Abstract
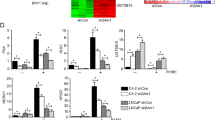
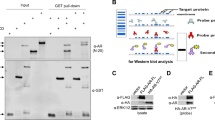
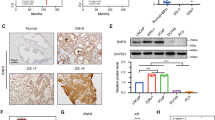
Aberrant transcriptional activity of androgen receptor (AR) is one of the dominant mechanisms for developing of castration-resistant prostate cancer (CRPC). Analyzing AR-transcriptional complex related to CRPC is therefore important towards understanding the mechanism of therapy resistance. While studying its mechanism, we observed that a transmembrane protein called neuropilin-2 (NRP2) plays a contributory role in forming a novel AR-transcriptional complex containing nuclear pore proteins. Using immunogold electron microscopy, high-resolution confocal microscopy, chromatin immunoprecipitation, proteomics, and other biochemical techniques, we delineated the molecular mechanism of how a specific splice variant of NRP2 becomes sumoylated upon ligand stimulation and translocates to the inner nuclear membrane. This splice variant of NRP2 then stabilizes the complex between AR and nuclear pore proteins to promote CRPC specific gene expression. Both full-length and splice variants of AR have been identified in this specific transcriptional complex. In vitro cell line-based assays indicated that depletion of NRP2 not only destabilizes the AR-nuclear pore protein interaction but also inhibits the transcriptional activities of AR. Using an in vivo bone metastasis model, we showed that the inhibition of NRP2 led to the sensitization of CRPC cells toward established anti-AR therapies such as enzalutamide. Overall, our finding emphasize the importance of combinatorial inhibition of NRP2 and AR as an effective therapeutic strategy against treatment refractory prostate cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Powers E, Karachaliou GS, Kao C, Harrison MR, Hoimes CJ, George DJ, et al. Novel therapies are changing treatment paradigms in metastatic prostate cancer. J Hematol Oncol. 2020;13:144.
Schmidt KT, Huitema ADR, Chau CH, Figg WD. Resistance to second-generation androgen receptor antagonists in prostate cancer. Nat Rev Urol. 2021;18:209–26.
Imamura Y, Sadar MD. Androgen receptor targeted therapies in castration-resistant prostate cancer: Bench to clinic. Int J Urol. 2016;23:654–65.
Tucci M, Zichi C, Buttigliero C, Vignani F, Scagliotti GV, Di Maio M. Enzalutamide-resistant castration-resistant prostate cancer: challenges and solutions. Onco Targets Ther. 2018;11:7353–68.
Giacinti S, Poti G, Roberto M, Macrini S, Bassanelli M, DIP F, et al. Molecular Basis of Drug Resistance and Insights for New Treatment Approaches in mCRPC. Anticancer Res. 2018;38:6029–39.
Roumiguie M, Paoletti X, Neuzillet Y, Mathieu R, Vincendeau S, Kleinclauss F, et al. Apalutamide, darolutamide and enzalutamide in nonmetastatic castration-resistant prostate cancer: a meta-analysis. Future Oncol. 2021;17:1811–23.
Tucci M, Leone G, Buttigliero C, Zichi C, DI Stefano RF, Pignataro D, et al. Hormonal treatment and quality of life of prostate cancer patients: new evidence. Minerva Urol Nefrol. 2018;70:144–51.
Vander Ark A, Cao J, Li X. Mechanisms and approaches for overcoming enzalutamide resistance in prostate cancer. Front Oncol. 2018;8:180.
He Y, Wei T, Ye Z, Orme JJ, Lin D, Sheng H, et al. A noncanonical AR addiction drives enzalutamide resistance in prostate cancer. Nat Commun. 2021;12:1521.
Schweizer MT, Haugk K, McKiernan JS, Gulati R, Cheng HH, Maes JL, et al. A phase I study of niclosamide in combination with enzalutamide in men with castration-resistant prostate cancer. PloS ONE. 2018;13:e0198389.
Shafran JS, Andrieu GP, Gyorffy B, Denis GV. BRD4 regulates metastatic potential of castration-resistant prostate cancer through AHNAK. Mol Cancer Res. 2019;17:1627–38.
Asangani IA, Dommeti VL, Wang X, Malik R, Cieslik M, Yang R, et al. Therapeutic targeting of BET bromodomain proteins in castration-resistant prostate cancer. Nature. 2014;510:278–82.
Coleman DJ, Gao L, Schwartzman J, Korkola JE, Sampson D, Derrick DS, et al. Maintenance of MYC expression promotes de novo resistance to BET bromodomain inhibition in castration-resistant prostate cancer. Sci Rep. 2019;9:3823.
Wang L, Xu M, Kao CY, Tsai SY, Tsai MJ. Small molecule JQ1 promotes prostate cancer invasion via BET-independent inactivation of FOXA1. J Clin Investig. 2020;130:1782–92.
Sulpice E, Plouet J, Berge M, Allanic D, Tobelem G, Merkulova-Rainon T. Neuropilin-1 and neuropilin-2 act as coreceptors, potentiating proangiogenic activity. Blood. 2008;111:2036–45.
