Abstract
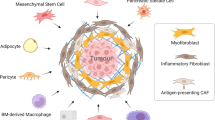
Pancreatic ductal adenocarcinoma (PDAC) is characterized by an extensive fibroinflammatory stroma and often experiences conditions of insufficient oxygen availability or hypoxia. Cancer-associated fibroblasts (CAF) are a predominant and heterogeneous population of stromal cells within the pancreatic tumor microenvironment. Here, we uncover a previously unrecognized role for hypoxia in driving an inflammatory phenotype in PDAC CAFs. We identify hypoxia as a strong inducer of tumor IL1ɑ expression, which is required for inflammatory CAF (iCAF) formation. Notably, iCAFs preferentially reside in hypoxic regions of PDAC. Our data implicate hypoxia as a critical regulator of CAF heterogeneity in PDAC.
Similar content being viewed by others
Introduction
Pancreatic ductal adenocarcinoma (PDAC) remains a deadly disease, with a 5-year survival rate of 11% [1]. A notable feature of PDAC is the presence of an abundant fibroinflammatory stroma that includes extracellular matrix (ECM), cancer-associated fibroblasts (CAF), and immune cells [2]. Recently, single-cell RNA sequencing (scRNA-seq) and other approaches have revealed transcriptionally and functionally distinct CAF subpopulations, myofibroblastic CAFs (myCAF), inflammatory CAFs (iCAF), and antigen-presenting CAFs (apCAF) [3,4,5,6,7]. The myCAF subset is involved in the production of ECM, whereas the iCAF subtype produces high levels of inflammatory cytokines and chemokines [7, 8]. The apCAF population is characterized by MHC class II expression [3]. Previous studies suggested a tumor-restrictive role for myCAFs and a tumor-promoting role for iCAFs and demonstrated that these subpopulations have the potential to interconvert [3, 6,7,8,9,10]. Mechanisms underlying CAF heterogeneity and plasticity as well as different roles of individual CAF subsets in pancreatic tumorigenesis are only beginning to be understood.
Hypoxia, or oxygen (O2) deprivation, occurs in solid tumors, including PDAC, because of their high oxygen/nutrient demand and aberrant vascularization [11,12,13]. Tumor hypoxia induces adaptive changes in cancer cells and surrounding stromal cells, and is associated with cancer progression and therapy resistance [14, 15]. Although hypoxia has been shown to promote fibrosis and angiogenesis by stimulating fibroblasts [16,17,18], the relationship between hypoxia and the recently defined CAF subsets in PDAC is unknown.
Here, we show that iCAFs are preferentially located in hypoxic regions of mouse PDAC in vivo and that the hypoxia-related gene signature is positively enriched in iCAFs in human PDAC samples. Using three-dimensional (3D) cocultures of pancreatic cancer cells and fibroblasts, we demonstrate that hypoxia promotes an iCAF state. Our study identifies hypoxia as a key environmental cue for inducing an iCAF phenotype, thus highlighting an instructive role of hypoxia in shaping the stromal microenvironment.
Results
CAF subtype proportions differ between normoxic and hypoxic tumor microenvironments
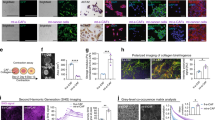
We and others have shown that there is considerable intratumoral heterogeneity of hypoxia in human and mouse PDAC tumors [19, 20]. To identify cells residing in hypoxic tumor areas in vivo, we injected Hypoxyprobe, an indicator of pO2 levels ≤ 1% [21], intraperitoneally into mice bearing orthotopic PDAC. In this model, pancreatic cancer cells derived from the KrasLSL-G12D/+;Trp53LSL-R172H/+;Pdx1-Cre (KPC) mouse model of PDAC [22] were injected into the pancreas of syngeneic C57/BL6 mice. Immunofluorescence staining for Hypoxyprobe in orthotopic PDAC showed patchy patterns of hypoxia (Fig. 1A), similar to those observed in human PDAC samples [19]. As expected, Hypoxyprobe was barely detectable in the normal pancreas (Fig. S1A). The average percentage of hypoxic cells in pancreatic tumors (defined as % Hypoxyprobe+ cells of total live cells) was 28% (Fig. 1B). One-third of total PDPN+ CAFs stained positively for Hypoxyprobe (Fig. 1C, D). Using a previously validated flow cytometry strategy for CAF subtypes [3, 6], we evaluated myCAFs (PDPN+Ly6C−MHCII−), iCAFs (PDPN+Ly6C+MHCII−), and apCAFs (PDPN+Ly6C−MHCII+) located within either hypoxic (Hypoxyprobe+) or normoxic (Hypoxyprobe−) tumor regions (Fig. 1E). Importantly, the distributions of CAF subpopulations from normoxic and hypoxic tumor microenvironments significantly differed (Fig. 1E–J). Although myCAFs were the prevalent CAF subset in both normoxic and hypoxic tumor regions (Fig. 1F, J), hypoxic areas contained a significantly higher fraction of iCAFs compared with normoxic areas (Fig. 1G, J) and exhibited pronounced increases in the iCAF/myCAF ratio (Fig. 1H). We further confirmed the preferential localization of iCAFs in hypoxic regions in orthotopic tumors using a different KPC PDAC cell line, 4662 (Fig. S1B).
