Abstract
Background
Preterm birth and its complications are the primary cause of death among children under the age of 5. Among the survivors, morbidity both perinatally and later in life is common. The dawn of novel technical platforms for comprehensive and sensitive analysis of protein profiles in blood has opened up new possibilities to study both health and disease with significant clinical accuracy, here used to study the preterm infant and the physiological changes of the transition from intrauterine to extrauterine life.
Methods
We have performed in-depth analysis of the protein profiles of 14 extremely preterm infants using longitudinal sampling. Medical variables were integrated with extensive profiling of 448 unique protein targets.
Results
The preterm infants have a distinct unified protein profile in blood directly at birth regardless of clinical background; however, the pattern changed profoundly postnatally, expressing more diverse profiles only 1 week later and further on up to term-equivalent age. Clusters of proteins depending on temporal trend were identified.
Conclusion
The protein profiles and the temporal trends here described will contribute to the understanding of the physiological changes in the intrauterine−extrauterine transition, which is essential to adjust early-in-life interventions to prone a normal development in the vulnerable preterm infants.
Impact
-
We have performed longitudinal and in-depth analysis of the protein profiles of 14 extremely preterm infants using a novel multiplex protein analysis platform.
-
The preterm infants had a distinct unified protein profile in blood directly at birth regardless of clinical background.
-
The pattern changed dramatically postnatally, expressing more diverse profiles only 1 week later and further on up to term-equivalent age.
-
Certain clusters of proteins were identified depending on their temporal trend, including several liver and immune proteins.
-
The study contributes to the understanding of the physiological changes in the intrauterine−extrauterine transition.
Similar content being viewed by others
Introduction
The number of preterm births is rising worldwide; nowadays 15 million infants are estimated to be born preterm each year, before 37 weeks of gestation.1,2 The aetiology behind preterm birth as well as the rising numbers is multifactorial though not yet fully understood.3,4 Preterm birth and its complications, such as respiratory insufficiency, intraventricular haemorrhages and necrotizing enterocolitis, are causing approximately one million deaths each year, making it the primary cause of death among children under the age of 5 (18%).5,6,7 Among the survivors, numerous preterm infants develop chronic diseases of heart, lungs, intestines, eyes and the neural system among others, resulting in preterm birth being one of most common reasons for morbidity both perinatally and later in life.6 Today, it is not known which infants are at risk of developing which disease, since few reliable predictors exist. Nonetheless, the first weeks and even days in life are believed to be determinant even though the precise physiological pathways are not yet identified.8,9
Recently, several studies have been conducted to explore omics profiles in preterm infants. Olin et al. showed that the immune system in human newborn infants converged in response to diverse postnatal exposures, based on the analysis of the immune cell population using mass cytometry and the protein profiles using proximity extension assays (PEA).8,10 Both cells and proteins were shown to undergo drastic changes after birth, following a stereotypic pattern. Similarly, Lynch et al.11 measured more than 1000 plasma proteins in the early neonatal period in infants at risk for retinopathy of prematurity (ROP) using an aptamer-based proteomic technology to identify proteins predictive of subsequent development of clinically significant ROP. The study identified several proteins that may be targets for future therapeutic interventions. In addition, Suski et al.12 showed by mass spectrometry-based analysis of the plasma proteomes that preterm infants have a multitude of time-related differences in protein abundances. They suggested profound impact of gestation duration on the ability to bridge the gap in systemic homoeostasis after preterm delivery. These results demonstrate the power of comprehensive omics profiles to identify proteins of potential for precision medicine efforts, although follow-up studies are needed to validate the targets for either diagnostic or therapeutic applications.
Here, we have used a large panel of protein assays based on the PEA principle to investigate the protein profiles in extremely preterm infants born between gestation weeks 22 and 27.10 The infants were sampled repeatably during several months to allow a comprehensive analysis of changes in protein concentrations in blood during the first months after birth.
