Abstract
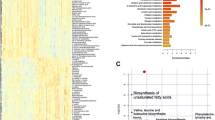
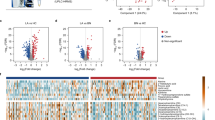
Discrimination of malignancy from thyroid nodules poses challenges in clinical practice. We aimed to identify the plasma metabolomic biomarkers in discriminating papillary thyroid cancer (PTC) from benign thyroid nodule (BTN). Metabolomics profiling of plasma was performed in two independent cohorts of 651 subjects of PTC (n = 215), BTN (n = 230), and healthy controls (n = 206). In addition, 132 patients with thyroid micronodules (<1 cm) and 44 patients with BTN suspected malignancy by ultrasound were used for biomarker validation. Recursive feature elimination algorithm was used for metabolic biomarkers selecting. Significant differential metabolites were demonstrated in patients with thyroid nodules (PTC and BTN) from healthy controls (P = 0.0001). A metabolic biomarker panel (17 differential metabolites) was identified to discriminate PTC from BTN with an AUC of 97.03% (95% CI: 95.28–98.79%), 91.89% sensitivity, and 92.63% specificity in discovery cohort. The panel had an AUC of 92.72% (95% CI: 87.46–97.99%), 86.57% sensitivity, and 92.50% specificity in validation cohort. The metabolic biomarker signature could correctly identify 84.09% patients whose nodules were suspected malignant by ultrasonography but finally histological benign. Moreover, high accuracy of 87.88% for diagnosis of papillary thyroid microcarcinoma was displayed by this panel and showed significant improvement in accuracy, AUC and specificity when compared with ultrasound. We identified a novel metabolic biomarker signature to discriminate PTC from BTN. The clinical use of this biomarker panel would have improved diagnosis stratification of thyroid microcarcinoma in comparison to ultrasound.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The metabolomics sequencing data and additional data related to this article will be shared on reasonable request to the corresponding author.
References
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66:115–32.
Howlader NNA, Krapcho M, Miller D, Brest A, Yu M, Ruhl J, et al. SEER cancer statistics review, 1975–2017. National Cancer Institute. 2020. Available from: https://seer.cancer.gov/csr/1975_2017.
Kitahara CM, Sosa JA. The changing incidence of thyroid cancer. Nat Rev Endocrinol. 2016;12:646–53.
Cabanillas ME, McFadden DG, Durante C. Thyroid cancer. Lancet. 2016;388:2783–95.
Brito JP, Gionfriddo MR, Al Nofal A, Boehmer KR, Leppin AL, Reading C, et al. The accuracy of thyroid nodule ultrasound to predict thyroid cancer: systematic review and meta-analysis. J Clin Endocrinol Metab. 2014;99:1253–63.
Tessler FN, Middleton WD, Grant EG, Hoang JK, Berland LL, Teefey SA, et al. ACR Thyroid Imaging, Reporting and Data System (TI-RADS): white paper of the ACR TI-RADS Committee. J Am Coll Radiol. 2017;14:587–95.
Burman KD, Wartofsky L. CLINICAL PRACTICE. Thyroid Nodules. N Engl J Med. 2015;373:2347–56.
Yang R, Zou X, Zeng H, Zhao Y, Ma X. Comparison of diagnostic performance of five different ultrasound TI-RADS classification guidelines for thyroid nodules. Front Oncol. 2020;10:598225.
Haugen BR, Alexander EK, Bible KC, Doherty GM, Mandel SJ, Nikiforov YE, et al. 2015 American Thyroid Association Management Guidelines for Adult Patients with Thyroid Nodules and Differentiated Thyroid Cancer: The American Thyroid Association Guidelines Task Force on Thyroid Nodules and Differentiated Thyroid Cancer. Thyroid. 2016;26:1–133.
Manning AM, Yang H, Falciglia M, Mark JR, Steward DL. Thyroid ultrasound-guided fine-needle aspiration cytology results: observed increase in indeterminate rate over the past decade. Otolaryngol Head Neck Surg. 2017;156:611–5.
Cibas ES, Baloch ZW, Fellegara G, LiVolsi VA, Raab SS, Rosai J, et al. A prospective assessment defining the limitations of thyroid nodule pathologic evaluation. Ann Intern Med. 2013;159:325–32.
Davies L, Welch HG. Increasing incidence of thyroid cancer in the United States, 1973-2002. JAMA. 2006;295:2164–7.
Leboulleux S, Tuttle RM, Pacini F, Schlumberger M. Papillary thyroid microcarcinoma: time to shift from surgery to active surveillance? Lancet Diabetes Endocrinol. 2016;4:933–42.
Mazeh H, Chen H. Advances in surgical therapy for thyroid cancer. Nat Rev Endocrinol. 2011;7:581–8.
Yu XM, Wan Y, Sippel RS, Chen H. Should all papillary thyroid microcarcinomas be aggressively treated? An analysis of 18,445 cases. Ann Surg. 2011;254:653–60.
