Abstract
Bidirectional communication between the gut and brain is well recognized, with data now accruing for a specific role of the gut microbiota in that link, referred to as the microbiome–gut–brain axis. This review will discuss the emerging role of the gut microbiota in brain development and behavior. Animal studies have clearly demonstrated effects of the gut microbiota on gene expression and neurochemical metabolism impacting behavior and performance. Based on these changes, a modulating role of the gut microbiota has been demonstrated for a variety of neuropsychiatric disorders, including depression, anxiety, and movement including Parkinson’s, and importantly for the pediatric population autism. Critical developmental windows that influence early behavioral outcomes have been identified that include both the prenatal environment and early postnatal colonization periods. The clearest data regarding the role of the gut microbiota on neurodevelopment and psychiatric disorders is from animal studies; however, human data have begun to emerge, including an association between early colonization patterns and cognition. The importance of understanding the contribution of the gut microbiota to the development and functioning of the nervous system lies in the potential to intervene using novel microbial-based approaches to treating neurologic conditions. While pathways of communication between the gut and brain are well established, the gut microbiome is a new component of this axis. The way in which organisms that live in the gut influence the central nervous system (CNS) and host behavior is likely to be multifactorial in origin. This includes immunologic, endocrine, and metabolic mechanisms, all of which are pathways used for other microbial–host interactions. Germ-free (GF) mice are an important model system for understanding the impact of gut microbes on development and function of the nervous system. Alternative animal model systems have further clarified the role of the gut microbiota, including antibiotic treatment, fecal transplantation, and selective gut colonization with specific microbial organisms. Recently, researchers have started to examine the human host as well. This review will examine the components of the CNS potentially influenced by the gut microbiota, and the mechanisms mediating these effects. Links between gut microbial colonization patterns and host behavior relevant to a pediatric population will be examined, highlighting important developmental windows in utero or early in development.
Similar content being viewed by others
Behavioral alterations in animal models
Early on, investigators working with germ-free (GF) animals noted behavioral differences when compared to conventionally (CONV) raised mice,1 or mice raised free of specific disease-causing organisms (specific pathogen free; SPF).2 Since then a spectrum of behavioral changes have been identified in GF animals using standardized animal behavioral testing techniques in rodents. These altered behaviors cluster into four domains: social interactions including behavior paralleled to that seen in autism spectrum disorder (ASD); stress-related and anxiety-related responses; learning and memory; and motor control (for a review see Vuong et al.3) While GF mice offer a system to examine the impact of gut microbiota in isolation, there are clearly limitations in applicability, since a GF world in humans does not exist. Additionally in studies using fecal transplantation during early development, there is often a mismatch between age of the GF animal and donor contributing the sample, altering microbial developmental trajectories. As an alternative, orally administered, non-absorbable antimicrobials have been used to alter the gut microbiome in rodents (neomycin, bacitracin, and pimaricin)4,5,6 as well as in zebrafish (amphotericin, kanamycin, and ampicillin).7 Behavioral changes similar to GF animals are reported with oral but not intraperitoneal4 treatment particularly related to stress-associated and anxiety-associated phenotypes. Oral antibiotics have also been used to examine the contribution of the maternal gut microbiota during pregnancy to offspring behavior by maternal treatment either preconception (non-absorbable sulfonamide)5 or early in gestation (neomycin, pimaricin, bacitracin).6 Compared to controls, offspring of antibiotic-treated mothers demonstrated increased anxiety-like behaviors and diminished social interactions. Analysis of fecal samples from antibiotic-exposed offspring demonstrated a 50% decrease in the relative abundance of the order Lactobacillales and increase in the bacterial family Clostridium.6 The behavioral phenotypes in offspring can be only partly rescued by fostering them with normal dams beginning on postnatal day 1, implicating the perinatal period as a critical developmental window.6
The perinatal period
The perinatal period is a critical developmental window characterized by rapid evolution of gut microbial colonization alongside changes in neuronal organization. Given this co-evolution, it is perhaps not surprising that studies using GF mice have been used to examine the association between gut microbiota and central nervous system (CNS) structure and function (Table 1). In a landmark paper in 2004, Sudo et al.2 demonstrated the importance of the perinatal period, identifying that gut microbes were involved in programing the hypothalamic-pituitary-adrenal (HPA) system stress response. Compared to SPF mice, GF animals had an exaggerated HPA response in response to an acute-restraint stress, evidenced by elevated levels of stress response hormones adrenocorticotropic hormone (ACTH) and corticosterone. The exaggerated HPA response was reversed by re-colonizing the GF mice with fecal flora from SPF animals, but only if SPF fecal reconstitution occurred before 6 weeks of age. The presence of early developmental windows for the gut microbiota to influence animal behavior is evident in experiments examining anxiety-like behaviors.8 GF mice have decreased anxiety-like behavior as demonstrated by standardized testing including increased exploratory behavior in open field tests, and more time in the light of a light/dark box test than SPF controls.8,9,10 The perinatal period is again critical in forming this response as GF animal behavior could be “normalized” only by colonization of GF mothers with SPF microbiota 30 days prior to mating, while colonization of adult GF mice had no effect.8
Maternal diet and obesity
Both human epidemiologic and animal studies have identified maternal obesity as a risk factor for behavioral and neurodevelopmental abnormalities in offspring11 associated with an altered gut microbial community.12,13 Buffington et al.14 examined the contribution of the gut microbiome to behavioral changes in mice born to mothers fed a high fat diet (MHFD). For 8 weeks prior to mating, female mice were fed an MHFD, while controls received a regular diet (MRD). Offspring from MHFD mothers had fewer social interactions, no preference for social novelty, and impaired sociability compared to MRD controls. These behavioral changes corresponded to anatomic changes in the paraventricular nuclei (PVN) of the hypothalamus, with decreased number of cells producing oxytocin, a neuropeptide previously shown to be important in modulating social behavior.15 Offspring exposed to MHFD demonstrated decreased gut microbial diversity and differences in microbial community membership compared to MRD offspring. Among the most significant microbial reduction in MHFD offspring was Lactobacillus reuteri, decreased over ninefold. Colonization with live, but not heat-killed, L. reuteri ameliorated the behavioral change, and restored the number of oxytocin-producing neurons in the PVN to baseline levels. These findings demonstrate that the prenatal environment, in this case MHFD with associated maternal and infant gut microbial changes, influences offspring CNS functioning and behavior.
