Abstract
Background
The role of epithelial cells in eosinophilic esophagitis (EoE) is not well understood. In this study, our aim was to isolate, culture, and expand esophageal epithelial cells obtained from patients with or without EoE and characterize differences observed over time in culture.
Methods
Biopsies were obtained at the time of endoscopy from children with EoE or suspected to have EoE. We established patient-derived esophageal epithelial cell (PDEEC) lines utilizing conditional reprogramming methods. We determined integrin profiles, gene expression, MHC class II expression, and reactivity to antigen stimulation.
Results
The PDEECs were found to maintain their phenotype over several passages. There were differences in integrin profiles and gene expression levels in EoE-Active compared to normal controls and EoE-Remission patients. Once stimulated with antigens, PDEECs express MHC class II molecules on their surface, and when co-cultured with autologous T-cells, there is increased IL-6 and TNF-α secretion in EoE-Active patients vs. controls.
Conclusion
We are able to isolate, culture, and expand esophageal epithelial cells from pediatric patients with and without EoE. Once stimulated with antigens, these cells express MHC class II molecules and behave as non-professional antigen-presenting cells. This method will help us in developing an ex vivo, individualized, patient-specific model for diagnostic testing for causative antigens.
Similar content being viewed by others
Introduction
Eosinophilic esophagitis (EoE) is an allergic, immune-mediated, clinicopathologic entity that is challenging to treat in pediatric and adult patients. It is believed that EoE is mainly driven by a T helper type 2 (Th2) inflammatory response leading to esophageal eosinophilia.1,2 In our previous work,3 using esophageal biopsy culture supernatants, we identified a set of cytokines and chemokines that differentiated EoE-New/-Active from EoE-Remission and non-inflammatory controls. This set of analytes included epithelial and innate cytokines and chemokines, suggesting that esophageal epithelial cells and the innate immune system play a significant role in the pathogenesis, continuation, and persistence of inflammation in EoE. Interleukin (IL)-6 and tumor necrosis factor (TNF)-α were two of the most highly secreted cytokines in EoE patients compared to non-inflammatory controls.
Human epithelial cells have been isolated successfully from a variety of tissues in the human body that have been both diseased and non-diseased including: lung,4 breast,5 small intestine,6 colon,7 bladder,8 and esophagus.9 It is important to note that human primary cells have a finite number of passages in which the phenotype is maintained.10, Conditional reprogramming of normal and cancerous epithelial cells (epithelial reprogramming) utilizing feeder cells and ROCK inhibitor was described in 2012 by Liu et al.11 This technique is capable of expanding large numbers of epithelial cells indefinitely by “reprogramming” these cells to a stem- or progenitor-like state, which is highly proliferative. Therefore, this technique offers a large reservoir of patient-specific epithelial cells for disease investigation or tissue engineering without the need for immortalization. Lastly, it has been shown that epithelial cells from various tissues including nasal mucosa, airway mucosa, and esophageal mucosa may express major histocompatibility complex (MHC) class II molecules during inflammation and thus behave as non-professional antigen-presenting cells (NPAPCs).12,13 Their capability of taking on the role of NPAPCs allows us to investigate the mechanisms behind the initiation of EoE in vitro.
In EoE, it has been shown that there is an abnormal epithelial barrier with reduced expression of E-cadherin, claudin, occudin, desmoglein-1, and flaggrin.14 Integrins are transmembrane proteins that play a vital role in bidirectional cell–cell signaling and function. Integrins usually exist in herterodimers and there are 24 known integrin dimers in humans.15,16 Cell adhesion and motility are regulated by crosstalk between cadherin- and integrin-mediated cell–cell and cell–matrix adhesions.17,18 Integrin profiles in EoE have never been evaluated or investigated.
Lastly, gene expression, genome-wide association studies, and RNA sequencing studies have identified that IL-5,19 IL-13,20,21 IL-18,22,19 IL-33, Eotaxin-3,23,24 thymic stromal lymphopoietin (TSLP),25 and ALOX1527 are upregulated while CRISP 3 and SPINK 726 are downregulated in EoE compared to non-inflammatory controls and inactive EoE. We used these genes as our basis for our gene expression studies of biopsies and expanded cells.
Our goal in this study was to isolate, culture, expand, and characterize individualized patient-derived esophageal epithelial cells (PDEECs) obtained from children with EoE. We characterized the cells by looking at their morphology, measuring growth kinetics, and by determining integrin profiles and gene expression levels. We also determined their functionality by evaluating their expression of MHC class II molecules before and after exposure to antigens and measured cytokine response to antigens in the presence or absence of autologous T-cells. Our overarching hypothesis is that these cells maintain their characteristics, phenotype, and functionality thus allowing us to investigate the role of epithelial cells in EoE for future experiments.
Materials and methods
Patients and patient groups
We recruited and enrolled children between the ages of 4–17 years at the time of a medically indicated esophagogastroduodenoscopy (EGD) at Connecticut Children’s Medical Center (CCMC), Hartford, CT between 11/16/2016 and 12/1/2017. This study was reviewed and approved by the institutional review board (CCMC IRB# 16-043).