Methods
Patients and nutritional management
The Donna Mega Study was a randomized, open-label, controlled trial conducted at a single site in Sweden designed to compare the effects of two parenteral fatty acid solutions. The trial was approved by the Regional Ethical Board, Gothenburg (Dnr 303-11, T336-18) at the University of Gothenburg. Informed written consent was obtained for all participants from their parents or guardians. The protocol for the Donna Mega study is available at clinicaltrial.gov (Clinical trial NCT 02760472). The major outcome, evaluating the impact of parenteral nutrition with and without fish oil on morbidities and growth, was described in Najm et al.13 Infants born at a gestational age (GA) < 28 weeks were enrolled from the neonatal intensive care unit at Sahlgrenska University Hospital in Gothenburg, Sweden, from April 2013 to September 2015. Parents of 90 of 138 eligible infants agreed to participation after informed consent. The nutritional strategy has been described previously.13,14 Briefly, all infants received parenteral and enteral nutrition according to clinical routine. Parenteral nutrition was initiated as soon as possible after birth. The parenteral lipid dosing strategy was to deliver a dose of 2–3 g/kg body weight every 24 h. Enteral nutrition used either maternal or donor breast milk with individualized fortification. Minimal enteral feeding was started within 3 h of birth and administered every 2−3 h (1–2 mL/meal) with a gradual increase in volume. Seventy-eight subjects fulfilled the criteria for final evaluation, surviving to 40 weeks postmenstrual age (PMA). From these, 14 individuals were chosen to represent an even distribution of gestational age at birth, sex, size at birth, delivery mode, morbidities (bronchopulmonary dysplasia (BPD), necrotizing enterocolitis (NEC), retinopathy of prematurity (ROP), postnatal inflammation), days with parenteral nutrition, number of transfusions and availability of serial serum samples from all time points (n = 7), at postnatal days 1, 7, 14, and 28 and at PMA 32, 36, and 40 weeks.
Serum protein profiling
Serum proteins were analysed using a multiplex PEA technology (Olink Bioscience, Uppsala, Sweden) previously described.10 Briefly, each kit consisted of a microtiter plate for measuring 92 protein biomarkers in all 88 samples and each well contained 96 pairs of DNA-labeled antibody probes. To minimize inter- and intra-run variation, the data were normalized using both an internal control (extension control) and an interplate control, and then transformed using a pre-determined correction factor. The pre-processed data were provided in the arbitrary unit Normalized Protein Expression (NPX) on a log2 scale and where a high NPX represents high protein concentration. In this study five Olink panels have been used including Cardiometabolic, Cardiovascular II, Cardiovascular III, Development, and Metabolism resulting in a total number of 460 protein assays. Samples and proteins that did not pass the quality control were removed. A total of 448 unique proteins from 88 samples were kept for the analysis (see Supplementary Table S4 (online)).
Hierarchical clustering and UMAP analysis
For clustering, the expression profiling of each protein was first standardized with a standard deviation of 1 centred at 0. The scaled values from all 88 samples were used to create the Euclidean distance matrix for dendrogram generation. Dendrograms showing gene expression in heatmaps have been clustered using the Ward2 algorithm, an implementation of Ward’s minimum variance method implemented as “Ward.D2” in R package pheatmap.15,16 Uniform Manifold Approximation and Projection (UMAP) has been performed on NPX values of samples by using the R packages umap with default parameters.17
Quantification of the protein variability
Interquartile range (IQR) was used to evaluate the dispersion pattern of protein expression across seven visits. NPX values of proteins were first standardized with a standard deviation of 1 centred at 0 within subjects before applying IQR. The calculation of IQR was performed using the R package stats (see Supplementary Table S5 (online)).
Time-series expression analysis
Mixed-effect modelling was performed using the lme4 package7 and Kenward−Roger approximation was used to calculate p values which were subsequently adjusted for multiple testing based on false discovery rate.18 p values were considered significant if <0.05. In total, 338 significantly expressed proteins across seven visits were identified. For the hierarchical clustering analysis, the mean NPX of each protein was used to create the dendrogram based on Euclidean distance. Ward’s minimum variance method was implemented as “Ward.D2” in the hclust function in the R package stats, where clusters are chosen at each stage such that the increase in cluster variance is minimized after merging. As a result, six clusters were identified. For regression analysis, the expression profiling of each protein in the certain cluster was standardized with a standard deviation of 1 centred at 0. Non-parametric local weighted regression (LOESS) was applied to generate the regression curve.
