Abstract
Background
Exosomes are nanovesicles originating from multivesicular bodies that have complex functions and significant therapeutic effects in many diseases. In the present study, we successfully extracted exosomes from Pseudomonas aeruginosa and assessed the effect of those exosomes on the development of the allergic response in two types of classic asthma models.
Methods
Female BALB/c mice were administrated with P. aeruginosa-derived exosomes 1 week before ovalbumin (OVA) or house dust mite (HDM) sensitization. Bronchoalveolar lavage fluid, serums and lung tissues were collected and analyzed for pathophysiology and immune responses.
Results
Our results demonstrated that P. aeruginosa-derived exosomes inhibited the development of airway hyper-responsiveness (AHR), peribronchial and perivascular inflammation in lung tissues and the level of serum IgE. Moreover, this protective effect was associated with an increase in the regulatory T cell (Treg) response and a concomitant decreased Th2 response.
Conclusions
In conclusion, these observations demonstrated that P. aeruginosa-derived exosomes could induce protection against allergic sensitization in asthma mice, and our study provided a new insight to prevent allergic diseases.
Similar content being viewed by others
Introduction
Asthma is the most common chronic disease of childhood, and is characterized by airway hyper-responsiveness, mucus hypersecretion, airway inflammation, and increased eosinophils. At present, the treatments of asthma include mainly glucocorticoid, β2 receptor agonists, and leukotriene antagonists. Despite the use of the above prescription drugs, the incidence of asthma is still increasing, bringing a heavy burden to the family and society.1 Moreover, about 10% of asthmatic cannot be controlled by drugs.2 Therefore, it is essential to explore the underlying pathogenesis and to find new ways to prevent and treat asthma.
Asthma is a heterogeneous disease that encompasses many different etiologies and pathophysiologies, which have been called “asthma endotypes”. The most representative and common “endotype” is a T-cell immune imbalance disease associated with an imbalance in Th1/Th2 cells.3 Induction of the Th2 response leads to AHR, mucus hypersecretion and airway remodeling.4 In recent years, it has been shown that regulatory T (Treg) cells played an important role in asthma. Treg cells can restore the balance of Th1 and Th2 cell function.5 However, Treg cells are often impaired in asthma, and transfusion of Treg cells to sensitized mice has been shown to relieve asthma phenotypes.6 Thus, both increase in the level of Treg cells, along with the balance in Th1/Th2 response are important for prevention or treatment of asthma.
Exosomes have an endocytic origin and are formed by the reverse budding of the peripheral membrane of multivesicular bodies.7 Exosomes are between 30 and 100 nm in diameter, and are released by almost all cell types. The exosomes have been extracted from the urine, saliva, blood, and milk. Recently, exosomes were also extracted from two kinds of Gram-negative bacteria Pseudomonas aeruginosa and Escherichia coli.8 Previous studies have shown that exosomes played important role in intercellular communication and numerous other physiological processes, both locally and systemically. In addition to tissue repair and diagnosis, recent studies showed that exosomes from both immune cells and non-immune cells could effectively inhibit the immune response in autoimmune diseases,9,10 and their immunomodulatory effects were associated with Treg responses,11 which could further suppress the activation of effector T cells.
Studies have shown that bacterial CpG oligonucleotides can effectively induce mammalian immune response through the TLR9 signaling pathway and therefore can be used as a good immune adjuvant for the treatment of immune diseases. However, synthetic CpG oligonucleotides are often difficult to enter cells, and finding a stable, highly active and biocompatible carrier system is the key to the application of CpG drugs.12 Lipopolysaccharide (LPS) is a major component of the outer cell wall of Gram negative bacteria. In recent years, epidemiologic evidence has confirmed that there is lower incidence of asthma or atopic diseases among children who grow up on farms, especially those exposed to LPS, indicating a immunomodulatory effect of LPS on asthma and allergic diseases.13,14 Although LPS has a prominent protective effect on asthma, LPS or Gram-negative bacteria cannot be directly applied into the human body as a preventive measure of asthma and allergic diseases. As commercial LPS is pathogenicity and toxicity, and is not natural biological origin. In addition, the ingredients of Gram-negative bacteria is complex, and administration of Gram-negative bacteria is liable to lead to an infection as it is hard to control the appropriate dosage. Recently, some studies suggested that outer membrane vesicles secreted by Gram-negative bacteria could deliver LPS to the host cell cytosol, which provided a new insight for the protection of asthma and allergic diseases.15,16 Meanwhile, our group has successfully extracted exosomes from Pseudomonas aeruginosa, one of the Gram-negative bacteria. Therefore, in this study, P. aeruginosa-derived exosomes were used to mimic the natural environmental LPS exposure, and our aim was to study whether P. aeruginosa-derived exosomes can protect against allergic asthma.