Prenatal stress
Maternal psychosocial stress is another exposure that can alter fetal programming with behavioral and neurodevelopmental consequences.16,17 Neuroimaging studies of infants born to mothers facing adverse prenatal environments demonstrate reduced cortical gray matter, smaller hippocampal and amygdala volumes, and altered connectivity as early as 5 weeks of age.18,19 The mechanisms underlying the biologic embedding of this exposure are multifactorial, with evidence that the microbiome may be one contributor. Using an established mouse model of chronic variable stress, Jasarevic et al.20 demonstrated that early prenatal stress (PNS) modified maternal vaginal microbial communities, altering diversity as well as composition, with loss of the most abundant vaginal commensal organism Lactobacillus. Loss of maternal Lactobacillus resulted in diminished vertical transmission to the fetus, altering the offspring gut microbiota as well as their plasma metabolome. Changes clustered in pathways related to energy, sugar, and mitochondrial metabolism. Interestingly, there were corresponding sex-specific changes to amino acid profiles in the hypothalamic and limbic regions of the brain, affecting male but not female offspring.
Evidence that stress can have a direct effect on gut microbes dates back 40 years, when investigators identified a decrease in culturable lactobacilli from stools of adult mice moved to cages without water, food, or bedding.21 Similar results have since been obtained using other model systems.22,23,24 Reminiscent of the vaginal microbiota, a number of large and small animal models have demonstrated a relative decrease in the abundance of Lactobacillus from the stool samples of offspring born to mothers subjected to PNS, along with other gut microbial community disruptions.25,26,27 Using rhesus monkeys, Bailey et al.25 demonstrated that infant monkeys born to mothers exposed to PNS had decreased amounts of culturable lactobacilli and bifidobacteria in their stool, most evident with late pregnancy PNS exposure. Rodent models also report alterations in gut microbial membership in PNS-exposed offspring,26,27,28 with decreased amounts of Lactobacillus and Bifidobacterium. Associated behavioral changes were long lasting to adulthood26,28 and sex-specific, with female but not male offspring demonstrating increased anxiety-like behaviors and diminished cognitive function.28 These changes in the gut microbial community were associated with increased levels of interleukin-1β (IL-1β) and decreased brain-derived neurotropic factor (BDNF), a neuronal growth factor suppressed by IL-1β, specifically in the amygdala, an area important in processing emotions and memory.28 Beyond animal models, one small human study of 56 vaginally born infants also found that levels of Lactobacillus was inversely related to stress during pregnancy as measured by a series of self-reported stress questionnaires and high cortisol levels.29 Mothers with high cumulative stress had significantly lower relative abundances of Lactobacillus and Bifidobacterium, with relatively higher levels of Gram-negative Proteobacteria. Taken together, these data indicate that PNS can alter both the maternal and infant microbiota, which is associated with behavioral changes in offspring. While clearly associative and not causal, the associated metabolic changes begin to implicate systemic responses involving metabolism and inflammation as potential mediators.
Blood–brain barrier and short-chain fatty acids
Within the gut, luminal production of short-chain fatty acids (SCFAs) by the gut microbiota is important in maintaining intestinal barrier integrity through regulation of tight junction proteins.30 There is now evidence that SCFAs have similar effects at a more distant site, the blood–brain barrier (BBB).31 Fetuses of GF pregnant mice had significantly increased BBB permeability, related to decreased expression of the tight junction proteins occludin and claudin-5. Fecal transfer from SPF mice to GF adult mice improved the integrity of the BBB and increased levels of occludin and claudin-5. Mono-colonization of GF mice with a single SCFA-producing bacteria, Clostridium tyrobutyricum or Bacteroides thetaiotaomicron, restored BBB integrity to levels similar to that found in SPF mice, as did treatment of GF mice with oral butyrate. These results indicate that metabolic byproducts of the gut microbiota act as signals to influence host physiology distant from their production, at the BBB.