Gene ontology (GO) enrichment analysis
GO enrichment analysis was performed using the R package clusterProfiler version 3.10.1.19 clusterProfiler implements a hypergeometric model to test for gene-set overrepresentation relative to a background gene-set. Enrichment of proteins in each cluster (1−6) from the time-course clustering analysis was analysed using the compareCluster function with fun = enrichGO, pvalueCutoff = 0.01, OrgDb = org.Hs.eg.db. Significantly enriched GO terms were simplified using gofilter function with level = 4.
Differential expression analysis
Differential expression analysis was conducted by using NPX with inflammatory conditions and the usage of steriods as covariates. The R package limma, which includes lmFit, eBayes, and topTable functions, was used for pairwise comparison of differentially expressed genes (DEGs).20 False discovery rate (FDR) was calculated by using p.adjust() function in R, which uses Benjamini−Hochberg method. Genes with FDRs < 0.01 and absolute log2 fold-change > 0.5 were considered as DEGs.
Tissue enrichment analysis
Tissue-specific gene sets are defined using the findings of tissue-elevated genes from the Human Protein Atlas.21,22 The Generally Applicable Gene-set Enrichment (GAGE) package in R was utilized to perform tissue enrichment analysis.22,23 All tissue-elevated genes were used as a reference. The global p values were then adjusted by using Benjamini & Hochberg method. Tissues with p values < 0.01 were considered as significantly enriched tissues.
Results
The study cohort
The study cohort consisted of extremely preterm infants born before 28 weeks of gestation, dated at ultrasonography examination. A total of 14 infants were chosen from a larger cohort to represent a diversion of gestational age, sex, birth weight, birth size, delivery mode and morbidity among others (Supplementary Tables S1 and S2 (online)). During seven planned appointments from birth to term-equivalent age (40 weeks postmenstrual age), 88 peripheral serum samples were collected (Supplementary Table S3 (online)). Eight subjects were male and six were female, seven were delivered vaginally and seven by caesarean section (Fig. 1a). The birth weights varied from 415 to 1235 g (Fig. 1b, c).
a The study cohort consisted of 14 extremely preterm infants born between gestational week 22 + 6 to 27 + 4. During seven planned visits, from birth to term-equivalent age, 88 peripheral serum samples were collected. Eight subjects were male and six female, seven were delivered vaginally and seven by caesarean section. b The relationship between birth weight and gestational age in weeks for each of the 14 subjects. The birth weights varied from 415 to 1235 g. c Growth in weight (g) from birth to 40 weeks postmenstrual age. See Supplementary Figs. S1 and S2 for longitudinal head circumference and length measurements. d Protein levels of CR2 showed a unified pattern, increasing over the seven visits for all 14 subjects. e Protein levels of COLEC12 as an example of a decreasing trend over time for all 14 subjects.
Proteome profiling using protein extension assay
Blood serum from the subjects were analysed using five different PEA panels to obtain the relative protein expression levels of a total of 448 unique protein targets as normalized protein expression (NPX).10 The assays included panels of proteins related to metabolism, development, cardiometabolic and cardiovascular diseases. The results of all proteins in all samples are presented in Supplementary Table S4 (online). An example of temporal variation of serum protein concentration during the various visits is presented with complement C3d receptor 2 (CR2), displaying uniform increasing levels after birth in all subjects (Fig. 1d). This protein is a member-bound receptor involved in the defence against viral infections.24 An opposite example with decreased levels after birth is the collectin subfamily member 12 (COLEC12), a cell surface glycoprotein mainly expressed in placenta and umbilical cord, associated with several functions for host defence (Fig. 1e).25
Longitudinal analysis of the blood profiles
Unsupervised hierarchical clustering analysis was performed on the complete protein profiles from the 14 infants. Overall, the expression patterns of the 88 samples demonstrated clustering based on the time after birth, with samples from the first visit forming one branch that is distinct from the branch comprising later visits (Fig. 2a). This suggests profound changes in serum protein levels during the first week of life. The Uniform Manifold Approximation and Projection (UMAP) analysis also demonstrates a similar pattern, as the first visit stands out as an outlier compared to other visits regardless of clinical differences, sex and delivery mode among the subjects (Fig. 2b, Supplementary Figs. S3−S6 (online)).17
To assess the variability of protein expression over time, we quantified the dispersion level of each protein based on all samples by examining their interquartile ranges (IQRs) (Supplementary Table S5 (online)). The median expression of protein is independent of the variability of protein expression (Supplementary Fig. S7 (online)). In Fig. 2c, d, the most variable and most stable proteins, respectively, are presented as boxplots showing the expression levels across seven visits. The most variable protein in the study is fibroblast growth factor 21 (FGF-21) with increasing expression levels after birth, which is in line with the report that elevated FGF-21 serum levels during the first week of life are independently associated with postnatal growth failure in preterm infants.26