A diagnosis of EoE was confirmed, according to the 2011 EoE consensus statement27 and was based on clinical history, endoscopic findings, and histologic findings (≥15 eosinophils per high-powered field [eos/HPF], basal layer hyperplasia, and papillary elongation). We defined three study groups as follows: (1) EoE-New/-Active (n = 14): untreated/newly diagnosed or treated/known EoE who underwent endoscopy after treatment for 8–12 weeks and were found to have persistence of esophageal eosinophilia (≥15 eos/HPF); (2) EoE-Remission: known EoE, inflammation resolved after treatment with dairy-free diet (<15 eos/HPF) (n = 6); and (3) Controls: patients who underwent EGD for suspected EoE and who were found to have no visual (endoscopic) or histologic evidence of esophageal inflammation (<1 eos/HPF) (n = 8). We excluded children with other comorbidities.
Sample collection
During the endoscopy procedure, five biopsies were collected from each patient. Four biopsies (for cell isolation and expansion) were placed in a tube with keratinocyte serum-free medium (Life Technologies, Carlsbad, CA) with 1× primocin (Invivogen, San Diego, CA), placed on ice in a Styrofoam box, and transported to our laboratory at the University of Connecticut Health Center. A single biopsy was placed in RNA later (Qiagen, Valencia, CA) for gene expression studies. We collected 10 ml of blood from each patient at the time of intravenous line insertion before their endoscopy procedure (before anesthesia was given).
Isolation and expansion of human esophageal epithelial cells
The four biopsies transported to our laboratory were centrifuged and resuspended in 10 ml of 1 U/ml dispase solution and incubated for 15 min at 37 °C. The biopsies were centrifuged and resuspended in 5 ml of 0.05% Trypsin-EDTA (Life Technologies, Carlsbad, CA) and minced on a sterile 100 mm plate using 2 sterilized razor blades. The minced tissue and an additional 5 ml of trypsin were placed in a tube and incubated for 10 min at 37 °C. We added 3 ml of fetal bovine serum (FBS) (Denville Scientific, Holliston, MA) to neutralize the reaction and cells were then centrifuged and resuspended in epithelial conditional reprogramming medium.28 Cells were counted and plated on a 100 mm pre-coated with 0.1% gelatin plate along with 1.1 × 104 3T3 cells/cm2 (irradiated at 3000 rad), 10 ml of epithelial conditional reprogramming medium, and 10 μl of 10 mM ROCK inhibitor (Tocris, Bristol, UK).
After cells became confluent, they were expanded out until passage 3. They were washed with phosphate-buffered saline (PBS) and 0.05% Trypsin-EDTA (Life Technologies, Carlsbad, CA) was added for 2–3 min at 37 °C to remove the 3T3 cells. After aspirating the Trypsin-EDTA, 0.25% Trypsin-EDTA (Life Technologies, Carlsbad, CA) was added and incubated for 5–10 min at 37 °C to lift the epithelial cells from the plate. Equal amount of medium was added to neutralize the reaction. Cells were centrifuged, resuspended, and then counted on a hemocytometer. Approximately, 1 million cells were plated onto a new pre-coated 0.1% gelatin plate along with irradiated 3T3 cells, epithelial reprogramming medium, and ROCK inhibitor. The rest of the cells were frozen with 10% dimethyl sulfoxide (DMSO) and epithelial conditional reprogramming medium and stored in liquid nitrogen. 3T3 cells were used at every passage whenever PDEECs were plated. 3T3 cells were removed when PDEECs were collected to be stored at −80 °C or for testing.
Immunofluorescence staining
Cells from passages 1 and 3 were fixed with 4% paraformaldehyde (PFA) solution29 for 10 min and washed with PBS. Cells were blocked with staining solution containing 1× PBS, 0.1% Triton X-100 (Fisher Scientific, Hampton, NH), 0.1% Tween 20 (Sigma Aldrich, St. Louis, MO), and 5% FBS for 45 min. After blocking, primary antibodies ZO-1 (BD Bioscience, San Jose, CA, 610966), E-cadherin (Santa Cruz, Dallas, TX, SC-7870), Ki67 (BD Bioscience, San Jose, CA, 556003), and P63 (Santa Cruz, Dallas, TX, SC-8343) were diluted at 1:100 with the staining solution and incubated overnight at 4 °C. Cells were rinsed with PBS, stained with secondary antibody, diluted at 1:1000 with staining buffer, and incubated for 1 h at room temperature in the dark. Cells were again rinsed and stained with 4,6-diamidino-2-phenylindole, diluted at 1:100 with 1× PBS, for 10 min at room temperature in the dark. Cells were rinsed with PBS and imaged using fluorescence microscope.
Growth kinetics
Four six-well cell culture dishes were coated with 0.1% gelatin for over an hour at 37 °C. Approximately 1.25 × 104 patient cells and 7.5 × 103 irradiated 3T3 feeder cells were plated into 3 triplicate wells with 2 ml of epithelial conditional reprogramming medium and 2 μl of ROCK inhibitor. At time intervals of 24, 48, 72, and 96 h, 1 plate was rinsed with PBS (Fisher Scientific, Hampton, NH) and incubated for 3 min with 1 ml 0.05% Trypsin-EDTA. It was then incubated for 5–8 min with 1 ml 0.25% Trypsin-EDTA until nearly all the cells had lifted off the plate. Cells were centrifuged at 300 × g for 5 min. The cells were then resuspended and counted using a hemocytometer.