Ma HJ, Yang JC, Leng ZP, Chang Y, Kang H, Teng LH. Preoperative prediction of papillary thyroid microcarcinoma via multiparameter ultrasound. Acta Radiol. 2017;58:1303–11.
de Meer SG, Schreinemakers JM, Zelissen PM, Stapper G, Sie-Go DM, Rinkes IH, et al. Fine-needle aspiration of thyroid tumors: identifying factors associated with adequacy rate in a large academic center in the Netherlands. Diagn Cytopathol. 2012;40 Suppl 1:E21–6.
Davies L, Morris LG, Haymart M, Chen AY, Goldenberg D, Morris J, et al. American Association of Clinical Endocrinologists and American College of Endocrinology Disease State Clinical Review: the increasing incidence of thyroid cancer. Endocr Pract. 2015;21:686–96.
Cradic KW, Milosevic D, Rosenberg AM, Erickson LA, McIver B, Grebe SK. Mutant BRAF(T1799A) can be detected in the blood of papillary thyroid carcinoma patients and correlates with disease status. J Clin Endocrinol Metab. 2009;94:5001–9.
Hu S, Ewertz M, Tufano RP, Brait M, Carvalho AL, Liu D, et al. Detection of serum deoxyribonucleic acid methylation markers: a novel diagnostic tool for thyroid cancer. J Clin Endocrinol Metab. 2006;91:98–104.
Lee JC, Zhao JT, Clifton-Bligh RJ, Gill A, Gundara JS, Ip JC, et al. MicroRNA-222 and microRNA-146b are tissue and circulating biomarkers of recurrent papillary thyroid cancer. Cancer. 2013;119:4358–65.
Mayerle J, Kalthoff H, Reszka R, Kamlage B, Peter E, Schniewind B, et al. Metabolic biomarker signature to differentiate pancreatic ductal adenocarcinoma from chronic pancreatitis. Gut. 2018;67:128–37.
Li J, Li J, Wang H, Qi LW, Zhu Y, Lai M. Tyrosine and glutamine-leucine are metabolic markers of early-stage colorectal cancers. Gastroenterology. 2019;157:257–9.
Banales JM, Inarrairaegui M, Arbelaiz A, Milkiewicz P, Muntane J, Munoz-Bellvis L, et al. Serum metabolites as diagnostic biomarkers for cholangiocarcinoma, hepatocellular carcinoma, and primary sclerosing cholangitis. Hepatology. 2019;70:547–62.
Luo P, Yin P, Hua R, Tan Y, Li Z, Qiu G, et al. A Large-scale, multicenter serum metabolite biomarker identification study for the early detection of hepatocellular carcinoma. Hepatology. 2018;67:662–75.
Farrokhi Yekta R, Rezaei Tavirani M, Arefi Oskouie A, Mohajeri-Tehrani MR, Soroush AR, Akbarzadeh Baghban A. Serum-based metabolic alterations in patients with papillary thyroid carcinoma unveiled by non-targeted 1H-NMR metabolomics approach. Iran J Basic Med Sci. 2018;21:1140–7.
Huang FQ, Li J, Jiang L, Wang FX, Alolga RN, Wang MJ, et al. Serum-plasma matched metabolomics for comprehensive characterization of benign thyroid nodule and papillary thyroid carcinoma. Int J Cancer. 2019;144:868–76.
Chen J, Hu Q, Hou H, Wang S, Zhang Y, Luo Y, et al. Metabolite analysis-aided diagnosis of papillary thyroid cancer. Endocr Relat Cancer. 2019;26:829–41.
Abooshahab R, Hooshmand K, Razavi SA, Gholami M, Sanoie M, Hedayati M. Plasma metabolic profiling of human thyroid nodules by gas chromatography-mass spectrometry (GC-MS)-based untargeted metabolomics. Front Cell Dev Biol. 2020;8:385.
Klubo-Gwiezdzinska J, Wartofsky L. The role of molecular diagnostics in the management of indeterminate thyroid nodules. J Clin Endocrinol Metab. 2018;103:3507–10.
Steward DL, Carty SE, Sippel RS, Yang SP, Sosa JA, Sipos JA, et al. Performance of a multigene genomic classifier in thyroid nodules with indeterminate cytology: a prospective blinded multicenter study. JAMA Oncol. 2019;5:204–12.
Nishino M, Nikiforova M. Update on molecular testing for cytologically indeterminate thyroid nodules. Arch Pathol Lab Med. 2018;142:446–57.
Nikiforova MN, Mercurio S, Wald AI, Barbi de Moura M, Callenberg K, Santana-Santos L, et al. Analytical performance of the ThyroSeq v3 genomic classifier for cancer diagnosis in thyroid nodules. Cancer. 2018;124:1682–90.
Valvo V, Iesato A, Kavanagh TR, Priolo C, Zsengeller Z, Pontecorvi A, et al. Fine-tuning lipid metabolism by targeting mitochondria-associated acetyl-CoA-carboxylase 2 in BRAF(V600E) papillary. Thyroid Carcinoma Thyroid. 2021;31:1335–58.
Abraham A, Kattoor AJ, Saldeen T, Mehta JL. Vitamin E and its anticancer effects. Crit Rev Food Sci Nutr. 2019;59:2831–8.
