Abstract
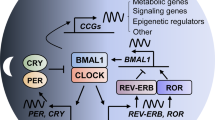
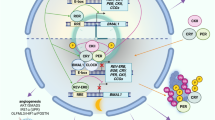
Disruption of the cellular pathway modulating endogenous 24-h rhythms, referred to as “the circadian clock”, has been recently proven to be associated with cancer risk, development, and progression. This pathway operates through a complex network of transcription-translation feedback loops generated by a set of interplaying proteins. The expression of core circadian clock genes is frequently dysregulated in human tumors; however, the specific effects and underlying mechanisms seem to vary depending on the cancer types and are not fully understood. In addition, specific oncogenes may differentially induce the dysregulation of the circadian clock in tumors. Pharmacological modulation of clock components has been shown to result in specific lethality in certain types of cancer cells, and thus holds great promise as a novel anti-cancer therapeutic approach. Here we present an overview of the rationale and current evidence for targeting the clock in cancer treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Roenneberg T, Merrow M. The circadian clock and human health. Curr Biol. 2016;26:R432–43.
Schibler U, Sassone-Corsi P. A web of circadian pacemakers. Cell. 2002;111:919–22.
Takahashi JS. Transcriptional architecture of the mammalian circadian clock. Nat Rev Genet. 2017;18:164–79.
Sulli G, Manoogian ENC, Taub PR, Panda S. Training the circadian clock, clocking the drugs, and drugging the clock to prevent, manage, and treat chronic diseases. Trends Pharmacol Sci. 2018;39:812–27.
Benna C, Helfrich-Forster C, Rajendran S, Monticelli H, Pilati P, Nitti D, et al. Genetic variation of clock genes and cancer risk: a field synopsis and meta-analysis. Oncotarget. 2017;8:23978–95.
Sancar A, Van Gelder RN. Clocks, cancer, and chronochemotherapy. Science. 2021;371:eabb0738.
Panda S, Sato TK, Castrucci AM, Rollag MD, DeGrip WJ, Hogenesch JB, et al. Melanopsin (Opn4) requirement for normal light-induced circadian phase shifting. Science. 2002;298:2213–6.
Mohawk JA, Green CB, Takahashi JS. Central and peripheral circadian clocks in mammals. Annu Rev Neurosci. 2012;35:445–62.
Kondratov RV, Chernov MV, Kondratova AA, Gorbacheva VY, Gudkov AV, Antoch MP. BMAL1-dependent circadian oscillation of nuclear CLOCK: posttranslational events induced by dimerization of transcriptional activators of the mammalian clock system. Genes Dev. 2003;17:1921–32.
Lee C, Etchegaray JP, Cagampang FR, Loudon AS, Reppert SM. Posttranslational mechanisms regulate the mammalian circadian clock. Cell. 2001;107:855–67.
Cao X, Yang Y, Selby CP, Liu Z, Sancar A. Molecular mechanism of the repressive phase of the mammalian circadian clock. Proc Natl Acad Sci USA. 2021;118:e2021174118.
Solt LA, Kojetin DJ, Burris TP. The REV-ERBs and RORs: molecular links between circadian rhythms and lipid homeostasis. Fut Med Chem. 2011;3:623–38.
Yoshitane H, Takao T, Satomi Y, Du NH, Okano T, Fukada Y. Roles of CLOCK phosphorylation in suppression of E-box-dependent transcription. Mol Cell Biol. 2009;29:3675–86.
Luciano AK, Zhou W, Santana JM, Kyriakides C, Velazquez H, Sessa WC. CLOCK phosphorylation by AKT regulates its nuclear accumulation and circadian gene expression in peripheral tissues. J Biol Chem. 2018;293:9126–36.
Liu N, Zhang EE. Phosphorylation regulating the ratio of intracellular CRY1 protein determines the circadian period. Front Neurol. 2016;7:159.
Tamaru T, Hirayama J, Isojima Y, Nagai K, Norioka S, Takamatsu K, et al. CK2alpha phosphorylates BMAL1 to regulate the mammalian clock. Nat Struct Mol Biol. 2009;16:446–8.
Tsuchiya Y, Akashi M, Matsuda M, Goto K, Miyata Y, Node K, et al. Involvement of the protein kinase CK2 in the regulation of mammalian circadian rhythms. Sci Signal. 2009;2:ra26.
Schernhammer ES, Kroenke CH, Laden F, Hankinson SE. Night work and risk of breast cancer. Epidemiology. 2006;17:108–11.