A–J Mice bearing 4-week orthotopic PDAC of mT3 tumor cells received an intraperitoneal injection with 60 mg/kg of Hypoxyprobe and were sacrificed 1.5–2 h later. A Immunofluorescence staining for Hypoxyprobe (green) and DAPI (blue) in orthotopic PDAC. Scale bar, 100 µm. B Percentage of Hypoxyprobe+ cells among total live cells from orthotopic PDAC, as analyzed by flow cytometry (n = 6). C Co-immunofluorescence staining for Hypoxyprobe (green), PDPN (red), and DAPI (blue) in orthotopic PDAC. Scale bar, 25 µm. D Percentage of Hypoxyprobe+ CAFs among total CAFs from orthotopic PDAC, as analyzed by flow cytometry (n = 6). E Schematic of flow cytometry strategy to identify CAF subsets residing in normoxic and hypoxic tumor regions. Representative flow plots showing the gating strategy for the analysis of normoxic (Hypoxyprobe−) and hypoxic (Hypoxyprobe+) CAF subsets from orthotopic PDAC. F–I Percentage of myCAFs (F), percentage of iCAFs (G), iCAF/myCAF ratio (H), and percentage of apCAFs (I) among normoxic and hypoxic CAFs from orthotopic PDAC, as analyzed by flow cytometry (n = 6). J Pie charts showing mean frequencies of the indicated subsets among normoxic and hypoxic CAFs from orthotopic PDAC, as analyzed by flow cytometry (n = 6). K Uniform manifold approximation and projection (UMAP) visualization of fibroblast clusters from human PDAC scRNA-seq (n = 16 patients merged). Different CAF subtype clusters are color-coded. Data are from Steele and colleagues [23], and annotations are from Kemp and colleagues [24]. L UMAP visualization of human PDAC fibroblasts from K colored by hypoxia gene set expression score. The hypoxia signature for analysis was obtained from MSigDB’s Hallmark collection. Red, highest score of hypoxia signature; blue, lowest score of hypoxia signature. The symbols in B, D, F–I represent individual mice, and horizontal lines represent the means. P-values were determined by student’s t-test. *p < 0.05; **p < 0.01; ***p < 0.001.
To address the relevance of the correlation between hypoxia and an iCAF phenotype in human PDAC, we interrogated the expression profiles of iCAFs and myCAFs from a scRNA-seq dataset [23] that includes 16 PDAC patient tumor samples. Populations of myCAFs and iCAFs were both present in this dataset, with the majority of fibroblasts falling into the myCAF group (Fig. 1K, as annotated in ref. [24]). The expression profiles of each cell were then scored using the Hallmark Hypoxia gene set (MSigDB) as a readout of exposure to hypoxia. We found that most iCAFs exhibit a robust hypoxia profile (76%, 177 of 234 total cells scored above the median of signature expression), while only a subset of myCAFs met this threshold (45%, 565 of 1250 total cells scored above the median of signature expression) (Fig. 1L, Fig. S1C). Compared with myCAFs, iCAFs were also enriched for the hypoxia-inducible factor 1 (HIF1) signaling pathway, a key hypoxic adaptation pathway (Fig. S1D). These observations suggest that the iCAF phenotype is linked with the hypoxic tumor microenvironment of PDAC.
Hypoxia promotes the induction of an inflammatory phenotype in CAFs by modulating their interactions with tumor cells
Based on the correlation between PDAC hypoxia and iCAF enrichment, we set out to determine whether hypoxia regulates an iCAF phenotype. When pancreatic stellate cells (PSCs), a precursor population of CAFs, are seeded in Matrigel in a transwell insert and cultured with PDAC tumor organoids in the lower compartment of the plate, they acquire the inflammatory features characteristic of iCAFs [7, 8]. On the other hand, PSCs cultured alone in Matrigel maintain a quiescent state [7, 8]. To examine the effects of hypoxia on an iCAF phenotype, we exposed the cocultures of mouse PDAC tumor organoids and PSCs to either normoxia (21% O2) or hypoxia (1% O2) (Fig. 2A) and measured the expression of CAF subset markers in PSCs. To model hypoxia in vitro, 1% O2 was used as the level of O2 in human pancreatic cancers measured by a polarographic electrode is ~1% [11]. Expression of the iCAF markers Il6, Cxcl1, and Lif was markedly elevated in hypoxic PSCs cocultured with tumor organoids relative to their normoxic counterparts (Fig. 2B, C). On the other hand, expression of the myCAF markers Acta2 (α-SMA gene) and Transgelin (Tagln) in PSCs cocultured with tumor organoids was not affected by hypoxia (Fig. 2D). Importantly, hypoxic induction of the iCAF markers in PSCs only occurred when cocultured with tumor organoids but not when cultured alone (Fig. 2B). These observations were reproduced in the cocultures of tumor cells and a CAF line derived from mouse PDAC (Fig. S2A, B). In addition, hypoxia increased the expression of iCAF marker genes in PSCs cocultured with another PDAC cell line, 4662 (Fig. S2C), suggesting that hypoxic induction of the iCAF phenotype is not limited to a specific cell line. Of note, in the context of 2D cocultures, hypoxia elevated the expression of Il6 and Cxcl1 in PSCs (P = 0.08 for Cxcl1 between normoxia and hypoxia) but not of Lif (Fig. S2D), implicating partial induction of the iCAF phenotype by hypoxia in 2D culture.