Stevens VL, Wang Y, Carter BD, Gaudet MM, Gapstur SM. Serum metabolomic profiles associated with postmenopausal hormone use. Metabolomics. 2018;14:97.
Cheng Y, Schlosser P, Hertel J, Sekula P, Oefner PJ, Spiekerkoetter U, et al. Rare genetic variants affecting urine metabolite levels link population variation to inborn errors of metabolism. Nat Commun. 2021;12:964.
Pavlova NN, Thompson CB. The emerging hallmarks of cancer metabolism. Cell Metab. 2016;23:27–47.
Hofland J, Zandee WT, de Herder WW. Role of biomarker tests for diagnosis of neuroendocrine tumours. Nat Rev Endocrinol. 2018;14:656–69.
Wagner K, Arciaga R, Siperstein A, Milas M, Warshawsky I, Sethu S, et al. Thyrotropin receptor/thyroglobulin messenger ribonucleic acid in peripheral blood and fine-needle aspiration cytology: diagnostic synergy for detecting thyroid cancer. J Clin Endocrinol Metab. 2005;90:1921–4.
Nixon AM, Provatopoulou X, Kalogera E, Zografos GN, Gounaris A. Circulating thyroid cancer biomarkers: current limitations and future prospects. Clin Endocrinol. 2017;87:117–26.
Xu SL, Tian YY, Zhou Y, Liu LQ. Diagnostic value of circulating microRNAs in thyroid carcinoma: a systematic review and meta-analysis. Clin Endocrinol. 2020;93:489–98.
Lamartina L, Grani G, Durante C, Filetti S, Cooper DS. Screening for differentiated thyroid cancer in selected populations. Lancet Diabetes Endocrinol. 2020;8:81–88.
Fussey JM, Bryant JL, Batis N, Spruce RJ, Hartley A, Good JS, et al. The clinical utility of cell-free DNA measurement in differentiated thyroid cancer: a systematic review. Front Oncol. 2018;8:132.
Cao S, Yu S, Yin Y, Su L, Hong S, Gong Y, et al. Genetic alterations in cfDNA of benign and malignant thyroid nodules based on amplicon-based next-generation sequencing. Ann Transl Med. 2020;8:1225.
Wada N, Duh QY, Sugino K, Iwasaki H, Kameyama K, Mimura T, et al. Lymph node metastasis from 259 papillary thyroid microcarcinomas: frequency, pattern of occurrence and recurrence, and optimal strategy for neck dissection. Ann Surg. 2003;237:399–407.
Dunn WB, Broadhurst D, Begley P, Zelena E, Francis-McIntyre S, Anderson N, et al. Procedures for large-scale metabolic profiling of serum and plasma using gas chromatography and liquid chromatography coupled to mass spectrometry. Nat Protoc. 2011;6:1060–83.
Smith CA, Want EJ, O’Maille G, Abagyan R, Siuzdak G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal Chem. 2006;78:779–87.
Chong J, Wishart DS, Xia J. Using MetaboAnalyst 4.0 for comprehensive and integrative metabolomics data analysis. Curr Protoc Bioinf. 2019;68:e86.
Rohart F, Gautier B, Singh A, Le Cao KA. mixOmics: an R package for ‘omics feature selection and multiple data integration. PLoS Comput Biol. 2017;13:e1005752.
Chong J, Yamamoto M, Xia J. MetaboAnalystR 2.0: from raw spectra to biological insights. Metabolites. 2019;9:57.
Benjamini YH, Controlling Y. the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc. 1995;57:289–300.
Kuhn M. Building predictive models in R using the caret package. J Stat Softw. 2008;28:1–26.
Robin X, Turck N, Hainard A, Tiberti N, Lisacek F, Sanchez JC, et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 2011;12:77.
Gu Z, Eils R, Schlesner M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics. 2016;32:2847–9.
Acknowledgements
We thank Zhen Cheng, XueJie Wang, Weiman He, and Fenghua Lai for help with the clinical database. We thank Yihao Liu for help in manuscript preparation, Wei Wang for technical assistance, and Bo Lin for access to thyroid cancer samples.
Author information
Authors and Affiliations
Contributions
SY, YTH, and JL collected the clinical samples, analyzed the data, and drafted the manuscript; CAL performed the bioinformatic analysis and drafted the manuscript; ZMG, XWC, LYZ, SP, SBH, LXX, XXL, RYL, SWC, BL, ZPW, and YBL collected and analyzed the data and commented on the study; WML collected human samples and commented on the study; JY and HPX designed and supervised the study and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Yu, S., Liu, C., Hou, Y. et al. Integrative metabolomic characterization identifies plasma metabolomic signature in the diagnosis of papillary thyroid cancer. Oncogene 41, 2422–2430 (2022). https://doi.org/10.1038/s41388-022-02254-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02254-5
This article is cited by
-
Cancer metabolites: promising biomarkers for cancer liquid biopsy
Biomarker Research (2023)
-
Small molecule metabolites: discovery of biomarkers and therapeutic targets
Signal Transduction and Targeted Therapy (2023)