Wegrzyn LR, Tamimi RM, Rosner BA, Brown SB, Stevens RG, Eliassen AH, et al. Rotating night-shift work and the risk of breast cancer in the nurses’ health studies. Am J Epidemiol. 2017;186:532–40.
Walasa WM, Carey RN, Si S, Fritschi L, Heyworth JS, Fernandez RC, et al. Association between shiftwork and the risk of colorectal cancer in females: a population-based case–control study. Occup Environ Med. 2018;75:344–50.
Viswanathan AN, Hankinson SE, Schernhammer ES. Night shift work and the risk of endometrial cancer. Cancer Res. 2007;67:10618–22.
Flynn-Evans EE, Mucci L, Stevens RG, Lockley SW. Shiftwork and prostate-specific antigen in the national health and nutrition examination survey. J Natl Cancer Inst. 2013;105:1292–7.
Straif K, Baan R, Grosse Y, Secretan B, El Ghissassi F, Bouvard V, et al. Carcinogenicity of shift-work, painting, and fire-fighting. Lancet Oncol. 2007;8:1065–6.
Kettner NM, Voicu H, Finegold MJ, Coarfa C, Sreekumar A, Putluri N, et al. Circadian homeostasis of liver metabolism suppresses hepatocarcinogenesis. Cancer Cell. 2016;30:909–24.
Lee S, Donehower LA, Herron AJ, Moore DD, Fu L. Disrupting circadian homeostasis of sympathetic signaling promotes tumor development in mice. PLoS ONE. 2010;5:e10995.
Fu L, Pelicano H, Liu J, Huang P, Lee CC. The circadian gene period2 plays an important role in tumor suppression and DNA damage response in vivo. Cell. 2002;111:41–50.
Papagiannakopoulos T, Bauer MR, Davidson SM, Heimann M, Subbaraj L, Bhutkar A, et al. Circadian rhythm disruption promotes lung tumorigenesis. Cell Metab. 2016;24:324–31.
Pariollaud M, Lamia KA. Cancer in the fourth dimension: what is the impact of circadian disruption? Cancer Discov. 2020;10:1455–64.
Ye Y, Xiang Y, Ozguc FM, Kim Y, Liu CJ, Park PK, et al. The genomic landscape and pharmacogenomic interactions of clock genes in cancer chronotherapy. Cell Syst. 2018;6:314–28. e2
Sulli G, Lam MTY, Panda S. Interplay between circadian clock and cancer: new frontiers for cancer treatment. Trends Cancer. 2019;5:475–94.
de Assis LVM, Kinker GS, Moraes MN, Markus RP, Fernandes PA, Castrucci AML. Expression of the circadian clock gene BMAL1 positively correlates with antitumor immunity and patient survival in metastatic melanoma. Front Oncol. 2018;8:185.
Li W, Liu L, Liu D, Jin S, Yang Y, Tang W, et al. Decreased circadian component Bmal1 predicts tumor progression and poor prognosis in human pancreatic ductal adenocarcinoma. Biochem Biophys Res Commun. 2016;472:156–62.
Ramos CA, Ouyang C, Qi Y, Chung Y, Cheng CT, LaBarge MA, et al. A non-canonical function of BMAL1 metabolically limits obesity-promoted triple-negative breast cancer. iScience. 2020;23:100839.
Fekry B, Ribas-Latre A, Baumgartner C, Deans JR, Kwok C, Patel P, et al. Incompatibility of the circadian protein BMAL1 and HNF4α in hepatocellular carcinoma. Nat Commun. 2018;9:4349.
Taniguchi H, Fernández AF, Setién F, Ropero S, Ballestar E, Villanueva A, et al. Epigenetic inactivation of the circadian clock gene BMAL1 in hematologic malignancies. Cancer Res. 2009;69:8447–54.
Zhang S, Zhang J, Deng Z, Liu H, Mao W, Jiang F, et al. Circadian clock components RORα and Bmal1 mediate the anti-proliferative effect of MLN4924 in osteosarcoma cells. Oncotarget. 2016;7:66087–99.
Yeh CM, Shay J, Zeng TC, Chou JL, Huang TH, Lai HC, et al. Epigenetic silencing of ARNTL, a circadian gene and potential tumor suppressor in ovarian cancer. Int J Oncol. 2014;45:2101–7.