Flow cytometric staining for integrins
Cells from 14 EoE-Active, 6 EoE-Remission, and 5 non-inflammatory control patients were collected at the time of expanding to passages 1 and 3. Approximately 500,000 cells were taken and aliquoted into 2 tubes for experimental staining, 1 tube for unstained cells, and 1 tube for live/dead staining. The cells (minus the unstained cells) were resuspended in 1 ml PBS and 1 μl of Fixable Viability Dye e780 (eBiosciences, San Diego, CA, 65-0865-14) (10% stock e780 and PBS, determined in previous study) and incubated on ice for 30 min in the dark. Cells were centrifuged at 2200 RPM for 4 min. All cells were resuspended in 500 μl 4% PFA and incubated for 7 min on ice in the dark. Fluorescence-activated cell sorting (FACS) buffer (PBS+2% FBS) was added and cells were centrifuged at 2200 RPM for 4 min. FACS buffer was added to the experimental samples and stained for surface markers including: Alpha 2 (CD49b, BD Biosciences, San Jose, CA, 564119, 5 μl), Beta 1 (CD29, BD Biosciences, San Jose, CA, 563513, 5 μl), Beta 3 (CD61, eBiosciences, San Jose, CA, 11-0519-42, 5 μl), Alpha 5 (CD49e, eBiosciences, San Jose, CA, 12-4900-42, 5 μl), Beta 2 (CD104, eBiosciences, San Jose, CA, 50-1049-82, 5 μl), Alpha 1 (CD49a, eBiosciences, San Jose, CA, 46-9490-42, 5 μl), Alpha 6 (CD49f eBiosciences, San Jose, CA, 25-0495-82, 2.5 μl), E-cadherin (CD324, eBiosciences, San Jose, CA, 53-3249-82, 4 μl), EpCAM/CD326 (eBiosciences, San Jose, CA, 12-9326-42, 5 μl), vimentin (Biolegend, San Diego, CA, 677807), Ki67 (eBiosciences, San Jose, CA, 48-5699-42), and CD90 (eBiosciences, San Jose, CA, 17-0909-42), for 30 min. FACS buffer was added and cells were all centrifuged at 2200 RPM for 4 min. Cells were resuspended in 300–500 μl FACS buffer, filtered in a 35 µm flow cytometric tube, and run on an LSRII flow cytometric machine.
Quantitative real-time PCR
Biopsies and PDEECs from five non-inflammatory controls and nine EoE-Active patients were analyzed. The original biopsy and approximately 500,000 cells from passage 3 were frozen in 500 μl RLT buffer until processing. Using an RNEasy Qiagen Kit (Qiagen Valencia, CA), RNA was isolated and quantified using a NanoDrop spectrophotometer. cDNA was generated using iScript (Bio-Rad Laboratories) protocol for final concentration of 5 ng per PCR reaction. Quantitative PCR was performed using premade primers (ThermoFisher Scientific, Waltham, MA) for genes: CDH1 (Hs0123894), P63 (Hs00978340), TSLP (Hs00263639), EpCAM (Hs00158980), Crisp3 (Hs00195988), CCL26 (Hs00171146), Spink7 (Hs00261445), IL-18 (Hs01038788), IL-33 (Hs00369211), TJP1 (Hs01551861), IL-13 (Hs00174379), Alox15 (Hs00993765), IL-5 (hs01548712), IL-6 (Hs00985639), IL-10 (Hs00961622), and Actin-Beta (Hs01060665). Cq values were normalized to β-actin for comparison of biopsies and passage 3 (Δcq). The fold change (ΔΔcq) between P3 and biopsy was then calculated.
CD4+/CD8+ separation
Patient blood was collected using BD Vacutainer CPTs Mononuclear Cell Preparation tube (BD, Franklin Lakes, NJ) and centrifuged. The plasma layer was pipetted off. The peripheral blood mononuclear cell layer was pipetted off and centrifuged with RPMI medium, FBS, and Penstrep. The cells were resuspended with RPMI medium, FBS, and DMSO. Finally, the cells were frozen and stored in liquid nitrogen until ready to use.30
Using the CD4+ Microbead MACS Separation (Miltenyi Biotec, Aubrun, CA) protocol, mononuclear cells were thawed, centrifuged, and resuspended with a buffer solution as recommended by the manufacturer’s instructions (PBS, 0.5% BSA, 2 mM EDTA). Cells were incubated with CD4+ microbeads for 15 min at 4 °C, centrifuged, and resuspended with new buffer solution. The cell suspension was applied onto a MACS MS column and unlabeled cells that passed through were collected. The column was rinsed three times with buffer solution and the effluent of the unlabeled cells was collected. The column was taken off the magnetic stand and CD4+ cells were carefully eluted with buffer solution and the provided plunger. The unlabeled cells were collected and then used for CD8+ isolation using the CD8+ Microbeads and a new MACS MS separation column.