Changed protein levels after birth
To get a comprehensive overview of the dynamic changes of protein expression over time, linear mixed-effect modelling was performed based on the longitudinal proteome profiles, including sex and subjects as covariates. In total, 338 out of the 448 proteins were identified with significant differences across time during the longitudinal study (Supplementary Table S6 (online)). A cluster analysis further suggested that the expression profiles across these 338 proteins could be stratified into six expression clusters (Fig. 3a, Supplementary Table S7 (online)). Note that clusters 2 and 4 are highly similar showing an upward trend followed by a downward trend, while clusters 5 and 6 are also mutually similar, displaying an overall downward trend in protein levels throughout the study period. Examples of longitudinal expression levels for proteins belonging to each of the six clusters are presented in Fig. 3b. A functional analysis using Gene Ontology (GO) term enrichment of proteins within each pattern was performed and the results are presented in Supplementary Table S8 (online).27
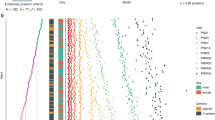
a Overview of the expression profiles of all proteins in each of the six clusters based on mean scaled NPX values for all subjects in each of the seven visits for a protein (grey). The coloured bold line represents the regression line based on all proteins and the total number of proteins in each cluster is also indicated in the top right part. b Examples of representative protein expression patterns for one protein from each cluster with the expression level for all 14 subjects. c Time-course expression analysis of significant proteins from mixed-effect modelling including a heatmap presenting the scaled expression based on mean NPX values for all subjects and visits. The dendrogram shows hierarchical clustering based on Euclidean distance of the protein expression (see Supplementary Fig. S14 for expression heatmap across postnatal age).
The first cluster (red) consisting of 97 proteins shows an upward trend during the first visits, the protein levels then stabilize or slightly decrease during later visits (Fig. 3a, Supplementary Fig. S8 (online)). An example (Fig. 3b) in this cluster is C−C motif chemokine ligand 24 (CCL24), a protein belonging to a family of secreted proteins involved in immunoregulatory and inflammatory processes.28 A GO analysis of all proteins in this cluster shows functions associated with blood coagulation and epithelial cell proliferation (Supplementary Table S8 (online)).27,29 The second cluster (light green) consisting of 34 proteins also presents an upward trend, but this is followed by a sharp downward trend during the later visits (Fig. 3a, Supplementary Fig. S9 (online)). An example from this cluster is the fibroblast activating protein alpha (FAP) (Fig. 3b), a serine protease thought to be involved in control of fibroblast growth and epithelial−mesenchymal interactions during development.30 A functional analysis of all proteins in cluster 2 shows that these proteins are enriched for extracellular structure organization, probably associated with endothelial cell adhesion (Supplementary Table S8 (online)).31 The third cluster (orange) consisting of 56 proteins displays increasing protein levels throughout the study period (Fig. 3a, Supplementary Fig. S10 (online)), exemplified by dipeptidyl peptidase 4 (DPP4) (Fig. 3b) involved in immune regulation. This protein has previously been reported to be elevated in preterm infants with no correlation to cord blood or venous blood of mothers.32 A functional GO analysis of all proteins in cluster 3 shows that these proteins are enriched for the process of leukocyte recruitment and the response to hypoxia (Supplementary Table S8 (online)). This is consistent with a previous report that suggested that the preterm immune system could generate an outburst of naive T and B lymphocytes from the thymus and bone marrow during the postnatal development.33 In addition, leptin (LEP) involved in weight gain can be found in this cluster.22 The fourth cluster (violet) consisting of 32 proteins is similar to cluster 2 (Fig. 3a, Supplementary Fig. S11 (online)) but reveals an even sharper decline in protein levels after the first visits as exemplified by surfactant protein D (PSP-D) (Fig. 3b), a protein invloved in the innate immune response and highly similar to surfactant B and C often given to preterm infants to improve lung capacity.34 The fifth cluster (blue) consisting of 56 proteins shows similar levels during the first visits and this is subsequently followed by a decreasing trend of protein levels (Fig. 3a, Supplementary Table S12 (online)). An example of protein in this cluster is delta-like non-canonical notch ligand 1 (DLK1) (Fig. 3b), which is a regulator of cell growth involved in the induction of several cell types, including adipocytes.35 A functional analysis of all proteins in this cluster shows enrichment for labyrinthine layer morphogenesis, placenta morphogenesis and the regulation of external stimulus (Supplementary Table S8 (online)). The sixth cluster (green) consisting of 63 proteins presents decreasing protein levels throughout the study period (Fig. 3a, Supplementary Fig. S13 (online)) as exemplified by the alpha protein of glycoprotein hormone (CGA) (Fig. 3b), also known as FSHA, GPHa, HCG, LHA and TSHA. CGA is produced in placenta and pituatory glands.22 A functional analysis of all proteins in cluster 6 yields similar functions as cluster 5 (Supplementary Table S8 (online)).