Qu M, Duffy T, Hirota T, Kay SA. Nuclear receptor HNF4A transrepresses CLOCK:BMAL1 and modulates tissue-specific circadian networks. Proc Natl Acad Sci USA. 2018;115:E12305–e12.
Puram RV, Kowalczyk MS, de Boer CG, Schneider RK, Miller PG, McConkey M, et al. Core circadian clock genes regulate leukemia stem cells in AML. Cell 2016;165:303–16.
Dong Z, Zhang G, Qu M, Gimple RC, Wu Q, Qiu Z, et al. Targeting glioblastoma stem cells through disruption of the circadian clock. Cancer Discov. 2019;9:1556–73.
Chen P, Hsu WH, Chang A, Tan Z, Lan Z, Zhou A, et al. Circadian regulator CLOCK recruits immune-suppressive microglia into the GBM tumor microenvironment. Cancer Discov. 2020;10:371–81.
Kinouchi K, Sassone-Corsi P. Metabolic rivalry: circadian homeostasis and tumorigenesis. Nat Rev Cancer. 2020;20:645–61.
Yang MY, Lin PM, Hsiao HH, Hsu JF, Lin HY, Hsu CM, et al. Up-regulation of PER3 Expression Is correlated with better clinical outcome in acute leukemia. Anticancer Res. 2015;35:6615–22.
Cao Q, Gery S, Dashti A, Yin D, Zhou Y, Gu J, et al. A role for the clock gene per1 in prostate cancer. Cancer Res. 2009;69:7619–25.
Gery S, Virk RK, Chumakov K, Yu A, Koeffler HP. The clock gene Per2 links the circadian system to the estrogen receptor. Oncogene. 2007;26:7916–20.
Hwang-Verslues WW, Chang P-H, Jeng Y-M, Kuo W-H, Chiang P-H, Chang Y-C, et al. Loss of corepressor PER2 under hypoxia up-regulates OCT1-mediated EMT gene expression and enhances tumor malignancy. Proc Natl Acad Sci USA. 2013;110:2331–6.
Tong H, Liu X, Li T, Qiu W, Peng C, Shen B, et al. NR1D2 accelerates hepatocellular carcinoma progression by driving the epithelial-to-mesenchymal transition. OncoTargets Ther. 2020;13:3931–42.
Tao L, Yu H, Liang R, Jia R, Wang J, Jiang K, et al. Rev-erbα inhibits proliferation by reducing glycolytic flux and pentose phosphate pathway in human gastric cancer cells. Oncogenesis. 2019;8:57.
Sulli G, Rommel A, Wang X, Kolar MJ, Puca F, Saghatelian A, et al. Pharmacological activation of REV-ERBs is lethal in cancer and oncogene-induced senescence. Nature. 2018;553:351–5.
Yu H, Meng X, Wu J, Pan C, Ying X, Zhou Y, et al. Cryptochrome 1 overexpression correlates with tumor progression and poor prognosis in patients with colorectal cancer. PLoS ONE. 2013;8:e61679.
Shafi AA, McNair CM, McCann JJ, Alshalalfa M, Shostak A, Severson TM, et al. The circadian cryptochrome, CRY1, is a pro-tumorigenic factor that rhythmically modulates DNA repair. Nat Commun. 2021;12:401.
Cadenas C, van de Sandt L, Edlund K, Lohr M, Hellwig B, Marchan R, et al. Loss of circadian clock gene expression is associated with tumor progression in breast cancer. Cell Cycle. 2014;13:3282–91.
Mannic T, Meyer P, Triponez F, Pusztaszeri M, Le Martelot G, Mariani O, et al. Circadian clock characteristics are altered in human thyroid malignant nodules. J Clin Endocrinol Metab. 2013;98:4446–56.
Ozturk N, Lee JH, Gaddameedhi S, Sancar A. Loss of cryptochrome reduces cancer risk in p53 mutant mice. Proc Natl Acad Sci USA. 2009;106:2841–6.
van der Watt PJ, Roden LC, Davis KT, Parker MI, Leaner VD. Circadian oscillations persist in cervical and esophageal cancer cells displaying decreased expression of tumor-suppressing circadian clock genes. Mol Cancer Res. 2020;18:1340–53.
Shostak A, Ruppert B, Ha N, Bruns P, Toprak UH, Eils R, et al. MYC/MIZ1-dependent gene repression inversely coordinates the circadian clock with cell cycle and proliferation. Nat Commun. 2016;7:11807.