Mechanisms of action
Transcriptional and protein changes
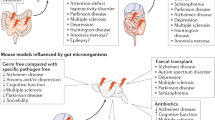
To elucidate the etiology of behavioral changes, comparisons in genetic signaling have been made across various regions of the brain in GF-raised, SPF-raised, and CONV-raised animals. Significant changes in gene expression and protein content are evident across virtually all anatomic components of the brain, from the more primitive structures of the limbic system, the amygdala and hippocampus, to more advanced regions of the frontal and prefrontal cortex, and are linked to changes in behavioral phenotype (Fig. 1). One of the most extensively studied is BDNF, a widely expressed neurotropin important to neuronal growth and survival and implicated in a variety of behavioral changes.32,33 Compared to CONV-raised or SPF mice, GF mice have decreased levels of BNDF mRNA and protein expression.2,8,34 Additionally, upstream regulators N-methyl-d-aspartate receptor subunits2,10 and downstream signaling molecule nerve growth factor-inducible clone A2,8,10,34 are also decreased. Anatomically, the site of these changes is in the hippocampus,2,4,8,10 and the cingulate cortex8 areas of the brain are important in the regulation of these behaviors. Studies examining variances in the brain transcriptome of GF animals compared to SPF and CONV identify significant anatomic site differentials in both neuronal-specific and more generalized cellular functions.8,35
Summary of neurophysiological abnormalities in microbiota-deficient animals. From Voung et al. Annu. Rev Neurosci. 2017.40:21–49 with permission. AM amygdala, BDNF brain-derived neurotrophic factor, CB cerebellum, CRF corticotropin-releasing factor 1, H hippocampus, HPA hypothalamic-pituitary-adrenal, HY hypothalamus, NC nasal cavity, NGFI-A nerve growth factor-inducible protein A, NR2B N-methyl-d-aspartate receptor subtype 2B, OB olfactory bulb, PFC prefrontal cortex, PSD-95 postsynaptic density protein-95, S striatum
Immuno-inflammatory mechanisms
The gut is one of the largest immune organs in the body and a major site of a microbial–host interface. The importance of the gut microbiota to immune development is profound and well established.36,37 Specific commensal bacteria in the gut are involved in inducing both innate and adaptive immune systems. These include T-helper type 1 (Th1), T-helper type 17 (Th17), and regulatory-T (Treg) cells that alter the background inflammatory environment in the gut both locally and systemically. Varieties of systemic autoimmune disorders have been shown to be modulated through altered immune signaling by the gut microbiota.38,39 The role of the gut microbiota in autoimmune CNS pathology was first described in experimental autoimmune encephalomyelitis (EAE).40 EAE is a T cell-mediated model for demyelinating diseases of the CNS, induced by eliciting an immune response to injected myelin-based proteins. During EAE, systemic T cells migrate to the CNS and pro-inflammatory Th1/Th17 responses facilitate demyelination and tissue damage.41 GF mice are highly resistant to development of EAE and are protected via altered T cell responses that have their origin in the gut microbiota. In the absence of bacteria under GF conditions, gut dendritic cells have diminished ability to stimulate local and systemic Th1/Th17 cells to produce pro-inflammatory cytokines. When these cells migrate to the CNS, there is diminished production of interferon-γ and IL-17a, with a concomitant increase in Treg cells that further dampens inflammation. The importance of the gut microbiota to this process is further clarified when GF animals are mono-colonized with segmented filamentous bacteria (SFB), which induce Th17 cells in the gut, and protection is lost. Animals became highly susceptible to EAE induction, and the pro-inflammatory CNS response is again robust. Thus, gut microbiota exert their effects on the CNS by mediating innate dendritic cell responses locally, resulting in altered T cell responses within the CNS.
The use of enteral antibiotics in the EAE animal model had comparable results to those found in GF animals.42 Similar to GF mice, CONV-raised mice pretreated with oral broad-spectrum antibiotics are resistant to development of EAE, with an associated decrease in systemic pro-inflammatory cytokines and increased levels of protective Treg cells. Transfer of Treg’s from animals treated with oral antibiotics conferred protection against EAE. As opposed to the loss of protection with SPF colonization, when the antibiotic-treated mice were re-colonized with a single commensal organism Bacteroides fragilis, resistance was maintained.43 Bacteroides fragilis contains a capsular polysaccharide complex, polysaccharide-A (PSA), which has an established role for immune modulation. Antibiotic-treated mice re-colonized with a mutated form of the organism deficient in capsular PSA lost resistance to EAE. These results implicate specific signaling components within microbes on the CNS pathology via immunomodulating pathways distal to the site of colonization.
Gut microbiota immunomodulation, fetal exposures, and social impairment
More pertinent to pediatrics than EAE, exposure of the fetus to maternal inflammation is a well-described risk factor for subsequent neuropathology,44,45 including ASD.46,47 A potential role of both the maternal and offspring gut microbiome in this process has been recently described using the maternal immune activation (MIA) animal model. In this model, pregnant mice are exposed to polyinoinic:polycytidylic (poly(I:C), an immunostimulant structurally similar to double-stranded RNA, mimicking viral infection. Offspring exhibit ASD-like behavior including reduced sociability, increased anxiety, repetitive stereotypical behaviors, and altered communication patterns, as well as abnormalities in cortical development. Using genetic mutants and blocking antibodies in mice, investigators identified that maternal IL-17a, secreted by Th17 cells, was required for the behavioral and cortical abnormalities to develop in the offspring.48 Based on the importance of gut microbiota to systemic immune responses, investigators next examined the contribution of maternal gut bacteria to the behavioral phenotype by pretreatment of poly(I:C)-exposed mothers with oral vancomycin.49 Pretreatment with this antibiotic decreased the proportion of Th17 cells in the small bowel, diminished maternal serum levels of IL-17a, and protected offspring from both the behavioral phenotype and cortical abnormalities. SFB, known to induce Th17 cells, was identified as the specific vancomycin-sensitive organism required for the phenotype to occur. Offspring of dams depleted of SFB during poly(I:C) exposure did not develop the behavioral changes or cortical pathology. Reconstitution of these dams with SFB through direct gavage or by co-housing with SFB-colonized animals once again resulted in offspring with the ASD phenotype. These data demonstrate that in this MIA model during pregnancy, maternal gut microbiota influences neurodevelopment in offspring through immune-mediated mechanisms involving Th17 induction.