Methods
Animals
Female BALB/c (5 weeks of age) mice were housed in individually filtered pathogen-free cages, maintained on a 12-h light/dark cycle with an ovalbumin (OVA)-free diet, under a constant room temperature (24 °C). Adequate amounts of sterile animal food and water were provided. Cages, food, bedding, and water were sterilized before use. The Institutional Animal Care and Research Advisory Committee at Chongqing Medical University reviewed and approved all animal procedures performed in this study. The use of animals in these experiments was in accordance with the guidelines issued by the Chinese Council on Animal Care.
Bacterial strains and growth media
The P. aeruginosa (ATCC 15692) reference laboratory strain was routinely grown in Luria-Bertani (LB) broth at 37 °C to the exponential growth phase.
Exosome isolation and concentration determination
P. aeruginosa-derived exosomes were isolated, as described with minor modifications.8 Isolated exosomes were quantified by measuring their protein content. Exosomes were re-suspended in PBS and stored at −80 °C until use.
Lipopolysaccharide (LPS) detection in the exosomes
LPS in the P. aeruginosa-derived exosomes was detected by Limulus Amebocyte Lysate (LAL) test according to the previous study17 in Supplemental Table S1.
Electron microscopy
Isolated exosomes were resuspended in PBS, adsorbed in formvar-coated nickel grids and negatively stained with 4% uranyl oxalate. Grids were air dried and observed with a Hitachi-7500 Transmission Electron Microscope. The exosomes were characterized by studying morphology and size (<100 nm) using an electron microscope.
Exosome administration and mouse models
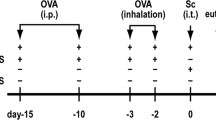
In a pilot experiment, intranasal (i.n.) pretreatments of P. aeruginosa-derived exosomes were performed to determine the optimal dose and time of exosomes administration. Through this approach, application of 50 μg i.n. of exosomes on 3 consecutive days was chosen as the optimal protocol. Anesthetized mice were i.n. pre-treated with exosomes from P. aeruginosa 1 week before OVA or HDM sensitizations and challenge. The OVA or HDM-induced asthma model was established according to the previous published methods18, and details were provided in Fig. 1. The experiments were performed with 7–8 mice/group.
Overview of the study protocol. Exosomes were introduced i.n. to 5 weeks old BALB/c mice for consecutively 3 days. At 7 and 21 days, after exosomes treatment, mice were either sensitized with OVA (a) or HDM (b) or were given PBS as a control. a From 28th days to 37th days, OVA or PBS-sensitized mice were challenged with either OVA or PBS. b At days 28, 30, 32, 34, and 36, HDM or PBS-sensitized mice were challenged with HDM or PBS. At days 38 (a) or 37 (b), all mice were killed
Airway hyper-responsiveness (AHR)
After 24 h of the final OVA/HDM challenge, lung resistance (LR) was measured using an invasive lung function test. Briefly, anesthetized mice were intubated and mechanically ventilated using a computer controlled piston ventilator (flexiVent, Scireq). Mice were then challenged with an aerosolized bronchoconstrictor, acetyl-β-methacholine (Sigma-Aldrich, Saint Louis, MO), at increasing doses: 0, 3.125, 6.25, 12.5, 25, and 50 mg/mL. At each dose, lung resistance was calculated using the single-compartment model.
Bronchoalveolar lavage and cell counting
Within 24 h of final OVA/HDM challenge, mice were killed and the trachea was cannulated, and bronchoalveolar lavage fluid (BALF) was obtained by flushing the lungs three times with 1.5 mL PBS. Total cell numbers in the BALF were counted microscopically. Differential cell counts were performed under Wright–Giemsa staining and based on standard morphologic and staining characteristics of at least 200 cells per sample. The supernatant was stored at −80 °C for further use. All slides were characterized by a single-blinded examiner to reduce evaluator bias.