A Schematic illustration of the 3D coculture platform to model tumor cell-PSC interactions under normoxia and hypoxia. B Quantitative RT-PCR analysis of iCAF markers in PSCs cultured alone or with mT3 tumor organoids under 21% O2 or 1% O2 for 48 h (n = 6). Expression levels were normalized by 18S rRNA. C Enzyme-linked immunosorbent assay (ELISA) of iCAF markers in conditioned media from 3D cocultures of PSCs and mT3 tumor cells under 21% O2 or 1% O2 for 72 h (n = 3). D Quantitative RT-PCR analysis of myCAF markers in PSCs cultured alone or with mT3 tumor organoids under 21% O2 or 1% O2 for 48 h (n = 6). Expression levels were normalized by 18S rRNA. E Gene set enrichment analysis (GSEA) showing significantly upregulated pathways in PSCs cultured with mT3 tumor organoids at 1% O2 compared with PSCs cultured with mT3 tumor organoids at 21% O2 for 48 h. NES normalized enrichment score, FDR false discovery rate. Each data point in B–D represents individual primary PSC lines. Data in B–D, mean ± SEM. P-values were determined by two-way ANOVA with Bonferroni post-test (B, D) and student’s t-test (C). NS not significant. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
To assess the effects of hypoxia on fibroblast phenotype in an unbiased fashion, we performed RNA-seq profiling of the hypoxic and normoxic PSCs cocultured with PDAC organoids. When using gene set enrichment analysis (GSEA), we found that “inflammatory response” and “IL6/JAK/STAT3”, in addition to the “hypoxia signature”, are top-ranked in association with the hypoxic PSCs (Fig. 2E, Fig. S2E). Altogether, these results indicate that hypoxia promotes an iCAF state and that the induction of an inflammatory fibroblast phenotype by hypoxia requires factors secreted by tumor cells.
Hypoxic regulation of the iCAF phenotype is independent of tumor HIF1ɑ or HIF2ɑ
Cellular adaptation to hypoxia is largely coordinated by hypoxia-inducible factors (HIFs) [25]. It has been shown that the major HIF isoforms, HIF1ɑ and HIF2ɑ, are expressed in human and mouse PDAC and play distinct roles in pancreatic tumorigenesis [20, 26,27,28]. Because hypoxic tumor cells are needed to establish the iCAF phenotype (Fig. 2B), we postulated that HIF1ɑ or HIF2ɑ in tumor cells may contribute to iCAF formation under hypoxia. To test this hypothesis, we knocked down HIF1ɑ, HIF2ɑ, or both in PDAC tumor cells using shRNAs (Fig. S3A, B) and cultured these tumor cells with PSCs under normoxia (21% O2) or hypoxia (1% O2). Unexpectedly, neither HIF1ɑ knockdown nor HIF2ɑ knockdown impaired induction of iCAF marker genes Il6, Cxcl1, and Lif in PSCs by hypoxia (Fig. 3A, B). Double knockdown of HIF1ɑ and HIF2ɑ abrogated the induction of Cxcl1 expression in PSCs exposed to hypoxia, but was not sufficient to fully prevent hypoxic induction of Il6 and Lif expression in PSCs (Fig. 3C). Moreover, knockdown of HIF1ɑ, HIF2ɑ, or in combination in another PDAC cell line 4662 failed to inhibit expression of Il6, Cxcl1, and Lif in PSCs under hypoxia (Fig. S3C–E). These data suggest that tumor HIF1ɑ and HIF2ɑ are largely dispensable for hypoxia-mediated iCAF formation.
Quantitative RT-PCR analysis of iCAF markers in PSCs cultured with mT3 tumor organoids expressing scrambled shRNA (shSCR) control, HIF1ɑ shRNA (shHIF1ɑ) (A), HIF2ɑ shRNA (shHIF2ɑ) (B), or both shHIF1ɑ and shHIF2ɑ (C) under 21% O2 or 1% O2 for 48 h (n = 4 shHIF1ɑ, n = 3 shHIF2ɑ, n = 5 shHIF1ɑ/shHIF2ɑ). Expression levels were normalized by 18S rRNA. Each data point represents individual primary PSC lines. Results show mean ± SEM. P-values were determined by two-way ANOVA with Bonferroni post-test. NS not significant. *p < 0.05; **p < 0.01; ***p < 0.001.
IL1ɑ in tumor cells mediates hypoxic induction of the iCAF phenotype
IL1ɑ secreted by pancreatic tumor cells and subsequent IL6/JAK/STAT3 activation in CAFs have been shown to trigger iCAF formation [7]. However, the mechanism underlying IL1ɑ induction in cancer cells has remained obscure. Because iCAF induction by hypoxia requires tumor cells, we measured IL1ɑ expression from tumor organoids exposed to either normoxia (21% O2) or hypoxia (1% O2). Hypoxia increased Il1ɑ mRNA levels in pancreatic cancer cells (Fig. 4A). IL1ɑ protein levels were also elevated in conditioned media from hypoxic cocultures relative to conditioned media from normoxic cocultures (Fig. 4B). Of note, although hypoxia significantly upregulated Il1ɑ expression in cancer cells cultured alone, an increase in Il1ɑ expression in cancer cells by hypoxia was even greater in the presence of PSCs (Fig. 4A), implicating bi-directional interactions between tumor cells and PSCs. Moreover, targeting IL1ɑ with a neutralizing antibody substantially reduced induction of iCAF marker genes Il6, Cxcl1, and Lif in PSCs but not myCAF marker gene Acta2 under hypoxia (Fig. 4C, Fig. S4A). To confirm the importance of IL1 signaling in iCAF formation, we treated cancer cell-PSC cocultures with an IL1 receptor (IL1R1)-neutralizing antibody, which resulted in the impaired acquisition of the iCAF phenotype under hypoxia (Fig. 4D). Consistent with a tumor HIF-independent mechanism for the hypoxic induction of iCAF formation, knockdown of HIF1ɑ, HIF2ɑ, or both did not affect upregulation of Il1ɑ in cancer cells in response to hypoxia (Fig. S4B–D). Collectively, our findings suggest that hypoxia induces IL1ɑ expression in tumor cells and that IL1ɑ is crucial for hypoxia-mediated iCAF formation.