In addition to maternal influences in MIA, offspring gut functioning and microbiota are also affected.50 MIA offspring have increased intestinal permeability related to altered gene expression of tight junction components, along with changes in gut microbial composition involving bacterial classes Clostridia and Bacteroidia. Both these changes are reminiscent of findings reported in limited subgroups of individuals with ASD.51,52,53,54 The altered microbial environment and leaky gut induced by MIA resulted in altered serum metabolites. The most significant increase was in 4-ethylphenylsulfate (4EPS), a predicted metabolite of gut microbes, which was increased 46-fold. Remarkably, systemic administration of wild-type mice with 4EPS alone resulted in behaviors similar to MIA-induced mice. Treatment with the commensal B. fragilis improved gut barrier integrity, restored the microbiota changes, and normalized cytokine and metabolic profiles, abrogating the increase in 4EPS profiles. Behaviors related to anxiety, stereotypical patterns, and vocalizations in response to social interactions were improved, while deficits in social performance were unchanged. In this case, improvement was not a function of microbial capsular PSA, as treatment with mutant bacteria had the same effect. The effect was also not specific to B. fragilis, as treatment with B. thetaiotaomicron also improved behaviors. It was however not a generalized microbial response, as Enterococcus faecalis had no effect. Bacteroides fragilis did not exert its effect directly by permanently colonizing the gut, as it could not be retrieved from feces, but rather correcting microbial community membership of the bacterial classes Clostridia and Bacteroidia. These results demonstrate an effect of the gut microbiome on behavior in this MIA model system, alterable by a common commensal organism, mediated at least in part by specific microbial metabolite.
Microbial metabolites, SCFAs
As illustrated in the MIA model, gut microbes produce a variety of metabolites that can act locally or enter host circulation to affect a variety of physiologic parameters.55 They are implicated in a variety of clinical disorders including inflammatory bowel disease,56 diabetes, atherosclerosis,57 and asthma,58 among others.59 Among the most intensively studied gut microbial metabolites are SCFAs. SCFAs are the fermentation products of undigested carbohydrates by the gut microbiota with the three major SCFAs produced being acetate, propionate, and butyrate. They exert an impact on the immune system by signalling through a variety of G-protein-coupled receptors, including free fatty acid receptors (FFARs) 2 and 3, that promote local intestinal dendritic cells, as well as T cell expansion, differentiation, and function.60
Microglia and SCFA
SCFAs produced by the gut microbiota have recently been shown to influence the CNS immune system by regulating micoglia maturation and function. Microglia are the principal immune cells of the CNS, playing an important role in CNS homeostasis by detecting pathogens and scavenging cellular debris from injured or dead tissue, similar to peripheral macrophages. Using three-dimensional imaging, Erny et al.61 demonstrated altered architecture and density of microglia in GF compared to SPF mice, indicating an immature or dysfunctional state. Comparison of genome-wide mRNA expression profiles between GF and SPF animals identified nearly 400 altered transcripts involving host defense, cell activation, and transcription. Transcripts normally present in immature microglia, including colony-stimulating factor 1 receptor (Csf1r), a cell surface marker with strong developmental regulation, were increased in GF microglia. The immune response of GF microglia when challenged with bacterial or viral stimulants was also immature, with significantly reduced expression of cytokine and chemokine pathways, as well as expression of genes directing cell differentiation, activation, and transformation. Treatment of SPF mice with oral broad-spectrum antibiotic resulted in microglial cell morphology and maturation markers similar to the immature phenotype of GF mice. Targeted re-colonization with specific organisms did not reverse the phenotype; rather, it required the presence of a complex and diverse gut microbial community, including SCFA producers. Remarkably, treatment of GF mice with a mixture of SCFA in the drinking water normalized microglial morphology, density, and maturity based on Csf1r expression. Additionally, mice deficient in the short-chain FFAR2 demonstrated morphologic changes in microglia similar to GF animals, implicating SCFA produced by gut microbes as a key mediator of microglia activation, maturation, and function.
Activation of microglia by SCFAs is also evident in a mouse model of Parkinson’s disease (PD) that overexpress human α-synuclein (ASO), a major constituent of Lewy bodies that are a hallmark of PD neuropathology.62 ASO mice harboring a standard SPF microbiota (SPF-ASO) have motor deficits and aggregation of ASO protein in the caudoputamen and substantia nigra, regions of the brain impacted by PD. Contributing to the phenotypes are activated microglia, producing neurotoxic cytokines. Removal of gut microbiota, through generation of GF animals or antibiotic treatment, protected against the PD phenotype. Feeding GF-ASO mice a mixture of SCFA resulted in loss of protection from the PD phenotype. SCFA-activated microglia, and motor function was again impaired. “Humanization” of standard GF animals (non-ASO) with fecal transplants from PD patients resulted in motor deficits not evident in GF animals transplanted with microbiota healthy human controls. The fecal samples from GF animals colonized with PD donor samples had higher relative abundance of propionate and butyrate, with lower amounts of acetate than that from animals colonized with healthy control sample. Taken together, these results provide compelling evidence that metabolic byproducts of the gut microbiota contribute to microglial homeostasis.
Neuroendocrine signaling
The work by Sudo et al.2 cited above illustrates interactions between gut microbes and endocrine signaling mechanisms in response to stress. Microbial–endocrine communication is also evident in work using a nonobese diabetic (NOD) mouse model where early microbial exposures contributed to sex hormone levels, and modified disease progression.63 Male NOD mice were colonized with a gender-dependent gut microbial community that elevated testosterone levels, and were protected from diabetes compared to GF and female mice. Fecal transfer from adult NOD male mice to female weanling, but not adult, mice altered the gut microbiota, elevated testosterone levels, and conferred protection.
