Abstract
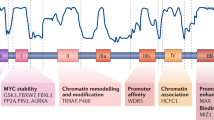
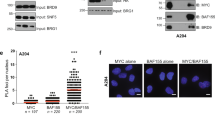
The PBAF complex, a member of SWI/SNF family of chromatin remodelers, plays an essential role in transcriptional regulation. We revealed a disease progression associated elevation of PHF10 subunit of PBAF in clinical melanoma samples. In melanoma cell lines, PHF10 interacts with MYC and facilitates the recruitment of PBAF complex to target gene promoters, therefore, augmenting MYC transcriptional activation of genes involved in the cell cycle progression. Depletion of either PHF10 or MYC induced G1 accumulation and a senescence-like phenotype. Our data identify PHF10 as a pro-oncogenic mechanism and an essential novel link between chromatin remodeling and MYC-dependent gene transcription.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Kadoch C, Crabtree GR. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci Adv. 2015;197:804–9.
Alfert A, Moreno N, Kerl K. The BAF complex in development and disease. Epigenetics Chromatin. 2019;12:1–15.
Centore RC, Sandoval GJ, Soares LMM, Kadoch C, Chan HM. Mammalian SWI/SNF chromatin remodeling complexes: emerging mechanisms and therapeutic strategies. Trends Genet. 2020;36:936–50.
Phelan ML, Sif S, Narlikar GJ, Kingston RE. Reconstitution of a core chromatin remodeling complex from SWI/SNF subunits. Mol Cell. 1999;3:247–53.
Trotter KW, Archer TK. The BRG1 transcriptional coregulator. Nucl Recept Signal. 2008;6:e004.
Ishizaka A, Mizutani T, Kobayashi K, Tando T, Sakurai K, Fujiwara T, et al. Double plant homeodomain (PHD) finger proteins DPF3a and -3b are required as transcriptional co-activators in SWI/SNF complex-dependent activation of NF-κB RelA/p50 heterodimer. J Biol Chem. 2012;287:11924–33.
Versteege I, Sevenet N, Lange J, Rousseau-Merck MF, Ambros P, Handgretinger R, et al. Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature. 1998;394:203–6.
Roberts СW, Galusha SA, McMenamin ME, Fletcher CD, Orkin SH. Haploinsufficiency of Snf5 (integrase interactor 1) predisposes to malignant rhabdoid tumors in mice. PNAS. 2000;5:13796–800.
Middeljans E, Wan X, Jansen PW, Sharma V, Stunnenberg HG, Logie C. SS18 together with animal-specific factors defines human BAF-type SWI/SNF complexes. PLoS ONE. 2012;7:e33834.
Crew AJ, Clark J, Fisher C, Gill S, Grimer R, Chand A, et al. Fusion of SYT to two genes, SSX1 and SSX2, encoding proteins with homology to the Kruppel-associated box in human synovial sarcoma. EMBO J. 1995;14:2333–40.
Kadoch C, Hargreaves DC, Hodges C, Elias L, Ho L, Ranish J, et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat Genet. 2013;45:592–602.
Poole CJ, van Riggelen J. MYC—master regulator of the cancer epigenome and transcriptome. Genes (Basel). 2017;8:142.
Dang CV. MYC on the path to cancer. Cell. 2012;149:22–35.
Brechalov AV, Georgieva SG, Soshnikova NV. Mammalian cells contain two functionally distinct PBAF complexes incorporating different isoforms of PHF10 signature subunit. Cell Cycle. 2014;13:1970–9.
Mashtalir N, D’Avino AR, Michel BC, Luo J, Pan J, Otto JE, et al. Modular organization and assembly of SWI/SNF family chromatin remodeling complexes. Cell. 2018;175:1272–88.e20.
Zeng L, Zhang Q, Li S, Plotnikov AN, Walsh MJ, Zhou M, et al. Mechanism and regulation of acetylated histone binding by the tandem PHD finger of DPF3b. Nature. 2010;466:258–62.
Soshnikova NV, Sheynov AA, Tatarskiy EV, Georgieva SG. The DPF domain as a unique structural unit participating in transcriptional activation, cell differentiation, and malignant. Transformation. 2020;12:115–23.
Shidlovskii YV, Krasnov AN, Nikolenko JV, Lebedeva LA, Kopantseva M, Ermolaeva MA, et al. A novel multidomain transcription coactivator SAYP can also repress transcription in heterochromatin. EMBO J. 2005;24:97–107.