MHC profiles and stimulation assay
Approximately, 250,000 PDEECs were collected at passages 1 and 3 from 5 EoE-Active, 5 EoE-Remission, and 5 non-inflammatory control patients. The cells were stained for flow cytometry as previously done except anti-human surface markers HLA-DR (BD Biosciences, San Jose, CA, 555558) and HLA-ABC (eBioscience, San Jose, CA 12-9983-42) were used. After staining, they were run on an LSRII flow cytometry machine for detection.
In a stimulation assay, from the same patients, 250,000 cells from passage 2, were stimulated in separate wells with dust mite antigen (Dermatophagoides farniae, Dermatophagoides pteronyssinus) (Greer Labs, Lenoir, NC), milk antigen (Greer Labs, Lenoir, NC), and lipopolysaccharide (LPS) for 24, 24, and 4 h, respectively. The cells were collected and stained for the same surface markers as above. After staining, they were analyzed on an LSRII flow cytometric machine for detection.
Stimulation assay and enzyme-linked immunosorbent assay (ELISA) procedures
Using passage 2 PDEECs from duplicate samples from 2 EoE-Active and 2 non-inflammatory control patients, we plated 25,000 PDEECs per well onto half of a 48-well (0.95 cm2) plate each. After allowing the cells to grow for 48 h with epithelial reprogramming medium, the cells were serum starved for 4 h. The medium was replaced with 0.25 ml fresh serum-starved medium. Using 8 wells for each patient, 50,000 previously isolated autologous CD4+ T-cells matched with the appropriate patient were added. Another 8 wells were used to add 50,000 isolated autologous CD8+ T-cells. The last eight wells were left without T-cells. At this time, the cells were stimulated, in duplicate for all conditions, with either 10 μl of LPS, milk protein, or dust mite extract for a final concentration of 10 μg/ml. A vehicle control of 10 μl was also plated in duplicate. After 72 h, supernatants were collected.
Using supernatants previously collected, ELISAs were performed for the cytokines IL-6 (DY206) and TNF-α (DY210) using the protocol provided (R&D Systems, Minneapolis, MN). Results were measured on Synergy Mx Microplate Reader (BioTek, Winooski, VT).
Statistical analysis
Using GraphPad Prism (GraphPad Software, LaJolla, CA), we performed non-parametric Kruskal–Wallis analysis and Dunn's multiple comparison post-test to analyze growth kinetics. Integrin profiles were analyzed with two-way analysis of variance (ANOVA) with Tukey’s multiple comparison test. We also performed non-parametric Mann–Whitney analysis on integrin dimers, quantitative reverse transcriptase PCR (qRT-PCR), and antigen-stimulation assays. MHC profiles were analyzed using a two-way ANOVA with Bonferroni multiple comparisons. A p-value of ≤0.05 was considered statistically significant (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001). Figures were generated using GraphPad Prism.
Results
Morphologic and phenotypic characterization of conditional reprogrammed PDEECs
Esophageal biopsies were obtained from 28 patients who were being screened for potential diagnosis of EoE or known to have EoE and were undergoing surveillance endoscopy.
Cells taken at passages 1 and 3 were characterized via immunofluorescence staining for phenotypic changes over time. Cells were evaluated for the epithelial marker E-cadherin and tight junction protein ZO-1 that demonstrated no change in staining over time. Cells were still dividing as indicated by the presence of Ki67, a mitotic protein involved in cell division, and expressing stem cell marker p63 indicating that these cells maintained a stem-like state (Fig. 1a). Cells did not demonstrate any morphologic differences regardless of disease state (Fig. 1b).
PDEECs from non-inflammatory control and EoE patients maintain phenotypic expression and morphologic characteristics over multiple passages in culture. a Cells plated at passages 1 and 3 from a non-inflammatory control and EoE-Active patients were also plated on coverslips and immunostained for epithelial, stem cell, and proliferation markers. After three passages, cells from both patients were proliferative and still portrayed an epithelial phenotype. Magnification ×100. b PDEECs isolated from biopsies obtained from a non-inflammatory control and EoE-Active patient showed similar morphology
Evaluation of growth kinetics across all patient groups
Population doubling time was calculated for each patient group in order to evaluate the effect of disease process on cell growth in culture. Doubling times for all 6 non-inflammatory controls, 8 EoE-Remission, and 14 EoE-Active had similar growth profiles. The doubling times were 30.5, 24.5, and 29 h respectively. There was no statistical significance (p < 0.05) between groups (data/figure not shown). It took approximately 2 weeks for the initial passage cells to become confluent and about 1 week for passage 1–3 cells to become confluent (data not shown).