Altman BJ, Hsieh AL, Sengupta A, Krishnanaiah SY, Stine ZE, Walton ZE, et al. MYC disrupts the circadian clock and metabolism in cancer cells. Cell Metab. 2015;22:1009–19.
Huber AL, Papp SJ, Chan AB, Henriksson E, Jordan SD, Kriebs A, et al. CRY2 and FBXL3 cooperatively degrade c-MYC. Mol Cell. 2016;64:774–89.
Noel R, Song X, Shin Y, Banerjee S, Kojetin D, Lin L, et al. Synthesis and SAR of tetrahydroisoquinolines as Rev-erbα agonists. Bioorg Med Chem Lett. 2012;22:3739–42.
Solt LA, Wang Y, Banerjee S, Hughes T, Kojetin DJ, Lundasen T, et al. Regulation of circadian behaviour and metabolism by synthetic REV-ERB agonists. Nature. 2012;485:62–8.
Shen W, Zhang W, Ye W, Wang H, Zhang Q, Shen J, et al. SR9009 induces a REV-ERB dependent anti-small-cell lung cancer effect through inhibition of autophagy. Theranostics. 2020;10:4466–80.
Dierickx P, Emmett MJ, Jiang C, Uehara K, Liu M, Adlanmerini M, et al. SR9009 has REV-ERB-independent effects on cell proliferation and metabolism. Proc Natl Acad Sci USA. 2019;116:12147–52.
Kojetin D, Wang Y, Kamenecka TM, Burris TP. Identification of SR8278, a synthetic antagonist of the nuclear heme receptor REV-ERB. ACS Chem Biol. 2011;6:131–4.
De Mei C, Ercolani L, Parodi C, Veronesi M, Lo Vecchio C, Bottegoni G. et al. Dual inhibition of REV-ERBβ and autophagy as a novel pharmacological approach to induce cytotoxicity in cancer cells. Oncogene. 2015;34:2597–608.
Amir M, Chaudhari S, Wang R, Campbell S, Mosure SA, Chopp LB, et al. REV-ERBα regulates T(H)17 cell development and autoimmunity. Cell Rep. 2018;25:3733–49. e8
Wang Y, Kumar N, Nuhant P, Cameron MD, Istrate MA, Roush WR, et al. Identification of SR1078, a synthetic agonist for the orphan nuclear receptors RORα and RORγ. ACS Chem Biol. 2010;5:1029–34.
Wang Y, Solt LA, Kojetin DJ, Burris TP. Regulation of p53 stability and apoptosis by a ROR agonist. PLoS ONE. 2012;7:e34921.
He B, Nohara K, Park N, Park YS, Guillory B, Zhao Z, et al. The small molecule nobiletin targets the molecular oscillator to enhance circadian rhythms and protect against metabolic syndrome. Cell Metab. 2016;23:610–21.
Ashrafizadeh M, Zarrabi A, Saberifar S, Hashemi F, Hushmandi K, Hashemi F, et al. Nobiletin in cancer therapy: how this plant derived-natural compound targets various oncogene and onco-suppressor pathways. Biomedicines. 2020;8:110.
Hirota T, Lee JW, St John PC, Sawa M, Iwaisako K, Noguchi T, et al. Identification of small molecule activators of cryptochrome. Science. 2012;337:1094–7.
Oshima T, Yamanaka I, Kumar A, Yamaguchi J, Nishiwaki-Ohkawa T, Muto K, et al. C-H activation generates period-shortening molecules that target cryptochrome in the mammalian circadian clock. Angew Chem. 2015;54:7193–7.
Lee JW, Hirota T, Kumar A, Kim NJ, Irle S, Kay SA. Development of small-molecule cryptochrome stabilizer derivatives as modulators of the circadian clock. ChemMedChem. 2015;10:1489–97.
Miller S, Son YL, Aikawa Y, Makino E, Nagai Y, Srivastava A, et al. Isoform-selective regulation of mammalian cryptochromes. Nat Chem Biol. 2020;16:676–85.
Miller S, Aikawa Y, Sugiyama A, Nagai Y, Hara A, Oshima T, et al. An isoform-selective modulator of cryptochrome 1 regulates circadian rhythms in mammals. Cell Chem Biol. 2020;27:1192–8. e5
Chun SK, Jang J, Chung S, Yun H, Kim NJ, Jung JW, et al. Identification and validation of cryptochrome inhibitors that modulate the molecular circadian clock. ACS Chem Biol. 2014;9:703–10.