Chalkley GE, Moshkin YM, Langenberg K, Bezstarosti K, Blastyak A, Gyurkovics H, et al. The transcriptional coactivator SAYP is a trithorax group signature subunit of the PBAP chromatin remodeling complex. Mol Cell Biol. 2008;28:2920–9.
Vorobyeva NE, Soshnikova NV, Nikolenko JV, Kuzmina JL, Nabirochkina EN, Georgieva S, et al. Transcription coactivator SAYP combines chromatin remodeler Brahma and transcription initiation factor TFIID into a single supercomplex. Proc Natl Acad Sci USA. 2009;106:11049–54.
Lessard J, Wu JI, Ranish JA, Wan M, Winslow MM, Staahl B, et al. An essential switch in subunit composition of a chromatin remodeling complex during neural development. Neuron. 2007;55:201–15.
Krasteva V, Crabtree GR, Lessard JA. The BAF45a/PHF10 subunit of SWI/SNF-like chromatin remodeling complexes is essential for hematopoietic stem cell maintenance. Exp Hematol. 2017;48:58–71.e15.
Viryasova GM, Tatarskiy VV Jr, Sheynov AA, Tatarskiy EV, Sud’ina GF, Georgieva SG, et al. PBAF lacking PHD domains maintains transcription in human neutrophils. Biochim Biophys Acta Mol Cell Res. 2019;1866:118525.
Cheng SW, Davies KP, Yung E, Beltran RJ, Yu J, Kalpana GV. c-MYC interacts with INI1/hSNF5 and requires the SWI/SNF complex for transactivation function. Nat Genet. 1999;22:102–5.
Stojanova A, Tu WB, Ponzielli R, Kotlyar M, Chan P, Boutros PC, et al. MYC interaction with the tumor suppressive SWI/SNF complex member INI1 regulates transcription and cellular transformation. Cell Cycle. 2016;15:1693–705.
Park J, Wood MA, Cole MD. BAF53 forms distinct nuclear complexes and functions as a critical c-Myc-interacting nuclear cofactor for oncogenic transformation. Mol Cell Biol. 2002;22:1307–16.
Zhuang D, Mannava S, Grachtchouk V, Tang W-H, Wawrzyniak JA, Berman AE, et al. c-MYC overexpression is required for continuous suppression of oncogene-induced senescence in melanoma cells. Oncogene. 2008;27:6623–34.
Sammak S, Allen MD, Hamdani N, Bycroft M, Zinzalla G. The structure of INI1/hSNF5 RPT1 and its interactions with the c-MYC:MAX heterodimer provide insights into the interplay between MYC and the SWI/SNF chromatin remodeling complex. FEBS J. 2018;285:4165–80.
Banga SS, Peng L, Dasgupta T, Palejwala V, Ozer HL. PHF10 is required for cell proliferation in normal and SV40-immortalized human fibroblast cells. Cytogenet Genome Res. 2010;126:227–42.
Panov VV, Kuzmina JL, Doronin SA, Kopantseva MR, Nabirochkina EN, Georgieva SG, et al. Transcription co-activator SAYP mediates the action of STAT activator. Nucleic Acids Res. 2012;40:2445–53.
Vorobyeva NE, Nikolenko JV, Nabirochkina EN, Krasnov AN, Shidlovskii YV, Georgieva SG, et al. SAYP and Brahma are important for ‘repressive’ and ‘transient’ Pol II pausing. Nucleic Acids Res. 2012;40:7319–31.
García-Gutiérrez L, Delgado MD, León J. Myc oncogene contributions to release of cell cycle brakes. Genes (Basel). 2019;10:244.
Mittal P, Roberts CWM. The SWI/SNF complex in cancer—biology, biomarkers and therapy. Nat Rev Clin Oncol. 2020;17:435–48.
Ribeiro-Silva C, Vermeulen W, Lans H. SWI/SNF: complex complexes in genome stability and cancer. DNA Repair (Amst). 2019;77:87–95.
Masliah-Planchon J, Bièche I, Guinebretière J-M, Bourdeaut F, Delattre O. SWI/SNF chromatin remodeling and human malignancies.Annu Rev Pathol. 2015;10:145–71. https://doi.org/10.1146/annurev-pathol-012414-040445.
Biegel JA, Zhou JY, Rorke LB, Stenstrom LB, Wainwright LM, Fogelgren B, et al. Germ-line and acquired mutations of INI1 in atypical teratoid and rhabdoid tumors. Cancer Res. 1999;59:74–79.