A Quantitative RT-PCR analysis of Il1ɑ in mT3 tumor organoids cultured alone or with PSCs under 21% O2 or 1% O2 for 48 h (n = 6). Expression levels were normalized by 18S rRNA. B ELISA of IL1ɑ in conditioned media from 3D cocultures of mT3 tumor cells and PSCs under 21% O2 or 1% O2 for 72 h (n = 4). C Quantitative RT-PCR analysis of iCAF markers in PSCs cultured with mT3 tumor organoids in the presence of IL1ɑ-neutralizing antibody or isotype control antibody under 21% O2 or 1% O2 for 72 h (n = 5). Expression levels were normalized by 18S rRNA. D Quantitative RT-PCR analysis of iCAF markers in PSCs cultured with mT3 tumor organoids in the presence of IL1R1-neutralizing antibody or isotype control antibody under 21% O2 or 1% O2 for 72 h (n = 3). Expression levels were normalized by 18S rRNA. Each data point represents individual primary PSC lines. Results show mean ± SEM. P-values were determined by Mann–Whitney test with Bonferroni post-test (A), Mann–Whitney test (B), and two-way ANOVA with Bonferroni post-test (C, D). NS not significant. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
Discussion
Hypoxia is a critical feature of the tumor microenvironment and predicts poor clinical outcomes [14, 15]. The impact of hypoxia on cancer cells has been well-characterized, yet much remains to be understood as to how hypoxia regulates stromal components and the tumor-stroma crosstalk. In this study, we demonstrate that intratumoral normoxic and hypoxic microenvironments differ in CAF composition in mouse PDAC and that iCAFs are linked to tumor hypoxia in human and mouse PDAC. By exposing 3D cocultures of pancreatic cancer cells and fibroblasts to either hypoxia or normoxia, we found that hypoxia promotes an inflammatory phenotype of fibroblasts. In addition, we showed that hypoxic induction of the iCAF phenotype requires IL1ɑ emanating from tumor cells, and the presence of hypoxic fibroblasts further elevates IL1ɑ levels in tumor cells, implicating hypoxia as a modulator of reciprocal interactions between cancer cells and fibroblasts.
HIFs are major mediators of adaptations to hypoxia; however, hypoxic responses can be HIF-independent [25, 29]. For instance, hypoxia regulates the NFκB pathway, which plays a central role in the control of immune responses, through HIF-dependent and HIF-independent mechanisms [30,31,32]. While our data suggest that upregulation of tumor IL1ɑ and iCAF induction by hypoxia are largely independent of HIF activation in tumor cells, the precise mechanism for tumor IL1ɑ regulation by hypoxia remains to be determined.
PSCs have been thought to give rise to the majority of PDAC CAFs. However, recent reports suggest that besides PSCs, PDAC CAFs can arise from multiple cell types [33, 34]. We observed that hypoxia propels an inflammatory phenotype in a PDAC CAF line as well as primary PSCs. It is unclear whether CAF populations of different developmental origins have a differential capacity to gain iCAF features.
A key finding in our current study is that iCAFs are enriched in hypoxic zones of PDAC compared with normoxic tumor regions. This spatial link between hypoxia and iCAF enrichment in vivo, together with our in vitro finding that hypoxia promotes iCAF induction, raises the possibility that hypoxia plays an active role in driving regional stromal heterogeneity. Notably, recent studies have observed a correlation between iCAF enrichment and immunosuppression [6, 10, 35], which warrants the investigation of the effects of hypoxia on CAF-immune crosstalk.
In summary, our study reveals that the normoxic and hypoxic microenvironments of PDAC exhibit distinct CAF compositions. We also show that hypoxia induces an inflammatory fibroblast phenotype through upregulation of tumor IL1ɑ, thus highlighting the significance of hypoxia in shaping the tumor stroma. A better understanding of the impact of hypoxia on CAF heterogeneity and function is needed to make stroma-targeting therapies clinically viable.