Parker MW, Linkugel AD, Goel HL, Wu T, Mercurio AM, Vander, et al. Structural basis for VEGF-C binding to neuropilin-2 and sequestration by a soluble splice form. Structure. 2015;23:677–87.
Rossignol M, Gagnon ML, Klagsbrun M. Genomic organization of human neuropilin-1 and neuropilin-2 genes: identification and distribution of splice variants and soluble isoforms. Genomics. 2000;70:211–22.
Fricker M, Hollinshead M, White N, Vaux D. Interphase nuclei of many mammalian cell types contain deep, dynamic, tubular membrane-bound invaginations of the nuclear envelope. J Cell Biol. 1997;136:531–44.
Drozdz MM, Vaux DJ. Shared mechanisms in physiological and pathological nucleoplasmic reticulum formation. Nucleus. 2017;8:34–45.
Ibarra A, Hetzer MW. Nuclear pore proteins and the control of genome functions. Genes Dev. 2015;29:337–49.
Rodriguez-Navarro S, Fischer T, Luo MJ, Antunez O, Brettschneider S, Lechner J, et al. Sus1, a functional component of the SAGA histone acetylase complex and the nuclear pore-associated mRNA export machinery. Cell. 2004;116:75–86.
Garcia-Oliver E, Garcia-Molinero V, Rodriguez-Navarro S. mRNA export and gene expression: the SAGA-TREX-2 connection. Biochim Biophys Acta. 2012;1819:555–65.
Labade AS, Karmodiya K, Sengupta K. HOXA repression is mediated by nucleoporin Nup93 assisted by its interactors Nup188 and Nup205. Epigenetics Chromatin. 2016;9:54.
Sump B, Brickner JH. Nup98 regulation of histone methylation promotes normal gene expression and may drive leukemogenesis. Genes Dev. 2017;31:2201–3.
Franks TM, Hetzer MW. The role of Nup98 in transcription regulation in healthy and diseased cells. Trends Cell Biol. 2013;23:112–7.
Liang Y, Franks TM, Marchetto MC, Gage FH, Hetzer MW. Dynamic association of NUP98 with the human genome. PLoS Genet. 2013;9:e1003308.
Dutta S, Roy S, Polavaram NS, Baretton GB, Muders MH, Batra S, et al. NRP2 transcriptionally regulates its downstream effector WDFY1. Sci Rep. 2016;6:23588.
Coutinho I, Day TK, Tilley WD, Selth LA. Androgen receptor signaling in castration-resistant prostate cancer: a lesson in persistence. Endocr Relat Cancer. 2016;23:T179–97.
Sharma NL, Massie CE, Ramos-Montoya A, Zecchini V, Scott HE, Lamb AD, et al. The androgen receptor induces a distinct transcriptional program in castration-resistant prostate cancer in man. Cancer Cell. 2013;23:35–47.
Massie CE, Lynch A, Ramos-Montoya A, Boren J, Stark R, Fazli L, et al. The androgen receptor fuels prostate cancer by regulating central metabolism and biosynthesis. EMBO J. 2011;30:2719–33.
Tan KN, Avery VM, Carrasco-Pozo C. Metabolic roles of androgen receptor and Tip60 in androgen-dependent prostate cancer. Int J Mol Sci. 2020;21:6622.
Borkowetz A, Froehner M, Rauner M, Conrad S, Erdmann K, Mayr T, et al. Neuropilin-2 is an independent prognostic factor for shorter cancer-specific survival in patients with Acinar adenocarcinoma of the prostate. Int J Cancer. 2020;146:2619–27.
Palancade B, Liu X, Garcia-Rubio M, Aguilera A, Zhao X, Doye V. Nucleoporins prevent DNA damage accumulation by modulating Ulp1-dependent sumoylation processes. Mol Biol Cell. 2007;18:2912–23.
Ruben GJ, Kirkland JG, MacDonough T, Chen M, Dubey RN, Gartenberg MR, et al. Nucleoporin mediated nuclear positioning and silencing of HMR. PloS ONE. 2011;6:e21923.
Radman-Livaja M, Ruben G, Weiner A, Friedman N, Kamakaka R, Rando OJ. Dynamics of Sir3 spreading in budding yeast: secondary recruitment sites and euchromatic localization. EMBO J. 2011;30:1012–26.
Kuhn TM, Capelson M. Nuclear pore proteins in regulation of chromatin state. Cells. 2019;8:1414.
Kuhn TM, Pascual-Garcia P, Gozalo A, Little SC, Capelson M. Chromatin targeting of nuclear pore proteins induces chromatin decondensation. J Cell Biol. 2019;218:2945–61.
Raices M, D’Angelo MA. Nuclear pore complexes and regulation of gene expression. Curr Opin cell Biol. 2017;46:26–32.
Ptak C, Aitchison JD, Wozniak RW. The multifunctional nuclear pore complex: a platform for controlling gene expression. Curr Opin Cell Biol. 2014;28:46–53.
