Abstract
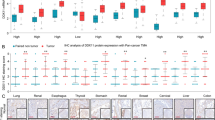
Chromodomain Y-like 2 (CDYL2), as a member of CDY family known to be involved in spermatogenesis, has been reported to participate in breast cancer development recently, but its exact biological role in hepatocellular carcinoma (HCC) remains unclear. Here, we observed that CDYL2 was down-regulated in human primary HCC tissues and the low levels of CDYL2 expression were correlated with poor survival. Gain- and loss-of-function experiments showed that CDYL2 inhibited the proliferation and metastasis of HCC cells in vitro and in vivo. Mechanistically, CDYL2 down-regulates solute carrier family 7 member 6 (SLC7A6) by decreasing the enrichment of H3K4me3 on the promoter region of SLC7A6. Additionally, we also found that signal transducer and activator of transcription 5A (STAT5A) could directly and positively regulate the expression of CDYL2. Thus, CDYL2 was regulated by STAT5A, and suppressed the amino acid transportation through down-regulation of SLC7A6, and then inhibits the mTORC1/S6K pathway, a master regulator of cell growth. Consistently, CDYL2 expression correlated significantly with STAT5A and SLC7A6 expression in HCC. Collectively, we propose a model for a STAT5A/CDYL2/SLC7A6 axis that provides novel insight into CDYL2, which may serve as a potential factor for predicting prognosis and a therapeutic target for HCC patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All datasets on which the conclusions of the paper rely are available to readers.
References
McGlynn KA, Petrick JL, El-Serag HB. Epidemiology of hepatocellular carcinoma. Hepatology. 2021;73:4–13.
Gunasekaran G, Bekki Y, Lourdusamy V, Schwartz M. Surgical treatments of hepatobiliary cancers. Hepatology. 2021;73:128–36.
Chidambaranathan-Reghupaty S, Fisher PB, Sarkar D. Hepatocellular carcinoma (HCC): epidemiology, etiology and molecular classification. Adv Cancer Res. 2021;149:1–61.
Dorus S, Gilbert SL, Foster ML, Barndt RJ, Lahn BT. The CDY-related gene family: coordinated evolution in copy number, expression profile and protein sequence. Hum Mol Genet. 2003;12:1643–50.
Wang A, Yasue H, Li L, Takashima M, de Leon FA, Liu WS. Molecular characterization of the bovine chromodomain Y-like genes. Anim Genet. 2008;39:207–16.
Zhang Y, Yang X, Gui B, Xie G, Zhang D, Shang Y, et al. Corepressor protein CDYL functions as a molecular bridge between polycomb repressor complex 2 and repressive chromatin mark trimethylated histone lysine 27. J Biol Chem. 2011;286:42414–25.
Mulligan P, Westbrook TF, Ottinger M, Pavlova N, Chang B, Macia E, et al. CDYL bridges REST and histone methyltransferases for gene repression and suppression of cellular transformation. Mol Cell. 2008;32:718–26.
Escamilla-Del-Arenal M, da Rocha ST, Spruijt CG, Masui O, Renaud O, Smits AH, et al. Cdyl, a new partner of the inactive X chromosome and potential reader of H3K27me3 and H3K9me2. Mol Cell Biol. 2013;33:5005–20.
Lahn BT, Tang ZL, Zhou J, Barndt RJ, Parvinen M, Allis CD, et al. Previously uncharacterized histone acetyltransferases implicated in mammalian spermatogenesis. Proc Natl Acad Sci USA. 2002;99:8707–12.
Wu H, Zhang H, Wang P, Mao Z, Feng L, Wang Y, et al. Short-Form CDYLb but not long-form CDYLa functions cooperatively with histone methyltransferase G9a in hepatocellular carcinomas. Genes Chromosome Cancer. 2013;52:644–55.
Siouda M, Dujardin AD, Barbollat-Boutrand L, Mendoza-Parra MA, Gibert B, Ouzounova M, et al. CDYL2 epigenetically regulates MIR124 to control NF-kappaB/STAT3-dependent breast cancer cell plasticity. iScience. 2020;23:101141.