Materials and methods
Mice
All animal protocols were reviewed and approved by the Institutional Animal Care and Use Committee of the University of Michigan. Wild-type (WT) C57/BL6 mice (stock # 000664) from Jackson Laboratory were used for PSC isolation and orthotopic transplantation experiments at 8–12 weeks of age, including male and female mice. For orthotopic transplantation, 7.5 × 104 mT3 (provided by Dr. David A. Tuveson) [36] or 7.5 × 104 4662 cells (provided by Robert H. Vonderheide) [37] derived from primary PDAC in KrasLSL-G12D/+; Trp53LSL-R172H/+; Pdx1-Cre (KPC) mice of a C57/BL6 genetic background, were resuspended as a 30 µl suspension of 50% Matrigel (#356231, Corning) in PBS and injected into the pancreas. At 4 weeks post-transplantation, mice received an intraperitoneal injection with 60 mg/kg of Hypoxyprobe (pimonidazole hydrochloride, Hypoxyprobe, Inc) and were sacrificed 1.5–2 h later for flow cytometry analysis.
Cell lines and cell culture
PSCs were isolated from WT mice by enzymatic digestion of pancreatic tissue and subsequent density gradient centrifugation as previously described [8, 38]. Primary PSC lines between passages 2 and 4 were used for all experiments. The FB1 CAF line was generated from an iKras* p53* mouse [39] by fluorescence-activated cell sorting of PDGFRα+;EPCAM− cells. The mT3 (provided by Dr. David A. Tuveson) [36] and 4662 (provided by Robert H. Vonderheide) [37] PDAC cell lines were derived from primary murine KPC PDAC. FB1, mT3, and 4662 cell lines were cultured no more than 20–25 passages. All cells were passaged in DMEM with 10% FBS and 1% penicillin/streptomycin (Thermo Fisher). For 3D cocultures, PSCs were seeded in Matrigel (#356231, Corning) in a transwell insert (#662610, Greiner Bio-One) and cultured with PDAC tumor organoids in the lower compartment of the 24-well plate in DMEM containing 5% FBS and 1% penicillin/streptomycin (Thermo Fisher). For IL1ɑ neutralization experiments, cocultures were treated with 3 µg/ml IL1ɑ-neutralizing antibody (#MAB4001, R&D Systems) or isotype control antibody (#400902, BioLegend) for 72 h. For IL1R1 neutralization experiments, cocultures were treated with 0.5 µg/ml IL1R1-neutralizing antibody (#PA5-47937, Invitrogen) or isotype control antibody (#AB108C, R&D Systems) for 72 h. Cell line authentication for FB1 and mT3 was not performed. The 4662 cells were authenticated by the Research Animal Diagnostic Laboratory (RADIL) at the University of Missouri. Mycoplasma testing (MycoAlert Detection Kit, Lonza) was performed monthly.
Lentiviral-mediated shRNA transduction
PDAC cell lines were transduced with lentivirus containing shRNA plasmids at optimized viral titers. Stable cell lines were established post-puromycin selection. The following shRNA plasmids were used: pGIPZ Scrambled shRNA (#RH4346, Horizon), pGIPZ HIF1α shRNA (#RMM4431-200404026, Horizon), pLKO.1 Scrambled shRNA (#1864, Addgene), pLKO.1 HIF2α shRNA (#TRCN0000082307, Sigma).
Quantitative RT-PCR
Total RNA was isolated from cells using the RNeasy mini kit (#74104, Qiagen). cDNA was synthesized using a High-Capacity cDNA Reverse Transcription Kit (#4368814, Applied Biosystems). PCR reactions were performed using SYBR Green PCR reagents (#A25742, Applied Biosystems) mixed with indicated cDNAs and primers (primer sequences are listed in Table S1) in a QuantStudio Real-Time PCR system (Applied Biosystems). Expression levels were normalized by 18S rRNA.
Immunofluorescence
Tissues were fixed in 4% paraformaldehyde/PBS (4 °C, overnight) and processed for paraffin embedding. For immunofluorescence, slides were boiled for 20 min in 10 mM sodium citrate (pH 6.0) for antigen retrieval and blocked with 5% serum/0.3% Triton X-100 for 1 h. Sections were incubated with FITC-conjugated Hypoxyprobe-1-MAb1 (4.3.11.3, #FITC-Mab, 1:500, Hypoxyprobe, Inc) and Alexa Fluor 594-conjugated PDPN antibody (8.1.1, #127414, 1:250) diluted in 1% BSA/0.3% Triton X-100 overnight at 4 °C. Slides were counterstained with DAPI (Invitrogen) and mounted in Prolong Gold antifade reagent (Invitrogen). Fluorescence images were acquired using an Olympus IX73 microscope.
Flow cytometry
Single-cell suspensions from mouse tissues were prepared as previously described [20]. Tumor tissues were cut in half, and one half was minced and processed for flow cytometry analysis. Cells were stained in PBS/0.5% FBS/2 mM EDTA with the following fluorochrome-conjugated antibodies: BV421-conjugated anti-Ly6C (HK1.4, #128031, 1:100), PerCP-Cy5.5-conjugated anti-CD45 (30-F11, #103132, 1:200), PE-conjugated anti-EPCAM (G8.8, #118205, 1:200), PE-conjugated anti-CD31 (390, #102407, 1:200), PE-Cy7-conjugated anti-PDPN (8.1.1, #127411, 1:100), APC-Cy7-conjugated anti-MHCII (M5/114.15.2, #107628, 1:300) (from BioLegend); FITC-conjugated Hypoxyprobe-1-MAb1 (4.3.11.3, #FITC-Mab, 1:200) (from Hypoxyprobe, Inc). The viability marker Zombie Aqua was purchased from BioLegend (#423102). Flow cytometry was performed on a ZE5 Cell Analyzer (Bio-Rad), and data were analyzed using FlowJo software.