In Fig. 3c, a time-course expression analysis is presented for the most significant variable genes from the mixed-effect modelling. Altogether expression levels of 338 proteins are shown as a heatmap throughout the seven visits. The clusters described in Fig. 3a can be observed as dendrograms presenting proteins with similar expression pattern.
Tissue origin of the variable proteins
In Fig. 4a, a volcano plot visualizes the proteins with significant increase or decrease in levels directly after birth. Two proteins showed the highest fold-change (increase) during the first week: C−C motif chemokine ligand 16 (CCL16) and the FGF-21, both annotated as secreted proteins specific (enriched) for the liver (Supplementary Table S11 (online)).22 The latter protein has been shown to stimulate the uptake of glucose in adipose tissues.36 The tissue origin of the variable proteins after birth was analysed based on the tissue specificity score of the Human Protein Atlas.22 The tissue enrichment analysis summarized in Fig. 4b shows that many proteins with decreasing levels of protein after birth are of placental origin, while many proteins increasing have been annotated of blood, lymphoid or liver origin. A chord diagram was therefore generated (Fig. 4c) in which the variable proteins are connected to their tissue of origin. The results confirm that many of the proteins increasing after birth have been classified as enriched in liver, lymphoid tissue or blood cells, while many of the proteins decreased have been classified as enriched in placenta.
a Volcano map of differentially expressed proteins in the first visit compared to the other visits based on NPX values of proteins from all subjects and visits. The X-axis represents log2 fold-change (FC) and the Y-axis represents log10 (adjusted p values). Increased proteins (log2FC > 0.5, adjusted p values < 0.01) after birth are shown as red dots, decreased proteins (log2FC < −0.5, adjusted p values < 0.01) after birth are shown as blue dots. b Tissue enrichment analysis results of differentially expressed proteins in the first visit. The X-axis represents the average fold-change of the tissue-specific gene-set and the Y-axis represents the significantly enriched tissues with adjusted p values (Benjamini and Hochberg method) <0.01. The size of the node represents the −log10 (adjusted p value) in a specific tissue. c Chord diagram presenting the tissue specificity of differentially expressed proteins in the first visit. Each link represents the number of proteins that are elevated in the linked tissue.
Proteins increased after birth
The analysis above shows that many proteins increasing after birth are of liver origin. In Fig. 5a, the 21 liver elevated proteins from the chord diagram are presented with a clear trend from low expression during the first day to increasing levels in the subsequent visits. This includes proteins involved in complementation and coagulation factors (complement C3d receptor 2 (C2), coagulation factor VII (F7) and coagulation factor XI (F11)), but also proteins mediating new blood vessel formation angiogenin (ANG).22 The tyrosine-binding protein (SERPINA7) is responsible for carrying thyroid hormones T3 and T4 in the blood, while inhibin beta C subunit (INHBC) is a member of the TGF-beta (transforming growth factor) superfamily of proteins (Fig. 5d).22 The list suggests that the liver is rapidly activated to generate proteins involved in immune response.
Heatmaps showing the increasing protein levels of a 21 liver elevated proteins and b 27 blood or lymphoid tissue-elevated proteins after birth as well as c decreasing protein levels of 8 placenta elevated proteins after birth. Examples of representative protein expression pattern for two proteins in d liver elevated proteins, e blood or lymphoid tissue-elevated proteins and f placenta elevated proteins with the expression levels for all 14 subjects.