Yang LF, Yang F, Zhang FL, Xie YF, Hu ZX, Huang SL, et al. Discrete functional and mechanistic roles of chromodomain Y-like 2 (CDYL2) transcript variants in breast cancer growth and metastasis. Theranostics. 2020;10:5242–58.
Kandasamy P, Gyimesi G, Kanai Y, Hediger MA. Amino acid transporters revisited: new views in health and disease. Trends Biochem Sci. 2018;43:752–89.
Wang Q, Holst J. L-type amino acid transport and cancer: targeting the mTORC1 pathway to inhibit neoplasia. Am J Cancer Res. 2015;5:1281–94.
Rask-Andersen M, Masuram S, Fredriksson R, Schiöth H. Solute carriers as drug targets: current use, clinical trials and prospective. Mol Asp Med. 2013;34:702–10.
Rotoli BM, Barilli A, Visigalli R, Ferrari F, Dall’Asta V. y + LAT1 and y + LAT2 contribution to arginine uptake in different human cell models: Implications in the pathophysiology of Lysinuric Protein Intolerance. J Cell Mol Med. 2020;24:921–9.
Jewell JL, Russell RC, Guan KL. Amino acid signalling upstream of mTOR. Nat Rev Mol Cell Biol. 2013;14:133–9.
Milewski K, Bogacińska-Karaś M, Fręśko I, Hilgier W, Jaźwiec R, Albrecht J, et al. Ammonia reduces intracellular asymmetric dimethylarginine in cultured astrocytes stimulating its y+LAT2 carrier-mediated loss. Int J Mol Sci. 2017;18:2308.
Liu P, Ge M, Hu J, Li X, Che L, Sun K, et al. A functional mammalian target of rapamycin complex 1 signaling is indispensable for c-Myc-driven hepatocarcinogenesis. Hepatology. 2017;66:167–81.
Lee TK, Man K, Poon RT, Lo CM, Yuen AP, Ng IO, et al. Signal transducers and activators of transcription 5b activation enhances hepatocellular carcinoma aggressiveness through induction of epithelial-mesenchymal transition. Cancer Res. 2006;66:9948–56.
Yu JH, Zhu BM, Wickre M, Riedlinger G, Chen W, Hosui A, et al. The transcription factors signal transducer and activator of transcription 5A (STAT5A) and STAT5B negatively regulate cell proliferation through the activation of cyclin-dependent kinase inhibitor (Cdkn2b) and Cdkn1a expression. Hepatology. 2010;52:1808–18.
Yu JH, Zhu BM, Riedlinger G, Kang K, Hennighausen L. The liver-specific tumor suppressor STAT5 controls expression of the reactive oxygen species-generating enzyme NOX4 and the proapoptotic proteins PUMA and BIM in mice. Hepatology. 2012;56:2375–86.
Audia JE, Campbell RM. Histone modifications and cancer. Cold Spring Harb Perspect Biol. 2016;8:a019521.
Sharma S, Kelly TK, Jones PA. Epigenetics in cancer. Carcinogenesis. 2010;31:27–36.
Qiu Z, Zhu W, Meng H, Tong L, Li X, Luo P, et al. CDYL promotes the chemoresistance of small cell lung cancer by regulating H3K27 trimethylation at the CDKN1C promoter. Theranostics. 2019;9:4717–29.
Vermeulen M, Eberl HC, Matarese F, Marks H, Denissov S, Butter F, et al. Quantitative interaction proteomics and genome-wide profiling of epigenetic histone marks and their readers. Cell. 2020;142:967–80.
Zhang Y, Yang X, Gui B, Xie G, Zhang D, Shang Y, et al. Corepressor protein CDYL functions as a molecular bridge between polycomb repressor complex 2 and repressive chromatin mark trimethylated histone lysine 27. J Biol Chem. 2011;286:42414–25.
Shen E, Shulha H, Weng Z, Akbarian S. Regulation of histone H3K4 methylation in brain development and disease. Philos Trans R Soc Lond Ser B Biol Sci. 2014;369:20130514.
Steve D, Gilbert SL, Forster ML, Barndt RJ, Lahn BT. The CDY-related gene family: coordinated evolution in copy number, expression profile and protein sequence. Hum Mol Genet. 2003;12:1643–50.