Histological analysis
After 24 h of the final OVA/HDM challenge, mice were killed and the lungs were harvested. Formalin-fixed lungs were dissected and embedded in paraffin. Paraffin blocks were serially sectioned at a thickness of 4 μm and stained with hematoxylin-eosin (H&E). Images were captured under a Nikon Eclipse E200 microscope connected to a Nikon Coolpix 995 camera (Nikon, Tokyo, Japan). The severity of inflammation was evaluated by assigning a value of 0 point for normal; 1 point for few cells; 2 points for a ring of inflammatory cells 1 cell layer deep; 3 points for a ring of inflammatory cells 2–4 cells deep; 4 points for a ring of inflammatory cells of >4 cells deep.
Flow cytometric analysis
The lungs were minced and incubated for 20 min at 37 °C in 1 mL of sterile PBS containing 0.2% collagenase I (Sigma-Aldrich). Single-pulmonary cell suspensions were obtained by forcing tissue through a 70 μm cell filter (Becton, Dickinson and Company, Franklin Lakes, NJ). Erythrocytes were lysed, and the remaining cells were resuspended in RPMI 1640 medium containing 10% fetal bovine serum. A single-cell suspension from lung (2 × 106 cells/mL) was incubated for 4–6 h at 37 °C and 5% CO2 in six-well flat-bottom plates (Nalgene of Thermo Fisher Scientific, Waltham, MA,) in 1 mL medium containing phorbol 12-myristate 13-acetate (50 ng/mL; Sigma-Aldrich), ionomycin (500 ng/mL; Sigma-Aldrich) and GolgiPlug-containing brefeldin A (Becton, Dickinson and Company). The cells were then collected, washed, pretreated with an Fc blocker, and subsequently stained for surface-associated CD4 (anti-CD4-FITC; Gk1.5, Pharmingen of Becton, Dickinson and Company), CD3 (anti-CD3-PerCP-Cy5.5; 145-2C11, Pharmingen), or CD25 (anti-CD25-PE; 3C7, Pharmingen). To detect the subsets of CD4+ T cells in lungs, the cells were stained for intracellular IFN-γ (anti-IFN-γ-BV421; XMG1.2, Pharmingen), IL-4 (anti-IL-4-APC; 11B11, Pharmingen), and FOXP3 (anti-FOXP3-APC; FJK-16s, eBioscience), and detected by flow cytometry (FACS Canto; Becton, Dickinson and Company). The data were analyzed with Cell Quest software (Becton, Dickinson and Company).
RNA extraction, reverse transcription, and quantitative PCR (qPCR)
Total RNA from mouse lung tissues was purified, and cDNA synthesis was performed using a PrimeScript RT Reagent Kit according to the manufacturer’s recommendations (Takara, Japan). Briefly, total RNA was extracted from different samples using Trizol reagent (Invitrogen; Carlsbad, Calif.) according to the manufacturer’s instructions. After quantification, the RNA was reverse transcribed using a PrimeScript RT reagent kit (Takara, Japan). Real-time quantitative PCR (q-PCR) was performed to detect the relative expression levels of Foxp3, T-bet and GATA3. The real-time PCR conditions were as follows: 95 °C for 3 min, followed by 40 cycles at 95 °C for 10 s and 60 °C for 30 s. GAPDH was used as an endogenous control. The primer sequences of Foxp3: 5′-TGGAACCACGGGCACTATCACA-3′ (forward) and 5′-GAGGCTGCGTATGATCAGTTATGC-3′ (reverse), T-bet were 5′-AGCAAGGACGGCGAATGTT-3′ (forward) and 5′-GGGTGGACATATAAGCGGTTC-3′ (reverse), GATA3 were 5′-CTCGGCCATTCGTACATGGAA-3′ (forward) and 5′-GGATACCTCTGCACCGTAGC-3′ (reverse), and GAPDH were 5′-AGCAATGCCTCCTGCACCACCAAC-‘3′ (forward) and 5′-CCGGAGGGGCCATCCACAGTCT-3′ (reverse).
BALF cytokine measurements
Cytokine concentrations in BALF were measured with commercial enzyme-linked immunosorbent assay (ELISA) kits according to the manufacturer’s instructions. ELISA kits were used to measure interferon (IFN)-γ (EMC101g.96, NeoBioscience, Shenzhen, China), interleukin (IL)-4 (EMC003.96, NeoBioscience, Shenzhen, China), IL-5 (EMC108.96, NeoBioscience, Shenzhen, China), IL-13 (EMC124.96, NeoBioscience, Shenzhen, China), IL-10 (EMC005.96, NeoBioscience, Shenzhen, China) and transforming growth factor (TGF)-β (EMC107b.96, NeoBioscience, Shenzhen, China). OVA-specific IgE was assessed by ELISA (439807, BioLegend, San Diego, CA). HDM-specific IgE expression was assessed using a modified protocol.19 In brief, HDM was coated onto 96-well plates (2.5 μg/well) and incubated overnight at 4 °C. After blocking with 5% FBS in PBS, undiluted serum was added and incubated overnight at 4 °C. After washing, biotin anti-mouse IgE (BD Bioscience-San Jose, CA) was added and incubated at 37 °C for 1 h.