Similarly, many immune molecules originating from blood are shown to be elevated after birth (Fig. 5b). Selectin L (SELL) (Fig. 5e) encodes a cell surface adhesion molecule required for leucocytes interaction with endothelial cells facilitating their migration into inflammation sites.22 The interleukin 2 receptor alpha (IL2-RA) (Fig. 5e) is also a molecule involved in immunological response being a component of the high affinity IL-2 receptor.22 Normally, this is an integral-membrane protein but soluble IL2-RA has been isolated from blood. Several immune-related molecules, such as C−C motif chemokine ligand 18 (CCL18), C−C motif chemokine ligand 24 (CCL24), Fc receptor like 1 (FCRL1), bone marrow stromal cell antigen 1 (BST1) and interleukin 7 receptor (IL7R), were also found to be elevated after the first week in the preterm infants. In addition, several complement related proteins (CR2, Glycoprotein VI platelet (GP6)) were elevated.
Proteins decreased after birth
The proteins decreasing the first week after preterm birth are visualized in the volcano plot (Fig. 4a). Interestingly, many of the proteins that demonstrate a steady decrease in concentration after birth are proteins that are expressed in the placenta.22 In Fig. 5c, a heatmap visualizes expression profiles for ten proteins all annotated to have elevated tissue specificity in the placenta.22 The relative protein levels across the seven visits are higher at birth in comparison with the subsequent visits. The Insulin-like growth factor binding protein 1 (IGFBP1) is a protein secreted to blood (Supplementary Table S12 (online)) mainly produced in placenta and liver.22 Interestingly, this protein is activated in inflammation, infection, hypoxia, and one can again speculate that the high levels at birth are due to an ongoing inflammation or infection, which might be implicated in the preterm birth. Similarly, follistatin like 3 (FSTL3), growth differentiation factor 15 (GDF15), glycoprotein hormone, alpha polypeptide (CGA) and placenta growth factor (PGF) are all produced in placenta and annotated to be secreted to blood (Supplementary Table S12 (online)).22 The individual trends across the preterm infants for GDF15 and PDF are shown in Fig. 5f. Interestingly, two of the proteins, the clathrin adaptor protein (DAB2) and the Xg blood group protein (XG), are not annotated to be secreted, but instead membrane-bound and thus presumably present on the surface of the placental cells and subsequently shreds and transfers to the fetus before birth (Supplementary Table S12 (online)).22 The fact that all of these proteins show a steady decrease in protein levels after birth supports the hypothesis that these proteins are of intrauterine placental origin.
Discussion
The results of this study show dramatic changes in protein profiles during the early weeks of life in extremely preterm infants. The magnitude of this early postnatal individual change in protein expression has to our knowledge not been described before. The analysis confirms patterns of well-known proteins involved in weight gain, including leptin which steadily increased in expression levels throughout development, but more interestingly many protein targets, not previously described in this context, were identified with significant changes in protein levels across time. Highest fold-change is observed for the inflammation marker interleukin 6 (IL-6), which is involved in chronic inflammation and autoimmunity.37 It is tempting to speculate that this is due to an ongoing inflammation of the preterm infants at birth. Different patterns (clusters) of protein expression profiles have been identified, involving more than 300 proteins with high similarities in longitudinal protein profiles across the analysed preterm infants. Particularly, we show a clear distinction in serum protein expression between the samples taken the first day after preterm birth compared to one to several weeks later.
It is important to point out that the study included a limited number of preterm infants, but the fact that the technical platform used only requires a few microlitres of blood will facilitate follow-up studies involving larger cohorts to validate the protein profiles. It should also be emphasized that many subjects received blood products, erythrocytes, plasma or thrombocytes one or several times during the study period and some of the observed trends might be affected by the transfer of blood proteins to the infants. Several follow-up studies to allow more in-depth analysis have been initiated involving larger cohorts and including the effect of various nutritional supplements.