Chun SK, Chung S, Kim HD, Lee JH, Jang J, Kim J, et al. A synthetic cryptochrome inhibitor induces anti-proliferative effects and increases chemosensitivity in human breast cancer cells. Biochem Biophys Res Commun. 2015;467:441–6.
Hirota T, Lee JW, Lewis WG, Zhang EE, Breton G, Liu X, et al. High-throughput chemical screen identifies a novel potent modulator of cellular circadian rhythms and reveals CKIα as a clock regulatory kinase. PLoS Biol. 2010;8:e1000559.
Xiong Y, Zhou L, Su Z, Song J, Sun Q, Liu SS, et al. Longdaysin inhibits Wnt/β-catenin signaling and exhibits antitumor activity against breast cancer. OncoTargets Ther. 2019;12:993–1005.
Lee JW, Hirota T, Ono D, Honma S, Honma KI, Park K, et al. Chemical control of mammalian circadian behavior through dual inhibition of casein kinase Iα and δ. J Med Chem. 2019;62:1989–98.
Mashhoon N, DeMaggio AJ, Tereshko V, Bergmeier SC, Egli M, Hoekstra MF, et al. Crystal structure of a conformation-selective casein kinase-1 inhibitor. J Biol Chem. 2000;275:20052–60.
Behrend L, Milne DM, Stöter M, Deppert W, Campbell LE, Meek DW, et al. IC261, a specific inhibitor of the protein kinases casein kinase 1-delta and -epsilon, triggers the mitotic checkpoint and induces p53-dependent postmitotic effects. Oncogene. 2000;19:5303–13.
Bibian M, Rahaim RJ, Choi JY, Noguchi Y, Schürer S, Chen W, et al. Development of highly selective casein kinase 1δ/1ε (CK1δ/ε) inhibitors with potent antiproliferative properties. Bioorg Med Chem Lett. 2013;23:4374–80.
Rosenberg LH, Lafitte M, Quereda V, Grant W, Chen W, Bibian M, et al. Therapeutic targeting of casein kinase 1δ in breast cancer. Sci Transl Med. 2015;7:318ra202.
Oshima T, Niwa Y, Kuwata K, Srivastava A, Hyoda T, Tsuchiya Y, et al. Cell-based screen identifies a new potent and highly selective CK2 inhibitor for modulation of circadian rhythms and cancer cell growth. Sci Adv. 2019;5:eaau9060.
Lian H, Su M, Zhu Y, Zhou Y, Soomro SH, Fu H. Protein kinase CK2, a potential therapeutic target in carcinoma management. Asian Pac J Cancer Prev. 2019;20:23–32.
Duda P, Akula SM, Abrams SL, Steelman LS, Martelli AM, Cocco L, et al. Targeting GSK3 and associated signaling pathways involved in cancer. Cells. 2020;9:1110.
Hirota T, Lewis WG, Liu AC, Lee JW, Schultz PG, Kay SA. A chemical biology approach reveals period shortening of the mammalian circadian clock by specific inhibition of GSK-3beta. Proc Natl Acad Sci USA. 2008;105:20746–51.
Battaglin F, Cao S, Millstein J, Puccini A, Togunaka R, Naseem M, et al. Polymorphism in the circadian clock pathway to predict outcome in patients (pts) with metastatic colorectal cancer (mCRC): data from TRIBE and FIRE-3 phase III trials. J Clin Oncol. 2018;36(15_suppl):3576.
Burgermeister E, Battaglin F, Eladly F, Wu W, Herweck F, Schulte N, et al. Aryl hydrocarbon receptor nuclear translocator-like (ARNTL/BMAL1) is associated with bevacizumab resistance in colorectal cancer via regulation of vascular endothelial growth factor A. EBioMedicine. 2019;45:139–54.
Qu F, Qiao Q, Wang N, Ji G, Zhao H, He L, et al. Genetic polymorphisms in circadian negative feedback regulation genes predict overall survival and response to chemotherapy in gastric cancer patients. Sci Rep. 2016;6:22424.
Wu Y, Tao B, Zhang T, Fan Y, Mao R. Pan-cancer analysis reveals disrupted circadian clock associates with T cell exhaustion. Front Immunol. 2019;10:2451.
Li Y, Basti A, Yalçin M, Relógio A. Circadian dysregulation of the TGFβ/SMAD4 pathway modulates metastatic properties and cell fate decisions in pancreatic cancer cells. iScience. 2020;23:101551.