Statistical analyses
Data are expressed as the mean and its standard error (SEM). The GraphPad Prism 4.0 program (GraphPad, San Diego, CA) was used for the statistical analyses. The statistical analyses were performed using two-way ANOVAs or two-way analysis of variance and Bonferroni’s post hoc test otherwise specified. P-values < 0.05 were considered to be statistically significant.
Results
P. aeruginosa-derived exosome identification and pre-treatment
Exosomes were isolated from P. aeruginosa by filtration and ultracentrifugation, and identified by electron microscopy (EM). EM analysis of P. aeruginosa-derived exosomes demonstrated round vesicles of 50–100 nm in diameter, each with a lipid bilayer membrane (Fig. 2). Therefore, the size and density of P. aeruginosa-derived exosomes corresponded to generally previous studies.20,21
Pre-treatment with P. aeruginosa-derived exosomes before OVA/HDM sensitization protects against allergic airway sensitization
Lung tissue from PBS-stimulated (PBS+PBS group) or exosome-stimulated (PBS+exosome group) mice had normal airways and the absence of inflammatory cells infiltrates (Figs. 3a, b and 4a, b, P < 0.05). OVA or HDM sensitized mice (OVA/HDM+PBS group) exhibited numerous inflammatory cells surrounding the airways and vessels. However, there was less cellular infiltration in OVA/HDM-sensitized mice that had received P. aeruginosa-derived exosomes (OVA/HDM+exosome group) (Figs. 3a, b and 4a, b, P < 0.05). AHR was measured to test the changes in lung function. As expected, AHR was significantly enhanced in the OVA/HDM+PBS group compared with the OVA/HDM+exosome and the PBS+PBS/exosome groups (Figs. 3c and 4c, P < 0.05). In parallel, OVA/HDM-specific immunoglobulin E (IgE) levels in the serum were measured with a standard enzyme-linked immunosorbent assay (ELISA). The mice in the OVA/HDM+PBS group had significantly higher levels of OVA/HDM-specific IgE in the serum compared with the OVA/HDM+exosome and the PBS+PBS/exosome mice (Figs. 3d and 4d, P < 0.05). In addition, total cell counts and differential cell counts in the BALF were determined. The total number of cells, neutrophils and eosinophils in the BALF increased significantly in OVA/HDM+PBS group in comparison with the OVA/HDM+exosome group and the PBS+PBS/exosome group (Figs. 3e and 4e, P < 0.05). These results confirmed that the P. aeruginosa-derived exosomes might responsible for the protection of allergic airway sensitization in the OVA/HDM-induced asthma model.
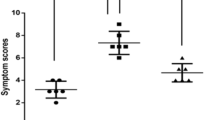
P. aeruginosa-derived exosomes inhibited the OVA-induced allergic response (Grouping: PBS+PBS, OVA+PBS, OVA+Exosome and PBS+Exosome). a Representative lung sections stained with H&E. Magnification: ×200. b After HE-stainning, lung tissue sections were scored for levels of alveolitis, perivasculitis and peribronchial inflammation. a Histological scores of pulmonary alveolitis; b Histological scores of pulmonary perivasculitis; c Histological scores of pulmonary peribronchiolitis. c Airway hyper-responsiveness was measured after challenge with methacholine. Data are represented as mean airway resistance in LR (m/(s*Pa)) ± SEM. LR lung resistance. d OVA-specific IgE (pg/mL) was measured with a standard ELISA. e The total number of cells a, neutrophils b, and eosinophils c in bronchoalveolar lavage fluid (BALF) were counted after Giemsa staining. The statistical analyses were performed using two-way ANOVAs (Panels b, d, e) and two-way analysis of variance and Bonferroni’s post hoc test (Panel c). *P < 0.05, **P < 0.01, ***P < 0.001 as compared with OVA+PBS mice, n = 6 mice per group