With a range in gestational ages from 22 to 27 weeks and birth weights from 400 to 1200 g, we found a distinct uniform protein expression at birth, without associations to degree of immaturity or degree of intrauterine growth retardation. Thereafter, during the first week of life, there were drastic changes in serum protein expression with large inter-individual variability. Many of the proteins showing an increase after birth have been annotated as of liver origin and the increase of these proteins in the infants’ blood was continuing through the study period. Another interesting observation is that many proteins annotated as of placenta origin were detected, suggesting an intrauterine origin from maternal−fetal interaction with a continuous decrease in levels during the study period.
In conclusion, the comprehensive and sensitive protein profiling platform presented here has allowed us to follow the molecular serum profiles of extremely preterm infants longitudinally after birth. The results show that the extremely preterm infants, regardless of duration or milieu of intrauterine life, have a common protein profile in peripheral blood directly at birth which dramatically changes during their first week of life, resulting in a distinctly altered profile only a week later.
Data availability
All the data used in the study are available in the supplementary material.
References
Chawanpaiboon, S. et al. Global, regional, and national estimates of levels of preterm birth in 2014: a systematic review and modelling analysis. Lancet Glob. Health 7, e37–e46 (2019).
Blencowe, H. et al. National, regional, and worldwide estimates of preterm birth rates in the year 2010 with time trends since 1990 for selected countries: a systematic analysis and implications. Lancet 379, 2162–2172 (2012).
Romero, R. et al. The preterm parturition syndrome. BJOG 113(Suppl 3), 17–42 (2006).
Goldenberg, R. L. et al. Epidemiology and causes of preterm birth. Lancet 371, 75–84 (2008).
Oza, S. et al. Neonatal cause-of-death estimates for the early and late neonatal periods for 194 countries: 2000−2013. Bull. World Health Organ. 93, 19–28 (2015).
Davey, M. A. et al. Risk-scoring systems for predicting preterm birth with the aim of reducing associated adverse outcomes. Cochrane Database Syst. Rev. p. CD004902 (2015).
You, D. et al. Global, regional, and national levels and trends in under-5 mortality between 1990 and 2015, with scenario-based projections to 2030: a systematic analysis by the UN Inter-agency Group for Child Mortality Estimation. Lancet 386, 2275–2286 (2015).
Olin, A. et al. Stereotypic immune system development in newborn children. Cell 174, 1277–1292.e1214 (2018).
Grisaru-Granovsky, S. et al. Population-based trends in mortality and neonatal morbidities among singleton, very preterm, very low birth weight infants over 16 years. Early Hum. Dev. 90, 821–827 (2014).
Assarsson, E. et al. Homogenous 96-plex PEA immunoassay exhibiting high sensitivity, specificity, and excellent scalability. PLoS ONE 9, e95192 (2014).
Lynch, A. M. et al. The relationship of novel plasma proteins in the early neonatal period with retinopathy of prematurity. Invest. Ophthalmol. Vis. Sci. 57, 5076–5082 (2016).
Suski, M. et al. Prospective plasma proteome changes in preterm infants with different gestational ages. Pediatr. Res. 84, 104–111 (2018).
Najm, S. et al. Effects of a lipid emulsion containing fish oil on polyunsaturated fatty acid profiles, growth and morbidities in extremely premature infants: a randomized controlled trial. Clin. Nutr. ESPEN 20, 17–23 (2017).
Hansen-Pupp, I. et al. Influence of insulin-like growth factor I and nutrition during phases of postnatal growth in very preterm infants. Pediatr. Res. 69, 448–453 (2011).
Kaufman, L. & Rousseeuw, P. J. Finding Groups in Data: An Introduction to Cluster Analysis (John Wiley & Sons, 2009).
Murtagh, F. & Legendre, P. Ward’s hierarchical agglomerative clustering method: which algorithms implement Ward’s criterion? J. Classification 31, 274–295 (2014).
McInnes, L., Healy, J. & Melville, J. Umap: uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Bates, D. et al. Fitting linear mixed-effects models using lme4. Preprint at https://arxiv.org/abs/1406.5823 (2014).
Yu, G. et al. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics: J. Integr. Biol. 16, 284–287 (2012).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47–e47 (2015).
Uhlen, M. et al. A genome-wide transcriptomic analysis of protein-coding genes in human blood cells. Science 366.6472 (2019).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347.6220, 1260419 (2015).
Luo, W. et al. GAGE: generally applicable gene set enrichment for pathway analysis. BMC Bioinform. 10, 161 (2009).