Changes of integrin profiles over time and between patient groups
Integrin profiles were evaluated via flow cytometry at passages 1 and 3 and data were expressed as percentages of viable cells positive. Integrin profile for non-inflammatory controls and EoE-Remission did not appear to change over time (Fig. 2a, b). However, EoE-Active PDEECs had a statistically significant less percentage of viable cells expressing integrin Alpha 1 (p < 0.05), Alpha 6 (p < 0.05), Beta 4 (p < 0.05), and Ki67 (p < 0.01) in passage 1 compared to non-inflammatory controls (Fig. 2a). EoE-Active PDEECs also had a statistically significant lower expression, compared to EoE-Remission, in Alpha 6 (p < 0.001) and Beta 4 (p < 0.05) (Fig. 2a). The percentage of viable cells increased in passage 3 to those seen in non-inflammatory controls and EoE-Remission (Fig. 2b). Expression of E-cadherin was lower in passage 1 in EoE-Active but increased to normal frequency in passage 3 (p < 0.05) (Fig. 2d). Alpha 6 (p < 0.001) and Beta 4 (p < 0.01) were significantly lower in EoE-Active in passage 1 vs. 3 (Fig. 2d). There was no statistical significance in integrin expression in the non-inflammatory controls and EoE-Remission groups in passage 1 or 3 (Fig. 2c, e).
Integrin profiles from Normal, EoE-Active, and EoE-Remission PDEECs at passages 1 and 3. a Passage 1 demonstrates lower Alpha 1, Alpha 6, and Beta 4 in EoE-Active. b Passage 3 demonstrates similar integrin profiles across all groups. Panels (c, d) demonstrate similar integrin profiles in passage 1 vs. 3 in non-inflammatory controls and EoE-Remission patients. Panel (e) demonstrate increased percentage of viable cells expressing Alpha 6 and Beta 4 in passage 3 compared to passage 1 in EoE-Active. In all figures, CD90 (mesenchymal marker) and vimentin (fibroblast marker) demonstrate low frequency. EpCam is high, which prove that our cultures are predominately epithelial cells. (5 Normal, 14 EoE-Active, 6 EoE-Remission)
Integrin dimers were examined by looking at double-positive signals for Alpha 6/Beta 4 (laminin receptor), Alpha 1/Beta 2 (collagen receptor), and Alpha 2/Beta 2 (collagen receptor). The percentage of viable cells expressing Alpha 6/Beta 4 showed little difference between non-inflammatory controls and EoE-Remission groups in passage 1 or 3 of the PDEECs. In EoE-Active patient cell lines, there is a lower expression of the Alpha 6/Beta 4 dimer at passage 1 but at passage 3 the dimer is consistent with the non-inflammatory control and EoE-Remission groups. Alpha 6/Beta 4 dimer was significantly lower in EoE-Active vs. non-inflammatory controls in passage 1 (p < 0.05) (Fig. 3a) but increased to normal frequency in passage 3 (Fig. 3b). Alpha 1/Beta 2 and Alpha 2/Beta 2 (not shown) indicated little difference in these double-positive populations among the groups. There was no statistical significance (p < 0.05) between the other groups.
Integrin dimer of passages 1 and 3 PDEECs. In passages 1 (a) and 3 (b), cells across all groups show a similar profile of the Alpha 6/Beta 4 dimer. At passage 1 (a), the percentage of viable EoE-Active cells expressing the dimer is significantly lower than in non-inflammatory controls. In EoE-Active, there is an increase of frequency in the Alpha 6/Beta 4 heterodimer at passage 3 compared to passage 1
Gene expression profiles of non-inflammatory controls and EoE patients demonstrate PDEECs maintain their gene expression when expanded over time
Gene expression was evaluated by using qRT-PCR comparing patient biopsy tissue to passage 3 PDEECs. Biopsies and PDEECs from 5 non-inflammatory controls and 9 EoE-Active patients were evaluated. Genes known to be associated with EoE (TSLP, Eotaxin-3/CCL26, and IL-33) were elevated in EoE-Active compared to non-inflammatory controls samples, although this did not reach statistical significance. Gene expression for CRISP3, SPINK7, IL-18, TJP1, EpCAM, and P63 were similar between the patient groups, while CDH1 was elevated in non-inflammatory controls compared to the EoE-Active groups. Expression of TJP1, in both groups, is significantly elevated indicating the successful isolation of epithelial cells from biopsy tissue (Fig. 4). There was no statistical significance (p < 0.05) between the EoE-Active and non-inflammatory control groups. IL-5 and Alox15 were not detectable (data not shown).
Identifying gene expression differences between biopsies and passage 3 PDEECs. Gene expression values were very similar between biopsies and passage 3 cells; however, there was an elevation in TJP1, which is observed mostly in epithelial cells. This result confirms that the passage 3 cells are epithelial cells. IL-33 was also elevated in EoE-Active (n = 9) patients in passage 3 compared to non-inflammatory control patients (n = 5)
Epithelial cells express MHC class II after stimulation with antigens
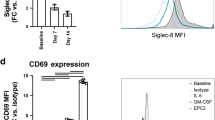
To determine whether PDEECs express MHC class II molecules before and after antigen stimulation, PDEECs from passage 2 from 5 non-inflammatory controls, 5 EoE-Active, and 5 EoE-Remission patients were stimulated with dust mite antigen, milk antigen, and LPS. Surface detection of MHC class II on viable positive cells was determined by flow cytometry. The results demonstrated little to no positive MHC class II markers in non-stimulated cells (<5% of viable cells), while all three stimulated groups had a significant increase in positive cells compared to the non-stimulated group (20–80% of viable cells in the non-inflammatory control, 20–60% of viable cells in the EoE-Active, and 20–80% in the EoE-Remission groups) (Fig. 5). All groups had a significant increase in viable MHC class II-positive cells after stimulation with antigens (p < 0.05).
