy Clin. Immunol. 135, 217–227 (2015).
Alroqi, F. J. et al. DOCK8 deficiency presenting as an IPEX-like disorder. J. Clin. Immunol. 37, 811–819 (2017).
Biggs, C. M., Keles, S. & Chatila, T. A. DOCK8 deficiency: insights into pathophysiology, clinical features and management. Clin. Immunol. 181, 75–82 (2017).
Charbit-Henrion, F. et al. Deficiency in mucosa-associated lymphoid tissue lymphoma translocation 1: a novel cause of IPEX-like syndrome. J. Pediatr. Gastroenterol. Nutr. 64, 378–384 (2017).
Brustle, A. et al. MALT1 is an intrinsic regulator of regulatory T cells. Cell Death Differ. 24, 1214–1223 (2017).
Liu, L. et al. Gain-of-function human STAT1 mutations impair IL-17 immunity and underlie chronic mucocutaneous candidiasis. J. Exp. Med. 208, 1635–1648 (2011).
Uzel, G. et al. Dominant gain-of-function STAT1 mutations in FOXP3 wild-type immune dysregulation-polyendocrinopathy-enteropathy-X-linked-like syndrome. J. Allergy Clin. Immunol. 131, 1611–1623 (2013).
Weinacht, K. G. et al. Ruxolitinib reverses dysregulated T helper cell responses and controls autoimmunity caused by a novel signal transducer and activator of transcription 1 (STAT1) gain-of-function mutation. J. Allergy Clin. Immunol. 139, 1629–1640 (2017).
Haapaniemi, E. M. et al. Autoimmunity, hypogammaglobulinemia, lymphoproliferation, and mycobacterial disease in patients with activating mutations in STAT3. Blood 125, 639–648 (2015).
Harris, T. J. et al. Cutting edge: an in vivo requirement for STAT3 signaling in TH17 development and TH17-dependent autoimmunity. J. Immunol. 179, 4313–4317 (2007).
Milner, J. D. et al. Early-onset lymphoproliferation and autoimmunity caused by germline STAT3 gain-of-function mutations. Blood 125, 591–599 (2015).
Jenks, J. A. et al. Differentiating the roles of STAT5B and STAT5A in human CD4+ T cells. Clin. Immunol. 148, 227–236 (2013).
Kanai, T., Jenks, J. & Nadeau, K. C. The STAT5b pathway defect and autoimmunity. Front. Immunol. 3, 234 (2012).
Anderson, M. S. et al. Projection of an immunological self shadow within the thymus by the aire protein. Science 298, 1395–1401 (2002).
Guo, C. J., Leung, P. S. C., Zhang, W., Ma, X. & Gershwin, M. E. The immunobiology and clinical features of type 1 autoimmune polyglandular syndrome (APS-1). Autoimmun. Rev. 17, 78–85 (2018).
Laakso, S. M. et al. Regulatory T cell defect in APECED patients is associated with loss of naive FOXP3(+) precursors and impaired activated population. J. Autoimmun. 35, 351–357 (2010).
Cassani, B. et al. Defect of regulatory T cells in patients with Omenn syndrome. J. Allergy Clin. Immunol. 125, 209–216 (2010).
Lee, Y. N. et al. Characterization of T and B cell repertoire diversity in patients with RAG deficiency. Sci. Immunol. 1, eaah6109 (2016).
de la Chapelle, A., Herva, R., Koivisto, M. & Aula, P. A deletion in chromosome 22 can cause DiGeorge syndrome. Hum. Genet. 57, 253–256 (1981).
McDonald-McGinn, D. M. et al. Phenotype of the 22q11.2 deletion in individuals identified through an affected relative: cast a wide FISHing net! Genet. Med. 3, 23–29 (2001).
Sullivan, K. E., McDonald-McGinn, D. & Zackai, E. H. CD4(+) CD25(+) T-cell production in healthy humans and in patients with thymic hypoplasia. Clin. Diagn. Lab. Immunol. 9, 1129–1131 (2002).
Adriani, M. et al. Impaired in vitro regulatory T cell function associated with Wiskott-Aldrich syndrome. Clin. Immunol. 124, 41–48 (2007).
Marangoni, F. et al. WASP regulates suppressor activity of human and murine CD4(+)CD25(+)FOXP3(+) natural regulatory T cells. J. Exp. Med. 204, 369–380 (2007).
Acknowledgements
The study was supported by NIH/NCCIH R01AT007083 and NIH/NIAID R03AI117442. The probiotic Lactobacillus reuteri DSM 17938 was obtained as a gift from BioGaia AB.
Author information
Authors and Affiliations
Contributions
Y.L. has contributed to the conception and the structure. J.F. and S.A.A. have contributed to the acquisition and summarization of literature. Y.L. drafted the article. D.QT. and J.M.R. revised it critically for important intellectual content. All authors have reviewed and edited the article and approved the final version to be published.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, Y., Freeborn, J., Armbrister, S.A. et al. Treg-associated monogenic autoimmune disorders and gut microbial dysbiosis. Pediatr Res 91, 35–43 (2022). https://doi.org/10.1038/s41390-021-01445-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-021-01445-2
This article is cited by
-
Probiotic-Derived Ecto-5'-Nucleotidase Produces Anti-Inflammatory Adenosine Metabolites in Treg-Deficient Scurfy Mice
Probiotics and Antimicrobial Proteins (2023)