Hannan, J. P. The structure-function relationships of complement receptor type 2 (CR2; CD21). Curr. Protein Pept. Sci. 17, 463–487 (2016).
Ma, Y. J. et al. Soluble collectin-12 (CL-12) is a pattern recognition molecule initiating complement activation via the alternative pathway. J. Immunol. 195, 3365–3373 (2015).
Guasti, L. et al. Elevated FGF21 leads to attenuated postnatal linear growth in preterm infants through GH resistance in chondrocytes. J. Clin. Endocrinol. Metab. 99, E2198–E2206 (2014).
Acencio, M. L., Lægreid, A. & Kuiper, M. et al. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330−D338 (2019).
Menzies-Gow, A. et al. Eotaxin (CCL11) and eotaxin-2 (CCL24) induce recruitment of eosinophils, basophils, neutrophils, and macrophages as well as features of early- and late-phase allergic reactions following cutaneous injection in human atopic and nonatopic volunteers. J. Immunol. 169, 2712–2718 (2002).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Wang, X. M. et al. Fibroblast activation protein increases apoptosis, cell adhesion, and migration by the LX-2 human stellate cell line. Hepatology 42, 935–945 (2005).
Wisgrill, L. et al. Endothelial cells of extremely premature infants display impaired immune response after proinflammatory stimulation. Pediatr. Res. 83, 128–134 (2018).
Yakovleva, A. A. et al. Dipeptidylpeptidase 4 (DPP4, CD26) activity in the blood serum of term and preterm neonates with cerebral ischemia. Neuropeptides 52, 113–117 (2015).
Walker, J. C. et al. Development of lymphocyte subpopulations in preterm infants. Scand. J. Immunol. 73, 53–58 (2011).
Soerensen, C. M. et al. Purification, characterization and immunolocalization of porcine surfactant protein D. Immunology 114, 72–82 (2005).
Traustadottir, G. A. et al. The imprinted gene Delta like non-canonical Notch ligand 1 (Dlk1) is conserved in mammals, and serves a growth modulatory role during tissue development and regeneration through Notch dependent and independent mechanisms. Cytokine Growth Factor Rev. 46, 17–27 (2019).
Tezze, C., Romanello, V. & Sandri, M. FGF21 as modulator of metabolism in health and disease. Front. Physiol. 10, 419 (2019).
Tanaka, T., Narazaki, M. & Kishimoto, T. IL-6 in inflammation, immunity, and disease. Cold Spring Harb. Perspect. Biol. 6, a016295 (2014).
Acknowledgements
The main funding was provided from the Erling Persson Foundation (KCAP). We would like to thank the Plasma Profiling Facility at SciLifeLab in Stockholm for conducting the Olink analyses. We acknowledge the entire staff of the Human Protein Atlas program and the Science for Life Laboratory for their valuable contributions.
Author information
Authors and Affiliations
Contributions
M.U., L.F. and A.H. conceived and designed the analysis. A.E., A.H., G.H., H.D. and M.J.K. collected and contributed data to the study. W.Z., H.D., A.T., L.F., M.J.K. and M.U. performed the data analysis. A.H., G.H. and A.E. supplied clinical material. M.U., W.Z., H.D. and L.F. drafted the manuscript. All authors discussed the analyses, results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
The trial was approved by the Regional Ethical Board, Gothenburg (Dnr 303-11, T336-18) at the University of Gothenburg. Informed written consent was obtained for all participants from their parents or guardians.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Zhong, W., Danielsson, H., Tebani, A. et al. Dramatic changes in blood protein levels during the first week of life in extremely preterm infants. Pediatr Res 89, 604–612 (2021). https://doi.org/10.1038/s41390-020-0912-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-020-0912-8
This article is cited by
-
Antisecretory factor in breastmilk is associated with reduced incidence of sepsis in preterm infants
Pediatric Research (2024)
-
The development of blood protein profiles in extremely preterm infants follows a stereotypic evolution pattern
Communications Medicine (2023)
-
Young infants display heterogeneous serological responses and extensive but reversible transcriptional changes following initial immunizations
Nature Communications (2023)
-
Changes in inflammatory proteins following platelet transfusion in a neonatal population
Pediatric Research (2023)
-
Blood protein profiles related to preterm birth and retinopathy of prematurity
Pediatric Research (2022)