ELISA
For ELISA of media, 3D cocultures were grown under 21% O2 or 1% O2 for 72 h. Media were collected, spun down, and assayed using the manufacturer’s protocol. ELISA assays used were IL1ɑ (#433404, BioLegend), CXCL1 (#EMCXCL1, Invitrogen), IL6 (#DY406-05, R&D Systems), and LIF (#445104, BioLegend).
RNA-seq and data analysis
Total RNA was isolated from cells using the RNeasy mini kit (#74104, Qiagen). Libraries were constructed using NEB polyA RNA ultra II and subsequently subjected to 150 cycles of sequencing on NovaSeq-6000 (Illumina). Adapters were trimmed using Cutadapt (v2.3). FastQC (v0.11.8) was used to ensure the quality of data. Reads were mapped to the mouse genome (GRCm38) using STAR (v2.6.1b) and assigned count estimates to genes with RSEM (v1.3.1). Alignment options followed ENCODE standards for RNA-seq. FastQC was used in an additional post-alignment step to ensure that only high-quality data were used for expression quantitation and differential expression. Differential gene expression analysis was performed using DESeq2, using a negative binomial generalized linear model (thresholds: linear fold change >1.5 or <−1.5, Benjamini–Hochberg FDR (Padj) < 0.05). GSEA was performed using GSEA 4.1.0.
Single-cell RNA-seq analysis
Human single-cell RNA-seq (scRNA-seq) data were previously published in [23], and fibroblasts were annotated in ref. [24]. Both raw and processed data are available at the NIH dbGaP database (accession #phs002071.v1.p1; [23]), with full clinical annotation. Downstream analysis was performed using Seurat V4.0.3 [40]. Hypoxia signature scoring was performed using Seurat’s “AddModuleScore” function.
Western blot analysis
Cells were lysed with 10 mmol/L Tris at pH 7.5, 150 mmol/L NaCl, 5 mmol/L EDTA, 0.1% SDS, and protease/phosphatase inhibitor cocktail (#78440, Thermo Fisher). Cell lysates were separated by SDS-PAGE, transferred to nitrocellulose membranes, blotted with primary antibodies overnight at 4 °C, and detected using horseradish peroxidase-conjugated secondary antibodies followed by exposure to chemiluminescence reagents (#PI34580, Thermo Fisher). The following antibodies were used: rabbit anti-HIF1α (#10006421, 1:500, Cayman), goat anti-HIF2α (#AF2997, 1 µg/ml, R&D Systems), mouse anti-beta actin (#MA1-91399, 1:50,000, Invitrogen), HRP-linked anti-rabbit IgG (#7074, 1:10,000, Cell Signaling), HRP-linked anti-mouse IgG (#7076, 1:60,000, Cell Signaling), and HRP-linked anti-goat IgG (#705035147, 1:20,000, Jackson ImmunoResearch).
Statistical analysis
Data were analyzed using GraphPad Prism 7 software. Statistical tests with normally distributed variables included two-tailed student’s t-test and two-way ANOVA. D’Agostino and Pearson test and/or Shapiro–Wilk test was used to test the normality of sample distribution. When variables were not normally distributed, we performed nonparametric Mann–Whitney test. Bonferroni correction was applied for multiple comparisons. P-value < 0.05 was considered statistically significant. No statistical method was used to predetermine sample sizes, experiments were not randomized, and the investigators were not blinded to allocation during experiments and outcome assessment.
Data availability
The RNA-seq data from this study are available at the Gene Expression Omnibus under the accession number GSE199012.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer Statistics, 2022. CA Cancer J Clin. 2022;72:7–33.
Zhang Y, Crawford HC, Pasca di Magliano M. Epithelial-stromal interactions in pancreatic cancer. Annu Rev Physiol. 2019;81:211–33.
Elyada E, Bolisetty M, Laise P, Flynn WF, Courtois ET, Burkhart RA, et al. Cross-species single-cell analysis of pancreatic ductal adenocarcinoma reveals antigen-presenting cancer-associated fibroblasts. Cancer Discov. 2019;9:1102–23.
Bernard V, Semaan A, Huang J, San Lucas FA, Mulu FC, Stephens BM, et al. Single-cell transcriptomics of pancreatic cancer precursors demonstrates epithelial and microenvironmental heterogeneity as an early event in neoplastic progression. Clin Cancer Res. 2019;25:2194–205.
Dominguez CX, Muller S, Keerthivasan S, Koeppen H, Hung J, Gierke S, et al. Single-cell RNA sequencing reveals stromal evolution into LRRC15(+) myofibroblasts as a determinant of patient response to cancer immunotherapy. Cancer Discov. 2020;10:232–53.
Steele NG, Biffi G, Kemp SB, Zhang Y, Drouillard D, Syu L, et al. Inhibition of Hedgehog signaling alters fibroblast composition in pancreatic cancer. Clin Cancer Res. 2021;27:2023–37.
Biffi G, Oni TE, Spielman B, Hao Y, Elyada E, Park Y, et al. IL1-induced JAK/STAT signaling is antagonized by TGFbeta to shape CAF heterogeneity in pancreatic ductal adenocarcinoma. Cancer Discov. 2019;9:282–301.
Ohlund D, Handly-Santana A, Biffi G, Elyada E, Almeida AS, Ponz-Sarvise M, et al. Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J Exp Med. 2017;214:579–96.