Dieppois G, Stutz F. Connecting the transcription site to the nuclear pore: a multi-tether process that regulates gene expression. J Cell Sci. 2010;123:1989–99.
Ibarra A, Benner C, Tyagi S, Cool J, Hetzer MW. Nucleoporin-mediated regulation of cell identity genes. Genes Dev. 2016;30:2253–8.
Kitazawa T, Rijli FM. Nuclear pore protein meets transcription factor in neural fate. Neuron. 2017;96:259–61.
Gomez-Cavazos JS, Hetzer MW. The nucleoporin gp210/Nup210 controls muscle differentiation by regulating nuclear envelope/ER homeostasis. J Cell Biol. 2015;208:671–81.
D’Angelo MA. Nuclear pore complexes as hubs for gene regulation. Nucleus. 2018;9:142–8.
Raices M, Bukata L, Sakuma S, Borlido J, Hernandez LS, Hart DO, et al. Nuclear pores regulate muscle development and maintenance by assembling a localized Mef2C complex. Dev Cell. 2017;41:540–54.e7.
Holzer K, Ori A, Cooke A, Dauch D, Drucker E, Riemenschneider P, et al. Nucleoporin Nup155 is part of the p53 network in liver cancer. Nat Commun. 2019;10:2147.
Rodriguez-Bravo V, Pippa R, Song WM, Carceles-Cordon M, Dominguez-Andres A, Fujiwara N, et al. Nuclear pores promote lethal prostate cancer by increasing POM121-driven E2F1, MYC, and AR nuclear import. Cell. 2018;174:1200–15.e20.
Su Y, Pelz C, Huang T, Torkenczy K, Wang X, Cherry A, et al. Post-translational modification localizes MYC to the nuclear pore basket to regulate a subset of target genes involved in cellular responses to environmental signals. Genes Dev. 2018;32:1398–419.
Gemmill RM, Nasarre P, Nair-Menon J, Cappuzzo F, Landi L, D'Incecco A, et al. The neuropilin 2 isoform NRP2b uniquely supports TGFbeta-mediated progression in lung cancer. Sci Signal. 2017;10:eaag0528.
Savoy RM, Chen L, Siddiqui S, Melgoza FU, Durbin-Johnson B, Drake C, et al. Transcription of Nrdp1 by the androgen receptor is regulated by nuclear filamin A in prostate cancer. Endocr Relat Cancer. 2015;22:369–86.
Stanton MJ, Dutta S, Zhang H, Polavaram NS, Leontovich AA, Honscheid P, et al. Autophagy control by the VEGF-C/NRP-2 axis in cancer and its implication for treatment resistance. Cancer research. 2013;73:160–71.
Polavaram NS, Dutta S, Islam R, Bag AK, Roy S, Poitz D, et al. Tumor- and osteoclast-derived NRP2 in prostate cancer bone metastases. Bone Res. 2021;9:24.
Kersemans V, Cornelissen B, Allen PD, Beech JS, Smart SC. Subcutaneous tumor volume measurement in the awake, manually restrained mouse using MRI. J Magn Reson Imaging. 2013;37:1499–504.
Acknowledgements
The authors thank all stuff members of the Advanced Microscopy Core and Genomics Core Facility of UNMC. We also extend our sincere thanks to the UNMC Bioinformatics and Bio-statistician Core for their support to analyze the data. The raw data has been deposited in Gene Expression Omnibus with the accession number GSE205150.
Funding
This work was supported by grants for SD (1R21CA241234-01, NE-LB506, Lageschulte Fund), KD (R01CA182435, R01CA239343, DoD W81XWH2110628), MHM and LHH (DFG project number 273676790), and MHM (DFG project number 416001651). MM is funded by the Rudolf-Becker-Foundation for his professorship. The construction of the prostate cancer tissue microarray was funded by the DFG Forschergruppe-1586 SKELMET to LCH and SF.
Author information
Authors and Affiliations
Contributions
SD and KD has designed the project and SD performed most of the work. NSP, RI, SB, SB, SR have assisted some of the work. TM has developed the NRP2-HA-tagged plasmid. SAA did the mass spec. AD took the electron microscopic images. MI, AB, SC, SF, MW, GBB, LCH, MHM were involved in TMA development and staining of NRP2. PG help us in promoter assay. PG, KJP, SKB and MHM critically evaluated the work and time to time provide there suggestion. DLK performed and analyzed the ChIP-seq.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dutta, S., Polavaram, N.S., Islam, R. et al. Neuropilin-2 regulates androgen-receptor transcriptional activity in advanced prostate cancer. Oncogene 41, 3747–3760 (2022). https://doi.org/10.1038/s41388-022-02382-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02382-y
This article is cited by
-
Neuropilin-2 promotes lineage plasticity and progression to neuroendocrine prostate cancer
Oncogene (2022)
-
Stem Cells as Target for Prostate cancer Therapy: Opportunities and Challenges
Stem Cell Reviews and Reports (2022)
-
Role of Neuropilin-2-mediated signaling axis in cancer progression and therapy resistance
Cancer and Metastasis Reviews (2022)