Xia X, Zhou X, Quan Y, Hu Y, Xing F, Li Z, et al. Germline deletion of Cdyl causes teratozoospermia and progressive infertility in male mice. Cell Death Dis. 2019;10:229.
Liu S, Yu H, Liu Y, Liu X, Shang Y. Chromodomain protein CDYL acts as a Crotonyl-CoA hydratase to regulate histone crotonylation and spermatogenesis. Mol Cell. 2017;67:853–66.
Mulligan P, Westbrook TF, Ottinger M, Pavlova N, Chang B, Macia E, et al. CDYL bridges REST and histone methyltransferases for gene repression and suppression of cellular transformation. Mol Cell. 2008;32:718–26.
Abu-Zhayia ER, Awwad SW, Ben-Oz BM, Khoury-Haddad H, Ayoub N. CDYL1 fosters double-strand break-induced transcription silencing and promotes homology-directed repair. J Mol Cell Biol. 2018;10:341–57.
Tang JZ, Zuo ZH, Kong XJ, Steiner M, Yin Z, Perry JK, et al. Signal transducer and activator of transcription (STAT)-5A and STAT5B differentially regulate human mammary carcinoma cell behavior. Endocrinology. 2010;12:43–55.
Bernaciak TM, Zareno J, Parsons JT, Silva CM. A novel role for signal transducer and activator of transcription 5b (STAT5b) in β1-integrin-mediated human breast cancer cell migration. Breast Cancer Res. 2009;11:1–11.
Kollmann S, Grundschober E, Maurer B, Warsch W, Grausenburger R, Edlinger L, et al. Twins with different personalities: STAT5B-but not STAT5A-has a key role in BCR/ABL-induced leukemia. Leukemia. 2019;33:1583–97.
Jiang Y, Tao Y, Zhang X, Wei X, Li M, He X, et al. Loss of STAT5A promotes glucose metabolism and tumor growth through miRNA AKT signaling in hepatocellular carcinoma. Mol Oncol. 2021;15:710–24.
Yue M, Jiang J, Gao P, Liu H, Qing G. Oncogenic MYC activates a feedforward regulatory loop promoting essential amino acid metabolism and tumorigenesis. Cell Rep. 2017;21:3819–32.
Yang J, Zhang L, Jiang Z, Ge C, Zhao F, Jiang J, et al. TCF12 promotes the tumorigenesis and metastasis of hepatocellular carcinoma via upregulation of CXCR4 expression. Theranostics. 2019;9:5810–27.
Zhang L, Huo Q, Ge C, Zhao F, Zhou Q, Chen X, et al. ZNF143-mediated H3K9 trimethylation upregulates CDC6 by activating MDIG in hepatocellular carcinoma. Cancer Res. 2020;80:2599–611.
Acknowledgements
The authors are grateful to Dr. Daqiang Li for donating pLVX-CDYL2 vector. This work was supported by the National Natural Science Foundation of China [Grant number 82173331, 81972580, 81773152].
Author information
Authors and Affiliations
Contributions
XC, JL performed study, writing and revision of the paper. ZW, XZ helped to perform this study. LZ, LZ, XL provided acquisition, analysis of data. CG performed immunohistochemical. FZ, MY carried out animal experiments. TC, HX, YC provided clinical samples. HT, HL, JL provided technical support. JL supervised this study. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Chen, X., Wang, Z., Zhao, X. et al. STAT5A modulates CDYL2/SLC7A6 pathway to inhibit the proliferation and invasion of hepatocellular carcinoma by targeting to mTORC1. Oncogene 41, 2492–2504 (2022). https://doi.org/10.1038/s41388-022-02273-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02273-2
This article is cited by
-
Baicalein improves the chemoresistance of ovarian cancer through regulation of CirSLC7A6
Journal of Ovarian Research (2023)
-
SL-scan identifies synthetic lethal interactions in cancer using metabolic networks
Scientific Reports (2023)
-
Chromodomain on Y-like 2 (CDYL2) implicated in mitosis and genome stability regulation via interaction with CHAMP1 and POGZ
Cellular and Molecular Life Sciences (2023)