Additional signaling mechanisms
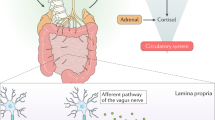
The vagus nerve, which is composed of both afferent and efferent fibers, offers an additional mechanism for bidirectional communication. Afferent fibers are stimulated directly or indirectly by microbial components or metabolites with local or central effects, and efferent fibers alter gut microbial community structure by affecting gut inflammation and permeability.64 Microbiota are also capable of producing neurotransmitters directly or by stimulating production by neurotransmitter-producing cells like enterochromaffin cells. The metabolite 5-hydroxytryptamine (5-HT) and its precursor the amino acid tryptophan are of particular interest as 5-HT has been important in the regulation of gut motility,65 and tryptophan can cross the BBB and participate in 5-HT metabolism centrally. Finally, host genotype also influences colonization patterns in human microbiomes (see for a review Goodrichet al.66). How genotype affects colonization patterns following delivery, and what if any relationship it has to genetic differences in neurologic disorders remains to be determined.
Human studies
The contribution of the gut microbiota to neurodevelopment and behavior in humans is a complex arena with many factors influencing outcomes including genetic, epigenetic, and environmental components, and studies of gut microbial contribution must be taken in this context. Human studies examining a contribution from the gut microbiome have utilized interventions including prebiotics and probiotics,67,68,69 fecal transplantation,70 and antibiotics71 to examine effects on both mood and emotions. There has been intense interest in examining individuals with ASD partly due to the frequent gastrointestinal symptomology associated with the disorder,72 and immune alterations (see for a review Vuong and Hsiao73). Studies have compared stools74,75,76,77,78 or intestinal biopsy samples79 between individuals with ASD to those without, across a variety of ages. While microbial dysbiosis is evident, no consistent microbial signature either in terms of specific taxa74,75,76,77,80 or metabolic byproducts78,81 has emerged across studies. Additionally, the cross-sectional design of many of these studies limits the ability to differentiate cause and effect. This is particularly problematic if looking to examine early origins of disease or behavior that may not be manifest until later in life.
Two small interventional studies endorse the concept that the gut microbiome may contribute to disease severity. In one, 11 children treated with 8 weeks of oral vancomycin demonstrated improved symptoms based on standardized blinded testing, which then reverted to baseline after treatment was stopped on follow-up in 2 to 8 months.71 In the second,70 18 children underwent microbial transplant therapy (MTT), consisting of an initial multi-staged bowel cleansing followed by daily administration of a standardized human gut microbiota used in recurrent Clostridium difficile infections,82 for 7–8 weeks. Behavioral ASD symptomatology significantly improved, with a sustained effect on follow-up at 8 weeks following completion of treatment. Longitudinal analysis of stool samples demonstrated a sustained increase in stool sample bacterial diversity following MTT, with evidence for continued engraftment of a number of taxa including Bifidobacterium, frequently reported as diminished in ASD population.80,81 The contribution of the viral elements, frequently overlooked, was also shifted toward communities closer to that of fecal donors.
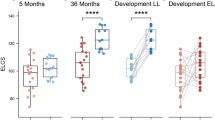
The first prospective human study to examine the relationship between the developing infant gut microbiome and cognition was recently reported.83 Fecal samples from 89 typically developing 1-year-old toddlers were analyzed for microbial community structure and tested for an association to cognitive functions. Infants clustered into three distinct gut microbial patterns, driven by differences in the relative abundance of Faeccalibacterium, Bacteroides, and an unclassified, Ruminococcaceae, reminiscent of previously described adult enterotypes.84 At 2 years, cognitive testing results differed significantly between clusters as measured by a composite score of cognitive function (the Mullen Scales of Early Learning). The Bacteroides cluster scored highest (90th percentile) and Faecalibacterium lowest, although still within normal range (72nd percentile). Receptive and expressive language domains showed the greatest differences, with no significant differences measured in motor (gross or fine) or visual reception skills, and no differences on neuroimaging between brain regions. Alpha diversity, the amount of within-sample microbial variation, was inversely related to cognitive function with the Bacteroides clusters demonstrating lowest alpha diversity. Breastfeeding is an important covariate associated with cognitive outcomes,85 and results in a gut microbiota with decreased diversity and Bacteroides dominant. In multivariate analysis that accounted for feeding type as well as other important clinical and demographic co-variates, cluster group, but not alpha diversity, remained significantly related to cognitive outcome at 2 years. While the results of this study remain associative and not causative, it is an important step in incorporating human subjects in examining the relationship between the gut microbiota and human neurodevelopment.
Perspective
Our understanding of the interactions between our gut microbes and the developing nervous system is in early stages (Fig. 2). Animal models have provided strong evidence that gut bacteria and their metabolites play a role in CNS homeostasis affecting behavior.86 Human studies have been primarily correlative, and caution must be applied before concluding cause and effect. Significant challenges remain in understanding what, if any, long-lasting effects early host–microbe interactions have on human neurodevelopmental and behavioral outcomes. Many of these challenges are rooted in our limited understanding of how early microbial colonization patterns interact with simultaneously evolving immune, neuroendocrine, and nervous systems. Questions remain regarding the importance of initial pioneering microbes, including viruses, fungi, and archaea during this co-evolution. Maternal and infant diet, stress, mode of delivery, intrapartum infection, and antibiotic exposures all shape early microbial colonization patterns. How long those patterns last, and what if any long-term phonotypic impact they have in humans disease and neurodevelopment is just now being examined.37,87 Human studies with a pediatric focus and systems-based approach, moving between animals and humans, will be required to understand the mechanism of host–microbe interactions and the impact on human health and development.
