Abstract
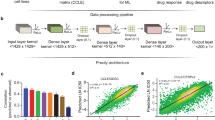
Recent advances in machine learning promise to yield novel insights by interrogation of large datasets ranging from gene expression and mutation data to CRISPR knockouts and drug screens. We combined existing and new algorithms with available experimental data to identify potentially clinically relevant relationships to provide a proof of principle for the promise of machine learning in oncological drug discovery. Specifically, we screened cell line data from the Cancer Dependency Map for the effects of azithromycin, which has been shown to kill cancer cells in vitro. Our findings demonstrate a strong relationship between Kallikrein Related Peptidase 6 (KLK6) mutation status and the ability of azithromycin to kill cancer cells in vitro. While the application of azithromycin showed no meaningful average effect in KLK6 wild-type cell lines, statistically significant enhancements of cell death are seen in multiple independent KLK6-mutated cancer cell lines. These findings suggest a potentially valuable clinical strategy in patients with KLK6-mutated malignancies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Travis J. Making the cut. Science. 2015;350:1456–7.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier EA. Programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337:816–21.
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, et al. (International Human Genome Sequencing Consortium (IHGSC)). Finishing the euchromatic sequence of the human genome. Nature. 2004;431:931–45.
Qiao X, Wang X, Shang Y, Li Y, Chen S. Azithromycin enhances anticancer activity of TRAIL by inhibiting autophagy and up- regulating the protein levels of DR4/5 in colon cancer cells in vitro and in vivo. Cancer Commun. 2018;38:43.
Fiorillo M, Toth F, Sotgia F, Lisanti MP. Doxycycline, azithromycin and vitamin C (DAV): a potent combination therapy for targeting mitochondria and eradicating cancer stem cells (CSCs). Aging. 2019;11:2202.
Lamb R, Ozsvari B, Lisanti CL, Tanowitz HB, Howell A, Martinez-Outschoorn UE, et al. Antibiotics that target mitochondria effectively eradicate cancer stem cells, across multiple tumor types: Treating cancer like an infectious disease. Oncotarget. 2015;6:4569.
Li F, Huang J, Ji D, Meng Q, Wang C, Chen S, et al. Azithromycin effectively inhibits tumor angiogenesis by suppressing vascular endothelial growth factor receptor 2 mediated signaling pathways in lung cancer. Oncol Lett. 2017;14:89–96.
Tsherniak A, Vazquez F, Montgomery PG, Weir BA, Kryukov G, Cowley GS, et al. Defining a cancer dependency map. 2017. Cell. 2017;170:564.
Ghandi M, Huang FW, Jané-Valbuena J, Kryukov GV, Lo CC, McDonald ER, et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature. 2019;569:503.
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, et al. Scikit-learn: Machine Learning in Python. J Mach Learn Res. 2011;12:2825.
Lundberg SM, Lee S. A unified approach to interpreting model predictions. In: Advances in neural information processing systems. Long Beach, CA, 30;2017.
Lundberg SM, Erion G, Chen H, DeGrave A, Prutkin JM, Nair B, et al. From local explanations to global understanding with explainable AI for trees. Nat Mach Intell. 2020;2:56.
Raileanu LE, Stoffel K. Theoretical Comparison between the Gini Index and Information Gain Criteria. Ann Math Artif Intell. 2004;41:77.
DepMap, Broad. DepMap 21Q1 Public. figshare. 2020. https://doi.org/10.6084/m9.figshare.13681534.v1.
Bazaga A, Leggate D, Weisser H. Genome-wide investigation of gene-cancer associations for the prediction of novel therapeutic targets in oncology. Sci Rep. 2020;10:10787.
Dezső Z, Ceccarelli M. Machine learning prediction of oncology drug targets based on protein and network properties. BMC Bioinform. 2020;21:104.
Madhukar NS, Khade PK, Huang L, Gayvert K, Galletti G, Stogniew M, et al. A Bayesian machine learning approach for drug target identification using diverse data types. Nat Commun. 2019;10:5221.
Lord CJ, Ashworth A. PARP inhibitors: synthetic lethality in the clinic. Science. 2017;355:1152.
Tamir A, Jag U, Sarojini S, Schindewolf C, Tanaka T, Gharbaran R, et al. Kallikrein family proteases KLK6 and KLK7 are potential early detection and diagnostic biomarkers for serous and papillary serous ovarian cancer subtypes. J Ovarian Res. 2014;7:109.
Haritos C, Michaelidou K, Mavridis K, Missitzis J, Ardavanis A, Griniatsos J, et al. Kallikrein-related peptidase 6 (KLK6) expression differentiates tumor subtypes and predicts clinical outcome in breast cancer patients. Clin Exp Med. 2018;18:203–13.
Ahmed N, Dorn J, Napieralski R, Drecoll E, Kotzsch M, Goettig P, et al. Clinical relevance of kallikrein-related peptidase 6 (KLK6) and 8 (KLK8) mRNA expression in advanced serous ovarian cancer. Biol Chem. 2016;397:1265.
Wang SM, Mao J, Li B, Wu W, Tang LL. Expression of KLK6 protein and mRNA in primary breast cancer and its clinical significance. Chin J Cell Mol Immunol. 2008;24:1087.
Chalmers ZR, Connelly CF, Fabrizio D, Gay L, Ali SM, Ennis R, et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 2017;9:34.
Jiang X, Baucom C, Elliott RL. Mitochondrial toxicity of azithromycin results in aerobic glycolysis and DNA damage of human mammary epithelia and fibroblasts. Antibiotics. 2019;8:E110.
Corsello SM, Nagari RT, Spangler RD, Rossen J, Kocak M, Bryan JG, et al. Discovering the anticancer potential of non-oncology drugs by systematic viability profiling. Nat Cancer. 2020;1:235–48.
Schrader CH, Kolb M, Zaoui K, Flechtenmacher C, Grabe N, Weber KJ, et al. Kallikrein-related peptidase 6 regulates epithelial-to-mesenchymal transition and serves as prognostic biomarker for head and neck squamous cell carcinoma patients. Mol Cancer. 2015;14:107.
Wang H, Unternaehrer JJ. Epithelial-mesenchymal transition and cancer stem cells: at the crossroads of differentiation and dedifferentiation. Developmental Dyn. 2019;248:10–20.
Shimada K, Muhlich JL, Mitchison TJ. A tool for browsing the Cancer Dependency Map reveals functional connections between genes and helps predict the efficacy and selectivity of candidate cancer drugs. 2019. https://www.biorxiv.org/content/10.1101/2019.12.13.874776v1.
Acknowledgements
The authors would like to thank Eoin McDonnell, Kris Wood, and Jonathan Mizrahi for relevant discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
JS, GV, and YB have roles at Red Cell Partners and Zephyr AI that involve the application of AI to cancer drug discovery. The authors all report no known financial interest in azithromycin.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sherman, J., Verstandig, G., Rowe, J.W. et al. Application of machine learning to large in vitro databases to identify drug–cancer cell interactions: azithromycin and KLK6 mutation status. Oncogene 40, 3766–3770 (2021). https://doi.org/10.1038/s41388-021-01807-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01807-4
This article is cited by
-
Azithromycin inhibits glioblastoma angiogenesis in mice via inducing mitochondrial dysfunction and oxidative stress
Cancer Chemotherapy and Pharmacology (2023)
-
Application of machine learning to large in-vitro databases to identify cancer cell characteristics: telomerase reverse transcriptase (TERT) expression
Oncogene (2021)