PDEECs express MHC class II molecules after stimulation. MHC class II-positive cells were not detected on non-stimulated cells, while 20–80% of stimulated cells expressed MHC class II molecules. All individual groups, compared to the non-stimulated, show a significant increase in the percentage of MHC class II-positive cells. Additionally, the profiles show non-significant differences between the stimulated groups
PDEECs react to antigens in the presence of T-cells
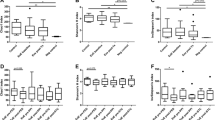
To evaluate whether the PDEECs were functional or not, we stimulated them with LPS, dust mite, and milk antigen, in the presence or absence of autologous T-cells in a co-culture system. There was higher cytokine production in EoE-Active (n = 2) patient samples compared to non-inflammatory controls (n = 2). Additionally, there was a more robust response when the epithelial cells were co-cultured with CD4+ or CD8+ T-cells (Fig. 6). Upon stimulation of PDEECs with LPS and dust mite antigen, IL-6 levels in EoE-Active were significantly higher than non-inflammatory controls (Fig. 6a, b). IL-6 levels were significantly higher in EoE-Active after stimulation with LPS, dust mite, and milk antigens in the presence of CD4+ T-cells (p < 0.05) (Fig. 6a–c). TNF-α was also significantly higher in EoE-active vs. non-inflammatory controls after stimulation with LPS with and without the presence of CD4+ or CD8+ T-cells (Fig. 6d). There were no significant differences in TNF-α secretion after stimulation with dust mite or milk in the presence or absence of T-cells (Fig. 6e, f).
Cytokine levels in supernatants of stimulated PDEECs in the presence or absence of autologous CD4+ and CD8+ cells. ELISA data for pro-inflammatory cytokines IL-6 (a, b, c) and TNF-α (d, e, f) are shown. These results show an increased cytokine level in all EoE-Active patients compared to non-inflammatory controls. There is also a more robust response when PDEECs were stimulated in the presence of CD4+ and CD8+ T-cells
Discussion
Prior to our study, EoE research has focused on immortalized cell lines or on biopsy gene expression levels. Most research studies have focused on developing a diagnostic test or identifying diagnostic biomarkers for EoE rather than in trying to develop a test that identifies the causative antigens. There has been limited work in using patient-specific cells and expanding these cells for the purpose of in vitro testing. In this study, we sought to characterize our PDEECs and investigate their functionality and possible role in EoE.
By developing individualized, PDEECs, we were able to characterize these cells and identify differences in integrin profiles and gene expression in EoE-Active vs. EoE-Remission and non-inflammatory controls. Epithelial integrin profiles have never been evaluated or investigated in EoE. Our hypothesis was that, since there is evidence of abnormal epithelial barrier,14 changes in tight junctions,31 and intracellular spaces32,33 in EoE, integrins must be affected as well. Integrins are able to mediate cell adhesion in modulating signaling cascades in cell motility, survival, proliferation, and differentiation. Integrins are found as heterodimeric transmembrane receptors consisting of α and β subunits. The α6β4 heterodimer, a laminin receptor, has been shown to play a role in signaling that stimulates epithelial cell migration and stabilizes or enhances the formation of actin-rich protrusions that are needed for cell movement.34 The major function of the α6β4 integrin is to maintain the integrity of epithelia.35,36 The decreased levels of α6β4 heterodimer in passage 1 of EoE-Active PDEECs may indicate that these cells are still in an inflammatory state, and as the cells are reprogrammed and expanded, the integrin profiles essentially normalize.
TSLP is an epithelial-derived cytokine that has been linked to Th2-type inflammatory responses in gastrointestinal diseases and is identified to be upregulated in EoE.37 Chandramouleeswaran et. al. showed that specific food antigens can trigger innate immune-mediated esophageal TSLP secretion, suggesting that esophageal epithelial cells at the barrier surface may play a significant role in the pathogenesis of EoE by regulating TSLP expression.37 Like TSLP, IL-33 has been found to be elevated in active EoE. IL-33 likely contributes to the Th2-type cell inflammatory response in EoE, specifically by stimulating eosinophil migration and activation via IL-13.38 IL-33 has also been shown to promote gastrointestinal allergy in a TSLP-independent manner.39 Increased gene expression levels of both IL-33 and TSLP at passage 3 in EoE-Active PDEECs suggest that there is a higher concentration of epithelial cells, due to the process of reprogramming, than at passage 1. In addition, the increased expression indicates that IL-33 and TSLP are playing a role in EoE-Active PDEECs that may be reflective of what is happening in vivo.
Professional APCs, such as dendritic cells and macrophages, mediate the cellular immune response by presenting antigens to T-cells via MHC class II molecules. In the squamous epithelium, there are little to no dendritic cells and macrophages. However, it has been demonstrated that epithelial cells, specifically in EoE, may behave as NPAPCs.13 We demonstrated that, when stimulated with dust mite, milk antigen, or LPS, there is an increase in the percentage of positive viable PDEECs with MHC class II surface markers. This confirms that these PDEECs appear to have the capability of taking on the role as NPAPCs and potentially assuming the role as their parent cells.