Sato F, Bhawal UK, Yoshimura T, Muragaki Y. DEC1 and DEC2 crosstalk between circadian rhythm and tumor progression. J Cancer. 2016;7:153–9.
Westermaier Y, Ruiz-Carmona S, Theret I, Perron-Sierra F, Poissonnet G, Dacquet C, et al. Binding mode prediction and MD/MMPBSA-based free energy ranking for agonists of REV-ERBα/NCoR. J Comput-Aided Mol Des. 2017;31:755–75.
Zou K, Li Z, Zhang Y, Zhang H-Y, Li B, Zhu W-L, et al. Advances in the study of berberine and its derivatives: a focus on anti-inflammatory and anti-tumor effects in the digestive system. Acta Pharm Sin. 2017;38:157–67.
Yuan F, Li D, Guo M, Fang T, Sun J, Qi F, et al. IC261 suppresses progression of hepatocellular carcinoma in a casein kinase 1 δ/ε independent manner. Biochem Biophys Res Commun. 2020;523:809–15.
Zhang Y, Zhang Y, Li M, Meng F, Yu Z, Chen Y, et al. Combination of SB431542, CHIR99021 and PD0325901 has a synergic effect on abrogating valproic acid‑induced epithelial‑mesenchymal transition and stemness in HeLa, 5637 and SCC‑15 cells. Oncol Rep. 2019;41:3545–54.
Kitabayashi T, Dong Y, Furuta T, Sabit H, Jiapaer S, Zhang J, et al. Identification of GSK3β inhibitor kenpaullone as a temozolomide enhancer against glioblastoma. Sci Rep. 2019;9:10049.
Tao NN, Zhang ZZ, Ren JH, Zhang J, Zhou YJ, Wai Wong VK, et al. Overexpression of ubiquitin-conjugating enzyme E2 L3 in hepatocellular carcinoma potentiates apoptosis evasion by inhibiting the GSK3β/p65 pathway. Cancer Lett. 2020;481:1–14.
Duffy DJ, Krstic A, Schwarzl T, Higgins DG, Kolch W. GSK3 inhibitors regulate MYCN mRNA levels and reduce neuroblastoma cell viability through multiple mechanisms, including p53 and Wnt signaling. Mol Cancer Therap. 2014;13:454–67.
Sha Z, Zhou J, Wu Y, Zhang T, Li C, Meng Q, et al. BYSL promotes glioblastoma cell migration, invasion, and mesenchymal transition through the GSK-3β/β-catenin signaling pathway. Front Oncol. 2020;10:565225.
Lo WY, Chang NW. An indirubin derivative, indirubin-3′-monoxime suppresses oral cancer tumorigenesis through the downregulation of survivin. PLoS ONE. 2013;8:e70198.
Acknowledgements
We thank Tsuyoshi Hirota and Jamie Cope for the critical reading of the manuscript.
Funding
Partly supported by the USC/Norris Translational Team Accelerator Project Initiative, the National Cancer Institute (grant numbers P30CA014089, UG1CA180830, R01CA23866201), the National Institute of Neurological Disorders and Stroke (grant number 1F31NS120654-01), the Gloria Borges WunderGlo Foundation, the Gene Gregg Pancreas Research Fund, and the Victoria Wilson Research Fund. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Cancer Institute of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
SAK serves on the board of Synchronicity Pharma and received research support from the company. Other authors declare no potential competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Battaglin, F., Chan, P., Pan, Y. et al. Clocking cancer: the circadian clock as a target in cancer therapy. Oncogene 40, 3187–3200 (2021). https://doi.org/10.1038/s41388-021-01778-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01778-6
This article is cited by
-
Targeted screening and identification of chlorhexidine as a pro-myogenic circadian clock activator
Stem Cell Research & Therapy (2023)
-
Integrated multi-omics analysis reveals the molecular interplay between circadian clocks and cancer pathogenesis
Scientific Reports (2023)
-
The circadian clock as a potential biomarker and therapeutic target in pancreatic cancer
Molecular and Cellular Biochemistry (2023)
-
ROR activation by Nobiletin enhances antitumor efficacy via suppression of IκB/NF-κB signaling in triple-negative breast cancer
Cell Death & Disease (2022)
-
Obesity, cancer risk, and time-restricted eating
Cancer and Metastasis Reviews (2022)