P. aeruginosa-derived exosomes inhibited the HDM-induced allergic response (Grouping: PBS+PBS, HDM+PBS, HDM+Exosome, and PBS+Exosome). a Representative lung sections stained with H&E. Magnification: ×200. b After HE-stainning, lung tissue sections were scored for levels of alveolitis, perivasculitis and peribronchial inflammation. a Histological scores of pulmonary alveolitis; b Histological scores of pulmonary perivasculitis; c Histological scores of pulmonary peribronchiolitis. c Airway hyper-responsiveness was measured after challenge with methacholine. Data are represented as mean airway resistance in LR (m/(s*Pa)) ± SEM. LR lung resistance. d HDM-specific IgE (pg/mL) was measured with a standard ELISA. e The total number of cells a, neutrophils b and eosinophils c in bronchoalveolar lavage fluid (BALF) were counted after Giemsa staining. The statistical analyses were performed using two-way ANOVAs (Panels b, d, e) and two-way analysis of variance and Bonferroni’s post hoc test (Panel c). *P < 0.05, **P < 0.01, ***P < 0.001 compared with HDM + PBS mice, n = 6 mice per group
Pretreatment with P. aeruginosa-derived exosomes induces Treg responses while suppressingTh2 responses
To determine the effect of the pretreatment with P. aeruginosa-derived exosomes on the T cell response, CD4+CD25+Foxp3+T (Treg), CD3+CD4+INF-γ+T (Th1) and CD3+CD4+IL-4+T (Th2), and their related cytokine production (TGF-β, IL-10, INF-γ, IL-4, IL-5 and IL-13) and transcription factor RNA (Foxp3 mRNA, T-bet mRNA and GATA3 mRNA) were measured. The CD4+CD25+Foxp3+ T (Treg) cells were significantly increased in the mice pre-treatment with P. aeruginosa-derived exosomes (OVA/HDM+exosome) compared with mice in OVA/HDM+PBS (Figs. 5a and 6a, P < 0.05). In the meanwhile, CD3+CD4+IL-4+ T (Th2) cells were significantly increased in mice that underwent OVA/HDM sensitization in comparison with the mice in control groups (PBS+PBS/exosomes). However, CD3+CD4+IL-4+ T (Th2) cells were significantly decreased after pre-treatment with P. aeruginosa-derived exosomes (OVA/HDM+exosome) (Figs. 5a and 6a, P < 0.05). No significant changes were detected in CD3+CD4+INF-γ+(Th1) levels between each group. Simultaneously, IL-5 and IL-13 production were decreased after pre-treatment with P. aeruginosa-derived exosomes in OVA-induced asthma model. IL-4, IL-5 and IL-13 production were decreased after pre-treatment with P. aeruginosa-derived exosomes in HDM-induced asthma model (Figs. 5b e, f and 6b d, e, f), P < 0.05). In addition, pre-treatment with P. aeruginosa-derived exosomes elevated the TGF-β and IL-10 production in OVA/HDM+exosome mice compared with mice in OVA/HDM+PBS (Figs. 5b a, b and 6b a, b), P < 0.05). No significant changes were detected in IFN-γ levels between each group (Figs. 5b, c and 6b, c), P < 0.05). In parallel, Foxp3 mRNA expression increased significantly in OVA/HDM sensitized mice receiving exosomes (OVA/HDM + exosome) and GATA3 mRNA expression was reduced in OVA/HDM+exosome mice compared with mice in OVA/HDM+PBS (Figs. 5c and 6c, P < 0.05). There were no difference in T-bet mRNA expression between these groups. Given these results, we demonstrated that the OVA/HDM sensitized mice which pre-treated with P. aeruginosa-derived exosomes had a higher overall proportion of regulatory T cells and lower Th2 cells responses.