References
Backhed, F., Manchester, J. K., Semenkovich, C. F. & Gordon, J. I. Mechanisms underlying the resistance to diet-induced obesity in germ-free mice. Proc. Natl Acad. Sci. USA 104, 979–984 (2007).
Sudo, N. et al. Postnatal microbial colonization programs the hypothalamic-pituitary-adrenal system for stress response in mice. J. Physiol. 558, 263–275 (2004).
Vuong, H. E., Yano, J. M., Fung, T. C. & Hsiao, E. Y. The microbiome and host behavior. Annu. Rev. Neurosci. 40, 21–49 (2017).
Bercik, P. et al. The intestinal microbiota affect central levels of brain-derived neurotropic factor and behavior in mice. Gastroenterology 141, 599–609 (2011).
Degroote, S., Hunting, D. J., Baccarelli, A. A. & Takser, L. Maternal gut and fetal brain connection: increased anxiety and reduced social interactions in Wistar rat offspring following peri-conceptional antibiotic exposure. Prog. Neuropsychopharmacol. Biol. Psychiatry 71, 76–82 (2016).
Tochitani, S. et al. Administration of non-absorbable antibiotics to pregnant mice to perturb the maternal gut microbiota is associated with alterations in offspring behavior. PLoS ONE 11, e0138293 (2016).
Phelps, D. et al. Microbial colonization is required for normal neurobehavioral development in zebrafish. Sci. Rep. 7, 11244 (2017).
Diaz Heijtz, R. et al. Normal gut microbiota modulates brain development and behavior. Proc. Natl Acad. Sci. USA 108, 3047–3052 (2011).
Clarke, G. et al. The microbiome–gut–brain axis during early life regulates the hippocampal serotonergic system in a sex-dependent manner. Mol. Psychiatry 18, 666–673 (2013).
Neufeld, K. M., Kang, N., Bienenstock, J. & Foster, J. A. Reduced anxiety-like behavior and central neurochemical change in germ-free mice. Neurogastroenterol. Motil. 23, 255–264, e119 (2011).
van der Burg, J. W. et al. The role of systemic inflammation linking maternal BMI to neurodevelopment in children. Pediatr. Res. 79, 3–12 (2016).
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Buffington, S. A. et al. Microbial reconstitution reverses maternal diet-induced social and synaptic deficits in offspring. Cell 165, 1762–1775 (2016).
Donaldson, Z. R. & Young, L. J. Oxytocin, vasopressin, and the neurogenetics of sociality. Science 322, 900–904 (2008).
Bale, T. L. et al. Early life programming and neurodevelopmental disorders. Biol. Psychiatry 68, 314–319 (2010).
Beijers, R., Buitelaar, J. K. & de Weerth, C. Mechanisms underlying the effects of prenatal psychosocial stress on child outcomes: beyond the HPA axis. Eur. Child Adolesc. Psychiatry 23, 943–956 (2014).
Betancourt, L. M. et al. Effect of socioeconomic status (SES) disparity on neural development in female African‐American infants at age 1 month. Dev. Sci. 19, 947–956 (2015).
Noble, K. G. et al. Family income, parental education and brain structure in children and adolescents. Nat. Neurosci. 18, 773–778 (2015).
Jasarevic, E., Howerton, C. L., Howard, C. D. & Bale, T. L. Alterations in the vaginal microbiome by maternal stress are associated with metabolic reprogramming of the offspring gut and brain. Endocrinology 156, 3265–3276 (2015).
Tannock, G. W. & Savage, D. C. Influences of dietary and environmental stress on microbial populations in the murine gastrointestinal tract. Infect. Immun. 9, 591–598 (1974).
Bailey, M. T. & Coe, C. L. Maternal separation disrupts the integrity of the intestinal microflora in infant rhesus monkeys. Dev. Psychobiol. 35, 146–155 (1999).
Bailey, M. T. et al. Exposure to a social stressor alters the structure of the intestinal microbiota: implications for stressor-induced immunomodulation. Brain Behav. Immun. 25, 397–407 (2011).
Galley, J. D. et al. Exposure to a social stressor disrupts the community structure of the colonic mucosa-associated microbiota. BMC Microbiol. 14, 189 (2014).
Bailey, M. T., Lubach, G. R. & Coe, C. L. Prenatal stress alters bacterial colonization of the gut in infant monkeys. J. Pediatr. Gastroenterol. Nutr. 38, 414–421 (2004).
Golubeva, A. V. et al. Prenatal stress-induced alterations in major physiological systems correlate with gut microbiota composition in adulthood. Psychoneuroendocrinology 60, 58–74 (2015).
Jasarevic, E., Howard, C. D., Misic, A. M., Beiting, D. P. & Bale, T. L. Stress during pregnancy alters temporal and spatial dynamics of the maternal and offspring microbiome in a sex-specific manner. Sci. Rep. 7, 44182 (2017).
Gur, T. L. et al. Prenatal stress affects placental cytokines and neurotrophins, commensal microbes, and anxiety-like behavior in adult female offspring. Brain Behav. Immun. 64, 50–58 (2017).