Chen Y, Kim J, Yang S, Wang H, Wu CJ, Sugimoto H, et al. Type I collagen deletion in alphaSMA(+) myofibroblasts augments immune suppression and accelerates progression of pancreatic cancer. Cancer Cell. 2021;39:548–65. e546
Lee BY, Hogg EKJ, Below CR, Kononov A, Blanco-Gomez A, Heider F, et al. Heterocellular OSM-OSMR signalling reprograms fibroblasts to promote pancreatic cancer growth and metastasis. Nat Commun. 2021;12:7336.
Koong AC, Mehta VK, Le QT, Fisher GA, Terris DJ, Brown JM, et al. Pancreatic tumors show high levels of hypoxia. Int J Radiat Oncol Biol Phys. 2000;48:919–22.
Olive KP, Jacobetz MA, Davidson CJ, Gopinathan A, McIntyre D, Honess D, et al. Inhibition of Hedgehog signaling enhances delivery of chemotherapy in a mouse model of pancreatic cancer. Science 2009;324:1457–61.
Kahn BM, Lucas A, Alur RG, Wengyn MD, Schwartz GW, Li J, et al. The vascular landscape of human cancer. J Clin Invest. 2021;131:e136655.
Bejarano L, Jordao MJC, Joyce JA. Therapeutic targeting of the tumor microenvironment. Cancer Discov. 2021;11:933–59.
Palazon A, Goldrath AW, Nizet V, Johnson RS. HIF transcription factors, inflammation, and immunity. Immunity 2014;41:518–28.
Masamune A, Kikuta K, Watanabe T, Satoh K, Hirota M, Shimosegawa T. Hypoxia stimulates pancreatic stellate cells to induce fibrosis and angiogenesis in pancreatic cancer. Am J Physiol Gastrointest Liver Physiol. 2008;295:G709–717.
Spivak-Kroizman TR, Hostetter G, Posner R, Aziz M, Hu C, Demeure MJ, et al. Hypoxia triggers hedgehog-mediated tumor-stromal interactions in pancreatic cancer. Cancer Res. 2013;73:3235–47.
Erkan M, Reiser-Erkan C, Michalski CW, Deucker S, Sauliunaite D, Streit S, et al. Cancer-stellate cell interactions perpetuate the hypoxia-fibrosis cycle in pancreatic ductal adenocarcinoma. Neoplasia 2009;11:497–508.
Dhani NC, Serra S, Pintilie M, Schwock J, Xu J, Gallinger S, et al. Analysis of the intra- and intertumoral heterogeneity of hypoxia in pancreatic cancer patients receiving the nitroimidazole tracer pimonidazole. Br J Cancer. 2015;113:864–71.
Lee KE, Spata M, Bayne LJ, Buza EL, Durham AC, Allman D, et al. Hif1a deletion reveals pro-neoplastic function of B cells in pancreatic neoplasia. Cancer Discov. 2016;6:256–69.
Raleigh JA, Chou SC, Arteel GE, Horsman MR. Comparisons among pimonidazole binding, oxygen electrode measurements, and radiation response in C3H mouse tumors. Radiat Res. 1999;151:580–9.
Hingorani SR, Wang L, Multani AS, Combs C, Deramaudt TB, Hruban RH, et al. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell. 2005;7:469–83.
Steele NG, Carpenter ES, Kemp SB, Sirihorachai V, The S, Delrosario L, et al. Multimodal mapping of the tumor and peripheral blood immune landscape in human pancreatic cancer. Nat Cancer. 2020;1:1097–112.
Kemp SB, Carpenter ES, Steele NG, Donahue KL, Nwosu ZC, Pacheco A, et al. Apolipoprotein E promotes immune suppression in pancreatic cancer through NF-kappaB-mediated production of CXCL1. Cancer Res. 2021;81:4305–18.
Lee KE, Simon MC. SnapShot: hypoxia-inducible factors. Cell 2015;163:1288–1288. e1281
Schofield HK, Tandon M, Park MJ, Halbrook CJ, Ramakrishnan SK, Kim EC, et al. Pancreatic HIF2alpha stabilization leads to chronic pancreatitis and predisposes to mucinous cystic neoplasm. Cell Mol Gastroenterol Hepatol. 2018;5:169–85.
Criscimanna A, Duan LJ, Rhodes JA, Fendrich V, Wickline E, Hartman DJ, et al. PanIN-specific regulation of Wnt signaling by HIF2α during early pancreatic tumorigenesis. Cancer Res. 2013;73:4781–90.
Tiwari A, Tashiro K, Dixit A, Soni A, Vogel K, Hall B, et al. Loss of HIF1A from pancreatic cancer cells increases expression of PPP1R1B and degradation of p53 to promote invasion and metastasis. Gastroenterology 2020;159:1882–97.
Lee P, Chandel NS, Simon MC. Cellular adaptation to hypoxia through hypoxia inducible factors and beyond. Nat Rev Mol Cell Biol. 2020;21:268–83.
Rius J, Guma M, Schachtrup C, Akassoglou K, Zinkernagel AS, Nizet V, et al. NF-kappaB links innate immunity to the hypoxic response through transcriptional regulation of HIF-1alpha. Nature 2008;453:807–11.