P. aeruginosa-derived exosomes regulated OVA-induced immunological responses. P. aeruginosa-derived exosomes up-regulated Treg responses but down-regulated Th2 responses. a CD4+ CD25+Foxp3+ T (Treg), CD3+ CD4+ INF-γ+ T (Th1) and CD3+ CD4+IL-4+ T (Th2) was analyzed by FCM in lung tissue. b Cytokine production (TGF-β (a), IL-10 (b), INF-γ (c), IL-4 (d), IL-5 (e), and IL-13 (f)) in BALF supernatants from OVA-induced mice was analyzed using ELISA. c Transcription factor RNA levels (Foxp3 mRNA, T-bet mRNA and GATA3 mRNA) in the lungs of OVA-induced mice were analyzed using q-PCR. Results were normalized using a GAPDH control and show mean fold increase over control±SEM. The statistical analyses were performed using two-way ANOVAs. *P < 0.05, **P < 0.01, ***P < 0.001 compared with OVA+PBS group, n = 6 mice per group
P. aeruginosa-derived exosomes regulated HDM-induced immunological responses. P. aeruginosa-derived exosomes up-regulated Treg responses but down-regulated Th2 responses. a CD4+CD25+Foxp3+ T (Treg), CD3+CD4+ INF-γ+ T (Th1) and CD3+CD4+IL-4+ T (Th2) was analyzed by FCM in lung tissue. b Cytokine production (TGF-β (a), IL-10 (b), INF-γ (c), IL-4 (d), IL-5 (e), and IL-13 (f)) in BALF supernatants from HDM-induced mice was analyzed using ELISA. C Transcription factor RNA levels (Foxp3 mRNA, T-bet mRNA and GATA3 mRNA) in the lungs of HDM-induced mice were analyzed using q-PCR. Results were normalized using a GAPDH control and show mean fold increase over control±SEM. The statistical analyses were performed using two-way ANOVAs. *P < 0.05, **P < 0.01, ***P < 0.001 compared with HDM+PBS group, n = 6 mice per group
Discussion
Exosomes are nano-sized membrane vesicles which are released extracellularly after fusion of multivesicular endosomes with the cell membrane. Previous studies suggested that exosomes played an important role in the diagnosis and treatment of various diseases.22,23 For instance, Khan demonstrated mouse embryonic stem cells-derived exosomes slowed down the death of H9C2 cells with hypoxia management, thus promoted the survival and proliferation of myocardial cells.24 Ho et al.25 reported that LRRK2 protein and DJ-1 protein were significantly increased in urine-derived exosomes from Parkinson’s patients, which may provide a new biological indicator for the diagnosis of Parkinson’s disease. Researchers now found exosomes released from different cellular sources might have a role in allergic diseases.26 Antigen-pulsed intestinal epithelial cell-derived exosomes will induce antigen specific tolerance and inhibit specific humoral or cellular immune responses.27 In addition, B cell-derived exosomes could present allergen-derived peptides and stimulated allergen-specific T cells, suggesting that exosomes played a role in allergy.28 Furthermore, it has been suggested that exosomes from mast cells might regulate the allergic immune response by the restoration of Th1/Th2 balance, through an induction of the Th1-type response.29 And serum containing exosomes from OVA-fed experimental allergic animals could induce immunoregulation to OVA through induction of regulatory CD25+ T cells, thus had a protective effect against hypersensitivity reaction.30 Recent studies have shown that bacteria-derived exosomes also had immunoregulatory effects. It was reported that enteropathogenic bacteria-secreted particles communicated with the established mammalian gut community and regulated gut immune homeostasis.31,32 In addition, mycobacterium tuberculosis membrane vesicles modulated the effector functions of infected macrophages and also circulated beyond the site of infection to further regulate immune responses.32
According to the hygiene hypothesis, exposure to LPS in established asthmatics may trigger exacerbations.33 However, pretreatment with endotoxins before the onset of asthma prevents subsequent asthma development.13 Our previous study also found that low-dose LPS prevented the development of allergic sensitization in an animal model of allergic asthma (Supplemental Figure S1 and S2). Therefore, we speculate that exosomes mediate the prophylactic and protective effects of bacteria on asthma sensitization. In our study, we succeeded in isolating exosomes from P. aeruginosa, and the immunomodulatory capacity of exosomes was tested in the OVA or HDM-induced mouse model. We found that P. aeruginosa-derived exosome pretreatment reduced AHR, peribronchial and perivascular inflammation, and serum OVA or HDM-specific IgE level in our two different experimental asthma models. Therefore, mice pretreated with P. aeruginosa-derived exosomes might protect against subsequent sensitization with OVA and HDM. Further, we explored the underlying mechanism of how P. aeruginosa-derived exosomes protected OVA or HDM-induced asthma models. Our results indicated that the P. aeruginosa-derived exosomes may have an immune regulatory role as they inhibit Th2 responses (CD3+ CD4+ IL-4+ T cells) and induce Treg cells (Foxp3+ CD4+ CD25+ T cells), which resulted in the increased secretion of TGF-β, IL-10 in OVA/HDM sensitized mice, decreased production of IL-5 and IL-13 during OVA sensitized mice, and decreased IL-4, IL-5, and IL-13 in HDM sensitized mice.