Since these cells are able to behave as NPAPCs with MHC class II surface molecules being activated, the addition of antigens could enact a response from the PDEECs. The inflammatory response that was shown of PDEECs with autologous CD4+ and CD8+ T-cells in the presence of a stimulating antigen has never been demonstrated. We have demonstrated that PDEECs from EoE-Active patients, compared to non-inflammatory controls, react to antigens and produce IL-6 in the presence of T-cells. It is also reassuring to see that the PDEECs from EoE-Active patients, as opposed to non-inflammatory controls, reacted to milk antigen. These EoE-Active patients eventually were responders to dairy-free diet. Lastly, PDEECs from EoE and non-inflammatory control patients expressed MHC class II molecules after antigen stimulation but only the EoE-Active patients respond by secreting cytokines in the presence of CD4+ T-cells. This is an indication that the autologous CD4+T-cells are already primed and the interaction between the PDEECs and T-cells is required. If these cells function similarly to their parent cells in vivo and behave as NPAPCs, their ability to respond to stimuli in vitro will allow us to expand the utility of these cells into diagnostic and therapeutic testing assays.
There are many strengths to our study. This was a prospective study performed on pediatric tissue obtained at the time of endoscopy and processed immediately with a large number of patients in the study expanding over different groups (EoE-Active, EoE-Remission, and normal non-inflammatory controls). This is the first study of its kind to develop patient-specific and PDEEC lines from patients with EoE.
In summary, we have demonstrated that we are able to successfully isolate esophageal epithelial cells, successfully expand these cells over multiple passages without any genetic modification, and thus develop patient-specific esophageal epithelial cell lines. PDEECs, while different between diseased and normal, essentially maintain their phenotype and genotype throughout. Upon activation, PDEECs will still function in vitro as NPAPCs allowing for further studies of patient-specific cells and their reactions to specific antigens that is similar to what is happening in vivo. This will allow for future use of these cells in either diagnostic and/or therapeutic testing for EoE.
References
Mishra, A., Schlotman, J., Wang, M. & Rothenberg, M. E. Critical role for adaptive T cell immunity in experimental eosinophilic esophagitis in mice. J. Leukoc. Biol. 81, 916–924 (2007).
Straumann, A. Idiopathic eosinophilic gastrointestinal diseases in adults. Best Pract. Res. Clin. Gastroenterol. 22, 481–496 (2008).
Sayej, W. N. et al. Characterizing the inflammatory response in esophageal mucosal biopsies in children with eosinophilic esophagitis. Clin. Transl. Immunol. 5, e88 (2016).
Ehrhardt, C., Kim, K. J. & Lehr, C. M. Isolation and culture of human alveolar epithelial cells. Methods Mol. Med. 107, 207–216 (2005).
Zubeldia-Plazaola, A. et al. Comparison of methods for the isolation of human breast epithelial and myoepithelial cells. Front. Cell Dev. Biol. 3, 32 (2015).
Spurrier, R. G., Speer, A. L., Hou, X., El-Nachef, W. N. & Grikscheit, T. C. Murine and human tissue-engineered esophagus form from sufficient stem/progenitor cells and do not require microdesigned biomaterials. Tissue Eng. Part A 21, 906–915 (2015).
Roche, J. K. Isolation of a purified epithelial cell population from human colon. Methods Mol. Med. 50, 15–20 (2001).
Southgate, J., Hutton, K. A., Thomas, D. F. & Trejdosiewicz, L. K. Normal human urothelial cells in vitro: proliferation and induction of stratification. Lab. Invest. 71, 583–594 (1994).
Kalabis, J. et al. Isolation and characterization of mouse and human esophageal epithelial cells in 3D organotypic culture. Nat. Protoc. 7, 235–246 (2012).
Geraghty, R. J. et al. Guidelines for the use of cell lines in biomedical research. Br. J. Cancer 111, 1021–1046 (2014).
Liu, X. et al. ROCK inhibitor and feeder cells induce the conditional reprogramming of epithelial cells. Am. J. Pathol. 180, 599–607 (2012).
Arebro, J. et al. Antigen-presenting epithelial cells can play a pivotal role in airway allergy. J. Allergy Clin. Immunol. 137, 957–60 e7 (2016).
Mulder, D. J. et al. Antigen presentation and MHC class II expression by human esophageal epithelial cells: role in eosinophilic esophagitis. Am. J. Pathol. 178, 744–753 (2011).
Simon, D. et al. Evidence of an abnormal epithelial barrier in active, untreated and corticosteroid-treated eosinophilic esophagitis. Allergy 73, 239–247 (2018).
Sato, M. et al. Increased expression of integrins by heparin-binding EGF like growth factor in human esophageal cancer cells. Cancer Lett. 102, 183–191 (1996).
Akiyama, S. K. Integrins in cell adhesion and signaling. Hum. Cell 9, 181–186 (1996).
Borghi, N., Lowndes, M., Maruthamuthu, V., Gardel, M. L. & Nelson, W. J. Regulation of cell motile behavior by crosstalk between cadherin- and integrin-mediated adhesions. Proc. Natl. Acad. Sci. USA 107, 13324–13329 (2010).