Cummins EP, Berra E, Comerford KM, Ginouves A, Fitzgerald KT, Seeballuck F, et al. Prolyl hydroxylase-1 negatively regulates IkappaB kinase-beta, giving insight into hypoxia-induced NFkappaB activity. Proc Natl Acad Sci USA. 2006;103:18154–9.
Walmsley SR, Print C, Farahi N, Peyssonnaux C, Johnson RS, Cramer T, et al. Hypoxia-induced neutrophil survival is mediated by HIF-1alpha-dependent NF-kappaB activity. J Exp Med. 2005;201:105–15.
Garcia PE, Adoumie M, Kim EC, Zhang Y, Scales MK, El-Tawil YS, et al. Differential contribution of pancreatic fibroblast subsets to the pancreatic cancer stroma. Cell Mol Gastroenterol Hepatol. 2020;10:581–99.
Helms EJ, Berry MW, Chaw RC, DuFort CC, Sun D, Onate MK, et al. Mesenchymal lineage heterogeneity underlies nonredundant functions of pancreatic cancer-associated fibroblasts. Cancer Discov. 2022;12:484–501.
Somerville TD, Biffi G, Dassler-Plenker J, Hur SK, He XY, Vance KE, et al. Squamous trans-differentiation of pancreatic cancer cells promotes stromal inflammation. elife 2020;9:e53381.
Boj SF, Hwang CI, Baker LA, Chio II, Engle DD, Corbo V, et al. Organoid models of human and mouse ductal pancreatic cancer. Cell 2015;160:324–38.
Evans RA, Diamond MS, Rech AJ, Chao T, Richardson MW, Lin JH, et al. Lack of immunoediting in murine pancreatic cancer reversed with neoantigen. JCI Insight. 2016;1:e88328.
Apte MV, Haber PS, Applegate TL, Norton ID, McCaughan GW, Korsten MA, et al. Periacinar stellate shaped cells in rat pancreas: identification, isolation, and culture. Gut. 1998;43:128–33.
Collins MA, Brisset JC, Zhang Y, Bednar F, Pierre J, Heist KA, et al. Metastatic pancreatic cancer is dependent on oncogenic Kras in mice. PLoS ONE. 2012;7:e49707.
Hao Y, Hao S, Andersen-Nissen E, Mauck WM 3rd, Zheng S, Butler A, et al. Integrated analysis of multimodal single-cell data. Cell 2021;184:3573–87. e3529
Acknowledgements
We are grateful to Dr. D. A. Tuveson for providing the mT3 PDAC cell line. The 4662 PDAC cell line was a generous gift from Dr. R. H. Vonderheide. We acknowledge support from the Flow Cytometry Core, the Advanced Genomics Core, and the Bioinformatics Core of the University of Michigan Medical School’s Biomedical Research Core Facilities.
Funding
This project was supported by DOD Peer Reviewed Cancer Research Program grant W81XWH-20-1-0629, Concern Foundation Conquer Cancer Now Award, and Swim Across America Young Investigator Award to KEL. AM was supported by NIH grant T32AI007413. TN was supported by NIH grant T32GM140223. KLD was supported by NIH grant T32CA009676. AR was funded by NIH grant R37CA214955, the American Cancer Society RSG-005-016, MIDAS PODS funding, and institutional startup fund from the University of Michigan. ESC was funded by American College of Gastroenterology Junior Faculty Development Award and VA Career Development Award BLR&D. HCC was supported by NIH grant R01CA247516. MPdM was supported by NIH grants R01CA151588 and R01CA198074, and the American Cancer Society. JL, HCC, and MPdM were supported by the University of Michigan Cancer Center Support Grant P30CA046592. HCC and MPdM were supported by NIH grant U01CA224145.
Author information
Authors and Affiliations
Contributions
Conceptualization: KEL; Methodology: AM, TN, YL, KLD, ESC, HCC, MPdM, and KEL; Validation: AM, TN, and KEL; Formal analysis: AM, TN, KLD, JL, ESC, and KEL; Investigation: AM, TN, YL, and KEL; Resources: KLD, AR, HCC, MPdM, and KEL; Data curation: AM, TN, KLD, JL, ESC, and KEL; Visualization: AM, TN, KLD, JL, ESC, and KEL; Writing—original draft: KEL; Writing—review and editing: AM, TN, KLD, MPdM, and KEL; Funding acquisition: KEL; Supervision: KEL.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mello, A.M., Ngodup, T., Lee, Y. et al. Hypoxia promotes an inflammatory phenotype of fibroblasts in pancreatic cancer. Oncogenesis 11, 56 (2022). https://doi.org/10.1038/s41389-022-00434-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41389-022-00434-2
This article is cited by
-
Evidence of steady-state fibroblast subtypes in the normal human breast as cells-of-origin for perturbed-state fibroblasts in breast cancer
Breast Cancer Research (2024)
-
CAFs vs. TECs: when blood feuds fuel cancer progression, dissemination and therapeutic resistance
Cellular Oncology (2024)
-
Discovery of YS-1 as a cell line of gastric inflammatory cancer-associated fibroblasts
Molecular Biology Reports (2024)
-
Extracellular matrix remodeling in tumor progression and immune escape: from mechanisms to treatments
Molecular Cancer (2023)
-
Functional roles of SRC signaling in pancreatic cancer: Recent insights provide novel therapeutic opportunities
Oncogene (2023)