We speculate that exosomes play an immunomodulatory role in two ways. One speculation may be that exosomes transfer as an antigen between immune cells, contributing to the amplification of Treg responses thus balance the immunoregulation.6 On the other hand, the release of LPS from exosomes may be a mechanism by which the exosomes protect against OVA or HDM-induced asthma. We detected exosomes produced by P. aeruginosa as a vehicle of LPS. A recent study also demonstrated that outer membrane vesicles secreted from Gram-negative bacteria delivered LPS to the host cells.15 However, no matter how the P. aeruginosa-derived exosomes works as an immune modulating role, our research suggests that exosomes derived from Gram-negative bacteria showed an immunomodulatory effect on T-cells immune balance in OVA or HDM-induced asthma, providing new insights for the prevention and treatment of allergic diseases. Encouragingly, exosomes are natural Ag-transferring units between immune cells, allowing cross-presentation and contributing to amplify immune responses. These vesicles are rather stable and reduce the dose of Ag required to induce an immune response. Moreover, exosomes display technology permits manipulation of their protein composition and tailoring for different functions.34 Taken together, exosomes are natural biocompatible carriers and are more likely to be clinically used to prevent autoimmune diseases than other currently available synthetic drugs.
In conclusion, our results revealed that exosomes released from P. aeruginosa have an important role in immunoregulation through up-regulation of Treg responses and down-regulation of Th2 responses during an allergic immunoresponse. The characterization of P. aeruginosa-derived exosomes in allergic disease provides a new insight on the prevention and treatment of allergy diseases. Further studies need to clarify the exact mechanisms of how P. aeruginosa-derived exosomes influence the allergic reaction and different molecules in the exosomes that at work, which will lead to the development of exosomes for use in preventation and treatment of diseases.
Change history
25 February 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41390-020-0823-8
References
Clark, N. M. Community-based approaches to controlling childhood asthma. Annu. Rev. Public Health 33, 193–208 (2012).
Wenzel, S. E. & Busse, W. W. Severe asthma: lessons from the Severe Asthma Research Program. J. Allergy Clin. Immunol. 119, 14–21 (2007).
Lötvall, J. et al. Asthma endotypes: a new approach to classification of disease entities within the asthma syndrome. J. Allergy Clin. Immunol. 127, 355–360 (2011).
Mazzarella, G., Bianco, A., Catena, E., De Palma, R. & Abbate, G. F. Th1/Th2 lymphocyte polarization in asthma. Allergy 55, 6–9 (2000). Suppl 61.
Ray, A., Khare, A., Krishnamoorthy, N., Qi, Z. & Ray, P. Regulatory T cells in many flavors control asthma. Mucosal Immunol. 3, 216–229 (2010).
Hartl, D. et al. Quantitative and functional impairment of pulmonary CD4+ CD25hi regulatory T cells in pediatric asthma. J. Allergy Clin. Immunol. 119, 1258–1266 (2007).
Robbins, P. D. & Morelli, A. E. Regulation of immune responses by extracellular vesicles. Nat. Rev. Immunol. 14, 195–208 (2014).
Chutkan, H., Macdonald, I., Manning, A. & Kuehn, M. J. Quantitative and qualitative preparations of bacterial outer membrane vesicles. Methods Mol. Biol. 966, 259–272 (2013).
Hough, K. P., Chanda, D., Duncan, S. R., Thannickal, V. J. & Deshane, J. S. Exosomes in immunoregulation of chronic lung diseases. Allergy 72, 534–544 (2017).
Okoye, I. S. et al. MicroRNA-containing T-regulatory-cell-derived exosomes suppress pathogenic T helper 1 cells. Immunity 41, 89–103 (2014).
Wakkach, A. et al. Characterization of dendritic cells that induce tolerance and T regulatory 1 cell differentiation in vivo. Immunity 18, 605–617 (2003).
Mutwiri, G., van Drunen Littel-van den Hurk, S. & Babiuk, L. A. Approaches to enhancing immune responses stimulated by CpG oligodeoxynucleotides. Adv. Drug. Deliv. Rev. 61, 226–232 (2009).
Schuijs, M. J. et al. Farm dust and endotoxin protect against allergy through A20 induction in lung epithelial cells. Science 349, 1106–1110 (2015).
Stein, M. M. et al. Innate immunity and asthma risk in Amish And Hutterite farm children. N. Engl. J. Med. 375, 411–421 (2016).