Sévenet N, Sheridan E, Amram D, Schneider P, Handgretinger R, Delattre O. Constitutional mutations of the hSNF5/INI1 gene predispose to a variety of cancers. Am J Hum Genet. 1999;65:1342–8.
Lu B, Shi H. An in-depth look at small cell carcinoma of the ovary, hypercalcemic type (SCCOHT): Clinical implications from recent molecular findings. J Cancer. 2019;10:223–37.
Buscarlet M, Krasteva V, Ho L, Simon C, Hebert J, Wilhelm B, et al. Essential role of BRG, the ATPase subunit of BAF chromatin remodeling complexes, in leukemia maintenance. Blood. 2014;123:1720–8.
Saladi SV, de la Serna IL. ATP dependent chromatin remodeling enzymes in embryonic stem cells. Bone. 2010;6:62–73.
Anbunathan H, Verstraten R, Singh AD, Harbour WJ, Bowcock AM. Integrative copy number analysis of uveal melanoma reveals novel candidate genes involved in tumorigenesis including a tumor suppressor role for PHF10/BAF45A. Clin Cancer Res. 2019;25:5156–66.
Bagnasco L, Tortolina L, Biasotti B, Castagnino N, Ponassi R, Tomati V, et al. Inhibition of a protein‐protein interaction between INI1 and c‐Myc by small peptidomimetic molecules inspired by Helix‐1 of c‐Myc: identification of a new target of potential antineoplastic interest. FASEB J. 2007;21:1256–63.
Shin KJ, Wall EA, Zavzavadjian JR, Santat LA, Liu J, Hwang J, et al. A single lentiviral vector platform for microRNA-based conditional RNA interference and coordinated transgene expression. Proc Natl Acad Sci USA. 2006;103:13759–64.
Stewart SA, Dykxhoorn DM, Palliser D, Mizuno H, Yu EY, An DS, et al. Lentivirus-delivered stable gene silencing by RNAi in primary cells. RNA. 2003;9:493–501.
Hermeking H, Rago C, Schuhmacher M, Li Q, Barrett JF, Obaya AJ, et al. Identification of CDK4 as a target of c-MYC. Proc Natl Acad Sci USA. 2000;97:2229–34.
Brechalov AV, Valieva ME, Georgieva SG, Soshnikova NV. PHF10 isoforms are phosphorylated in the PBAF mammalian chromatin remodeling complex. Mol Biol. 2016;50:278–83.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:10–12.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2009;26:139–40.
Robinson MD, Oshlack A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010;11:R25
Liao Y, Wang J, Jaehnig EJ, Shi Z, Zhang B. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 2019;47:W199–205.
Croft D, Mundo AF, Haw R, Milacic M, Weiser., Wu G, et al. The Reactome pathway knowledgebase. Nucleic Acids Res. 2014;42:472–7.
Ramírez F, Dündar F, Diehl S, Grüning BA, Manke T. DeepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 2014;42:187–91.
Egorov AA, Sakharova EA, Anisimova AS, Dmitriev SE, Gladyshev VN, Kulakovskiy IV. Svist4get: a simple visualization tool for genomic tracks from sequencing experiments. BMC Bioinforma. 2019;20:4–9.
Acknowledgements
We thank the Center for Precision Genome Editing and Genetic Technologies for Biomedicine (Institute of Gene Biology) supported by Ministry of Science and Higher Education of the Russian Federation (075-15-2019-1661) for providing the equipment.
Funding
This study was supported by the Russian Foundation of Basic Research (grant #17-54-33031 (to SGG) and NIH R21 CA220096 grant (to MAN).
Author information
Authors and Affiliations
Contributions
Conceptualization: NVS, MAN, SGG, AAS; methodology: NVS, VVT, NSK, AAS; validation: VVT, EVT, NSK; investigation: NVS, EVT, NSK, VVT; resources: EVT, SGG; data curation: NVS, VVT, NSK; writing (original draft): NVS, SGG, AAS, NSK; writing (review and editing): NVS, SGG, AAS, MAN; visualization: NVS, VVT, EVT, NSK; supervision: SGG; project administration: NVS, SGG, MAN; funding acquisition: SGG, MAN.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Reuven Agami.
Supplementary information
Rights and permissions
About this article
Cite this article
Soshnikova, N.V., Tatarskiy, E.V., Tatarskiy, V.V. et al. PHF10 subunit of PBAF complex mediates transcriptional activation by MYC. Oncogene 40, 6071–6080 (2021). https://doi.org/10.1038/s41388-021-01994-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01994-0