Zijlmans, M. A., Korpela, K., Riksen-Walraven, J. M., de Vos, W. M. & de Weerth, C. Maternal prenatal stress is associated with the infant intestinal microbiota. Psychoneuroendocrinology 53, 233–245 (2015).
Peng, L., Li, Z. R., Green, R. S., Holzman, I. R. & Lin, J. Butyrate enhances the intestinal barrier by facilitating tight junction assembly via activation of AMP-activated protein kinase in Caco-2 cell monolayers. J. Nutr. 139, 1619–1625 (2009).
Braniste, V. et al. The gut microbiota influences blood–brain barrier permeability in mice. Sci. Transl. Med. 6, 263ra158 (2014).
Govindarajan, A. et al. Transgenic brain-derived neurotrophic factor expression causes both anxiogenic and antidepressant effects. Proc. Natl Acad. Sci. USA 103, 13208–13213 (2006).
Martinowich, K., Manji, H. & Lu, B. New insights into BDNF function in depression and anxiety. Nat. Neurosci. 10, 1089–1093 (2007).
Arentsen, T., Raith, H., Qian, Y., Forssberg, H. & Diaz Heijtz, R. Host microbiota modulates development of social preference in mice. Microb. Ecol. Health Dis. 26, 29719 (2015).
Stilling, R. M. et al. Microbes and neurodevelopment—absence of microbiota during early life increases activity-related transcriptional pathways in the amygdala. Brain Behav. Immun. 50, 209–220 (2015).
Hooper, L. V., Littman, D. R. & Macpherson, A. J. Interactions between the microbiota and the immune system. Science 336, 1268–1273 (2012).
Tamburini, S., Shen, N., Wu, H. C. & Clemente, J. C. The microbiome in early life: implications for health outcomes. Nat. Med. 22, 713–722 (2016).
Abdollahi-Roodsaz, S. et al. Stimulation of TLR2 and TLR4 differentially skews the balance of T cells in a mouse model of arthritis. J. Clin. Invest. 118, 205–216 (2008).
Wen, L. & Duffy, A. Factors influencing the gut microbiota, inflammation, and type 2 diabetes. J. Nutr. 147, 1468S–1475SS (2017).
Lee, Y. K., Menezes, J. S., Umesaki, Y. & Mazmanian, S. K. Proinflammatory T-cell responses to gut microbiota promote experimental autoimmune encephalomyelitis. Proc. Natl Acad. Sci. USA 108(Suppl. 1), 4615–4622 (2011).
Kasper, L. H. & Shoemaker, J. Multiple sclerosis immunology: the healthy immune system vs the MS immune system. Neurology 74(Suppl. 1), S2–S8 (2010).
Ochoa-Reparaz, J. et al. Role of gut commensal microflora in the development of experimental autoimmune encephalomyelitis. J. Immunol. 183, 6041–6050 (2009).
Ochoa-Reparaz, J. et al. Central nervous system demyelinating disease protection by the human commensal Bacteroides fragilis depends on polysaccharide A expression. J. Immunol. 185, 4101–4108 (2010).
Armstrong-Wells, J. et al. Inflammatory predictors of neurologic disability after preterm premature rupture of membranes. Am. J. Obstet. Gynecol. 212, 212 e1–212 e9 (2015).
Rovira, N. et al. Impact of histological chorioamnionitis, funisitis and clinical chorioamnionitis on neurodevelopmental outcome of preterm infants. Early Hum. Dev. 87, 253–257 (2011).
Atladottir, H. O. et al. Maternal infection requiring hospitalization during pregnancy and autism spectrum disorders. J. Autism Dev. Disord. 40, 1423–1430 (2010).
Atladottir, H. O., Henriksen, T. B., Schendel, D. E. & Parner, E. T. Autism after infection, febrile episodes, and antibiotic use during pregnancy: an exploratory study. Pediatrics 130, e1447–e1454 (2012).
Choi, G. B. et al. The maternal interleukin-17a pathway in mice promotes autism-like phenotypes in offspring. Science 351, 933–939 (2016).
Kim, S. et al. Maternal gut bacteria promote neurodevelopmental abnormalities in mouse offspring. Nature 549, 528–532 (2017).
Hsiao, E. Y. et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 155, 1451–1463 (2013).
D’Eufemia, P. et al. Abnormal intestinal permeability in children with autism. Acta Paediatr. 85, 1076–1079 (1996).
de Magistris, L. et al. Alterations of the intestinal barrier in patients with autism spectrum disorders and in their first-degree relatives. J. Pediatr. Gastroenterol. Nutr. 51, 418–424 (2010).
Finegold, S. M., Downes, J. & Summanen, P. H. Microbiology of regressive autism. Anaerobe 18, 260–262 (2012).
Esnafoglu, E. et al. Increased serum zonulin levels as an intestinal permeability marker in autistic subjects. J. Pediatr. 188, 240–244 (2017).
Nicholson, J. K. et al. Host–gut microbiota metabolic interactions. Science 336, 1262–1267 (2012).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Koeth, R. A. et al. Intestinal microbiota metabolism of l-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat. Med. 19, 576–585 (2013).
Trompette, A. et al. Gut microbiota metabolism of dietary fiber influences allergic airway disease and hematopoiesis. Nat. Med. 20, 159–166 (2014).
Rooks, M. G. & Garrett, W. S. Gut microbiota, metabolites and host immunity. Nat. Rev. Immunol. 16, 341–352 (2016).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Backhed, F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016).
Erny, D. et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat. Neurosci. 18, 965–977 (2015).