Vanaja, S. K. et al. Bacterial outer membrane vesicles mediate cytosolic localization of LPS and Caspase-11 Activation. Cell 165, 1106–1119 (2016).
Choi, D. S. et al. Proteomic analysis of outer membrane vesicles derived from Pseudomonas aeruginosa. Proteomics 11, 3424–3429 (2011).
Chałupniak, A. et al. Application of quartz tuning forks for detection of endotoxins and Gram-negative bacterial cells by monitoring of Limulus Amebocyte Lysate coagulation. Biosens. Bioelectron 58, 132–137 (2014).
Niu, C. et al. Vitamin A maintains the airway epithelium in a murine model of asthma by suppressing glucocorticoid-induced leucine zipper. Clin. Exp. Allergy 46, 848–860 (2016).
Johnson, J. R. et al. Divergent immune responses to house dust mite lead to distinct structural-functional phenotypes. Am. J. Physiol. Lung. Cell Mol. Physiol. 293, L730–L739 (2007).
Théry, C., Amigorena, S., Raposo, G. & Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell. Biol. 3, 22 (2006).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Kamerkar, S. et al. Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature 546, 498–503 (2017).
Sharma, R., Huang, X., Brekken, R. A. & Schroit, A. J. Detection of phosphatidylserine-positive exosomes for the diagnosis of early-stage malignancies. Br. J. Cancer 117, 545–552 (2017).
Khan, M. et al. Embryonic stem cell-derived exosomes promote endogenous repair mechanisms and enhance cardiac function following myocardial infarction. Circ. Res. 117, 52–64 (2015).
Ho, D. H., Yi, S., Seo, H., Son, I. & Seol, W. Increased DJ-1 in urine exosome of Korean males with Parkinson’s disease. Biomed. Res. Int. 2014, 704678 (2014).
Admyre, C. et al. Exosomes - nanovesicles with possible roles in allergic inflammation. Allergy 63, 404–408 (2008).
Van Niel, G. et al. Intestinal epithelial exosomes carry MHC class II/peptides able to inform the immune system in mice. Gut 52, 1690–1697 (2003).
Admyre, C. et al. B cell-derived exosomes can present allergen peptides and activate allergen-specific T cells to proliferate and produce TH2-like cytokines. J. Allergy Clin. Immunol. 120, 1418–1424 (2007).
Skokos, D. et al. Mast cell-dependent B and T lymphocyte activation is mediated by the secretion of immunologically active exosomes. J. Immunol. 166, 868–876 (2001).
Karlsson, M. R., Kahu, H., Hanson, L. A., Telemo, E. & Dahlgren, U. I. An established immune response against ovalbumin is suppressed by a transferable serum factor produced after ovalbumin feeding: a role of CD25+ regulatory cells. Scand. J. Immunol. 55, 470–477 (2002).
Deng, Z. et al. Enterobacteria-secreted particles induce production of exosome-like S1P-containing particles by intestinal epithelium to drive Th17-mediated tumorigenesis. Nat. Commun. 6, 6956 (2015).
Kovacikova, Z., Neumannova, K., Rydlova, J., Bizovská, L. & Janura, M. The effect of balance training intervention on postural stability in children with asthma. J. Asthma 55, 502–510 (2017). 0.
Lowe, A. P. P. et al. Route of administration affects corticosteroid sensitivity of a combined ovalbumin and lipopolysaccharide model of asthma exacerbation in guinea pigs. J. Pharmacol. Exp. Ther. 362, 327–337 (2017).
Prado, N. et al. Exosomes from bronchoalveolar fluid of tolerized mice prevent allergic reaction. J. Immunol. 181, 1519–1525 (2008).
Acknowledgements
This work was supported by the Program for Innovative Research Team at Chongqing University, 2013.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Ding, FX., Liu, B., Zou, WJ. et al. Pseudomonas aeruginosa-derived exosomes ameliorates allergic reactions via inducing the Treg response in asthma. Pediatr Res 84, 125–133 (2018). https://doi.org/10.1038/s41390-018-0020-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-018-0020-1
This article is cited by
-
Pseudomonas aeruginosa outer membrane vesicles ameliorates lung ischemia–reperfusion injury by regulating the balance of regulatory T cells and Th17 cells through Tim-3 and TLR4/NF-κB pathway
Inflammation Research (2021)
-
Extracellular vesicles: novel communicators in lung diseases
Respiratory Research (2020)