Hynes, R. O. Integrins: bidirectional, allosteric signaling machines. Cell 110, 673–687 (2002).
Blanchard, C. et al. A striking local esophageal cytokine expression profile in eosinophilic esophagitis. J. Allergy Clin. Immunol. 127, 208–217 (2011).
Zuo, L. et al. IL-13 induces esophageal remodeling and gene expression by an eosinophil-independent, IL-13R alpha 2-inhibited pathway. J. Immunol. 185, 660–669 (2010).
Blanchard, C. et al. IL-13 involvement in eosinophilic esophagitis: transcriptome analysis and reversibility with glucocorticoids. J. Allergy Clin. Immunol. 120, 1292–1300 (2007).
Niranjan, R. et al. Involvement of interleukin-18 in the pathogenesis of human eosinophilic esophagitis. Clin. Immunol. 157, 103–113 (2015).
Blanchard, C. et al. Eotaxin-3 and a uniquely conserved gene-expression profile in eosinophilic esophagitis. J. Clin. Invest. 116, 536–547 (2006).
Bhattacharya, B. et al. Increased expression of eotaxin-3 distinguishes between eosinophilic esophagitis and gastroesophageal reflux disease. Hum. Pathol. 38, 1744–1753 (2007).
Sleiman, P. M. et al. GWAS identifies four novel eosinophilic esophagitis loci. Nat. Commun. 5, 5593 (2014).
Matoso, A. et al. Expression microarray analysis identifies novel epithelial-derived protein markers in eosinophilic esophagitis. Mod. Pathol. 26, 665–676 (2013).
Liacouras, C. A. et al. Eosinophilic esophagitis: updated consensus recommendations for children and adults. J. Allergy Clin. Immunol. 128, 3–20e6 (2011).
Jensen, T. J., Foster, C., Sayej, W. & Finck, C. M. Conditional reprogramming of pediatric human esophageal epithelial cells for use in tissue engineering and disease investigation. J. Vis. Exp. 121 (2017); doi: 10.3791/55243.
Janisch, H. D. et al. Histologic abnormalities in routine biopsies of patients with esophagitis and different gastric acid secretion. Appl. Pathol. 2, 272–276 (1984).
Freshney, R. I. Culture of Animal Cells A Manual of Basic Technique. (John Wiley & Sons, Hoboken, NJ, 2005).
Okimoto, K. et al. A prospective study of eosinophilic esophagitis and the expression of tight junction proteins in patients with gastroesophageal reflux disease symptoms. Gut Liver 12, 30–37 (2017).
Ravelli, A. et al. Dilated intercellular spaces in eosinophilic esophagitis. J. Pediatr. Gastroenterol. Nutr. 59, 589–593 (2014).
Ravelli, A. M. et al. Dilated intercellular spaces: a major morphological feature of esophagitis. J. Pediatr. Gastroenterol. Nutr. 42, 510–515 (2006).
Mercurio, A. M., Rabinovitz, I. & Shaw, L. M. The alpha 6 beta 4 integrin and epithelial cell migration. Curr. Opin. Cell Biol. 13, 541–545 (2001).
Larjava, H., Salo, T., Haapasalmi, K., Kramer, R. H. & Heino, J. Expression of integrins and basement membrane components by wound keratinocytes. J. Clin. Invest. 92, 1425–1435 (1993).
van der Neut, R., Krimpenfort, P., Calafat, J., Niessen, C. M. & Sonnenberg, A. Epithelial detachment due to absence of hemidesmosomes in integrin beta 4 null mice. Nat. Genet. 13, 366–369 (1996).
Chandramouleeswaran, P. M. et al. Preferential secretion of thymic stromal lymphopoietin (TSLP) by terminally differentiated esophageal epithelial cells: relevance to eosinophilic esophagitis (EoE). PLoS ONE 11, e0150968 (2016).
Judd, L. M. et al. Elevated IL-33 expression is associated with pediatric eosinophilic esophagitis, and exogenous IL-33 promotes eosinophilic esophagitis development in mice. Am. J. Physiol. Gastrointest. Liver Physiol. 310, G13–G25 (2016).
Han, H., et al. IL-33 promotes gastrointestinal allergy in a TSLP-independent manner. Mucosal Immunol. 11, 578 (2018).
Acknowledgements
We would like to thank our research coordinators Miriam Lincoln, Chih-Chen Jennifer Wong, and Krista Spada for all their help in recruiting patients. We also would like to thank Evan Jellison, PhD, from the flow cytometry core at the University of Connecticut Health for helping with the flow cytometric analysis. This work was supported by Adopt a Friend Grant from the Connecticut Children’s Medical Center Foundation and Connecticut Children’s Medical Center Strategic Research Funds.
Author contributions
All the authors contributed to writing, editing, and revising the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Sayej, W.N., Foster, C., Jensen, T. et al. Expanding and characterizing esophageal epithelial cells obtained from children with eosinophilic esophagitis. Pediatr Res 84, 306–313 (2018). https://doi.org/10.1038/s41390-018-0033-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-018-0033-9