Sampson, T. R. et al. Gut microbiota regulate motor deficits and neuroinflammation in a model of Parkinson’s disease. Cell 167, 1469–80 e12 (2016).
Markle, J. G. et al. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 339, 1084–1088 (2013).
Bonaz, B., Bazin, T. & Pellissier, S. The vagus nerve at the interface of the microbiota–gut–brain axis. Front. Neurosci. 12, 49 (2018).
Yano, J. M. et al. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell 161, 264–276 (2015).
Goodrich, J. K., Davenport, E. R., Clark, A. G. & Ley, R. E. The relationship between the human genome and microbiome comes into view. Annu. Rev. Genet. 51, 413–433 (2017).
Messaoudi, M. et al. Assessment of psychotropic-like properties of a probiotic formulation (Lactobacillus helveticus R0052 and Bifidobacterium longum R0175) in rats and human subjects. Br. J. Nutr. 105, 755–764 (2011).
Tillisch, K. et al. Consumption of fermented milk product with probiotic modulates brain activity. Gastroenterology 144, 1394–1401 (2013).
Steenbergen, L., Sellaro, R., van Hemert, S., Bosch, J. A. & Colzato, L. S. A randomized controlled trial to test the effect of multispecies probiotics on cognitive reactivity to sad mood. Brain Behav. Immun. 48, 258–264 (2015).
Kang, D. W. et al. Microbiota transfer therapy alters gut ecosystem and improves gastrointestinal and autism symptoms: an open-label study. Microbiome 5, 10 (2017).
Sandler, R. H. et al. Short-term benefit from oral vancomycin treatment of regressive-onset autism. J. Child Neurol. 15, 429–435 (2000).
Kohane, I. S. et al. The co-morbidity burden of children and young adults with autism spectrum disorders. PLoS ONE 7, e33224 (2012).
Vuong, H. E. & Hsiao, E. Y. Emerging roles for the gut microbiome in autism spectrum disorder. Biol. Psychiatry 81, 411–423 (2017).
Finegold, S. M. et al. Gastrointestinal microflora studies in late-onset autism. Clin. Infect. Dis. 35, S6–S16 (2002).
Song, Y., Liu, C. & Finegold, S. M. Real-time PCR quantitation of clostridia in feces of autistic children. Appl. Environ. Microbiol. 70, 6459–6465 (2004).
Parracho, H. M., Bingham, M. O., Gibson, G. R. & McCartney, A. L. Differences between the gut microflora of children with autistic spectrum disorders and that of healthy children. J. Med. Microbiol. 54, 987–991 (2005).
Finegold, S. M. et al. Pyrosequencing study of fecal microflora of autistic and control children. Anaerobe 16, 444–453 (2010).
Wang, L. et al. Elevated fecal short chain fatty acid and ammonia concentrations in children with autism spectrum disorder. Dig. Dis. Sci. 57, 2096–2102 (2012).
Williams, B. L. et al. Impaired carbohydrate digestion and transport and mucosal dysbiosis in the intestines of children with autism and gastrointestinal disturbances. PLoS ONE 6, e24585 (2011).
Wang, L. et al. Low relative abundances of the mucolytic bacterium Akkermansia muciniphila and Bifidobacterium spp. in feces of children with autism. Appl. Environ. Microbiol. 77, 6718–6721 (2011).
Adams, J. B., Johansen, L. J., Powell, L. D., Quig, D. & Rubin, R. A. Gastrointestinal flora and gastrointestinal status in children with autism—comparisons to typical children and correlation with autism severity. BMC Gastroenterol. 11, 22 (2011).
Hamilton, M. J., Weingarden, A. R., Sadowsky, M. J. & Khoruts, A. Standardized frozen preparation for transplantation of fecal microbiota for recurrent Clostridium difficile infection. Am. J. Gastroenterol. 107, 761–767 (2012).
Carlson, A. L. et al. Infant gut microbiome associated with cognitive development. Biol. Psychiatry 83, 148–159 (2018).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Girard, L. C., Doyle, O. & Tremblay, R. E. Breastfeeding, cognitive and noncognitive development in early childhood: a population study. Pediatrics 139, pii: e20161848, (2017).
Lim, E. S. et al. Early life dynamics of the human gut virome and bacterial microbiome in infants. Nat. Med. 21, 1228–1234 (2015).
Chu, D. M. et al. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat. Med. 23, 314–326 (2017).
Humann, J. et al. Bacterial peptidoglycan traverses the placenta to induce fetal neuroproliferation and aberrant postnatal behavior. Cell Host Microbe 19, 901 (2016).
Acknowledgments
Salary support for B.B.W. was provided in part by a grant from Children’s Discovery Institute at Washington University in St. Louis MD-II-2018-725.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Warner, B.B. The contribution of the gut microbiome to neurodevelopment and neuropsychiatric disorders. Pediatr Res 85, 216–224 (2019). https://doi.org/10.1038/s41390-018-0191-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-018-0191-9
This article is cited by
-
Establishing associated risk factors, including fungal and parasitic infections among Malaysians living with schizophrenia
Scientific Reports (2024)
-
Association between microbiome and the development of adverse posttraumatic neuropsychiatric sequelae after traumatic stress exposure
Translational Psychiatry (2023)
-
A prospective investigation into the association between the gut microbiome composition and cognitive performance among healthy young adults
Gut Pathogens (2022)
-
Sex-specific relationships of the infant microbiome and early-childhood behavioral outcomes
Pediatric Research (2022)
-
Depression phenotype identified by using single nucleotide exact amplicon sequence variants of the human gut microbiome
Molecular Psychiatry (2021)