Abstract
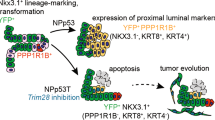
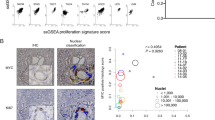
The molecular mechanisms of luminal cell differentiation are not understood well enough to determine how differentiation goes awry during oncogenesis. Using RNA-Seq analysis, we discovered that CREB1 plays a central role in maintaining new luminal cell survival and that oncogenesis dramatically changes the CREB1-induced transcriptome. CREB1 is active in luminal cells, but not basal cells. We identified ING4 and its E3 ligase, JFK, as CREB1 transcriptional targets in luminal cells. During luminal cell differentiation, transient induction of ING4 expression is followed by a peak in CREB1 activity, while JFK increases concomitantly with CREB1 activation. Transient expression of ING4 is required for luminal cell induction; however, failure to properly down-regulate ING4 leads to luminal cell death. Consequently, blocking CREB1 increased ING4 expression, suppressed JFK, and led to luminal cell death. Thus, CREB1 is responsible for the suppression of ING4 required for luminal cell survival and maintenance. Oncogenic transformation by suppressing PTEN resulted in constitutive activation of CREB1. However, the tumor cells could no longer fully differentiate into luminal cells, failed to express ING4, and displayed a unique CREB1 transcriptome. Blocking CREB1 in tumorigenic cells suppressed tumor growth in vivo, rescued ING4 expression, and restored luminal cell formation, but ultimately induced luminal cell death. IHC of primary prostate tumors demonstrated a strong correlation between loss of ING4 and loss of PTEN. This is the first study to define a molecular mechanism whereby oncogenic loss of PTEN, leading to aberrant CREB1 activation, suppresses ING4 expression causing disruption of luminal cell differentiation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Frank SB, Berger PL, Ljungman M, Miranti CK. Human prostate luminal cell differentiation requires NOTCH3 induction by p38-MAPK and MYC. J Cell Sci. 2017;130:1952–64.
Henry GH, Malewska A, Joseph DB, Malladi VS, Lee J, Torrealba J, et al. A cellular anatomy of the normal adult human prostate and prostatic urethra. Cell Rep. 2018;25:3530. e3535
Burger PE, Xiong X, Coetzee S, Salm SN, Moscatelli D, Goto K, et al. Sca-1 expression identifies stem cells in the proximal region of prostatic ducts with high capacity to reconstitute prostatic tissue. Proc Natl Acad Sci USA. 2005;102:7180–5.
Kwon OJ, Zhang L, Xin L. Stem Cell Antigen-1 identifies a distinct androgen-independent murine prostatic luminal cell lineage with bipotent potential. Stem Cells. 2016;34:191–202.
Lamb LE, Knudsen BS, Miranti CK. E-cadherin-mediated survival of androgen-receptor-expressing secretory prostate epithelial cells derived from a stratified in vitro differentiation model. J Cell Sci. 2010;123:266–76.
Xiao GQ, Golestani R, Pham H, Sherrod AE. Stratification of atypical intraepithelial prostatic lesions based on basal cell and architectural patterns. Am J Clin Pathol. 2020;153:407–16.
Abate-Shen C, Shen MM, Gelmann E. Integrating differentiation and cancer: the Nkx3.1 homeobox gene in prostate organogenesis and carcinogenesis. Differentiation. 2008;76:717–27.
Brandi F, Grupp K, Hube-Magg C, Kluth M, Lang D, Minner S, et al. High concordance of TMPRSS-ERG fusion between primary prostate cancer and its lymph node metastases. Oncol Lett. 2018;16:6238–44.
Thangapazham R, Saenz F, Katta S, Mohamed AA, Tan SH, Petrovics G, et al. Loss of the NKX3.1 tumorsuppressor promotes the TMPRSS2-ERG fusion gene expression in prostate cancer. BMC Cancer. 2014;14:16.
Liu W, Xie CC, Thomas CY, Kim ST, Lindberg J, Egevad L, et al. Genetic markers associated with early cancer-specific mortality following prostatectomy. Cancer. 2013;119:2405–12.
Chang H, Jung WY, Kang Y, Lee H, Kim A, Kim BH. Expression of ROR1, pAkt, and pCREB in gastric adenocarcinoma. Ann Diagn Pathol. 2015;19:330–4.
Wang S, Garcia AJ, Wu M, Lawson DA, Witte ON, Wu H. Pten deletion leads to the expansion of a prostatic stem/progenitor cell subpopulation and tumor initiation. Proc Natl Acad Sci USA. 2006;103:1480–5.
Koh CM, Bieberich CJ, Dang CV, Nelson WG, Yegnasubramanian S, De Marzo AM. MYC and prostate cancer. Genes Cancer. 2010;1:617–28.
Berger PL, Frank SB, Schulz VV, Nollet EA, Edick MJ, Holly B, et al. Transient induction of ING4 by Myc drives prostate epithelial cell differentiation and its disruption drives prostate tumorigenesis. Cancer Res. 2014;74:3357–68.
Hung T, Binda O, Champagne KS, Kuo AJ, Johnson K, Chang HY, et al. ING4 mediates crosstalk between histone H3 K4 trimethylation and H3 acetylation to attenuate cellular transformation. Mol Cell. 2009;33:248–56.
Lonze BE, Ginty DD. Function and regulation of CREB family transcription factors in the nervous system. Neuron. 2002;35:605–23.
Shankar DB, Cheng JC, Kinjo K, Federman N, Moore TB, Gill A, et al. The role of CREB as a proto-oncogene in hematopoiesis and in acute myeloid leukemia. Cancer Cell. 2005;7:351–62.
Thway K, Fisher C. Tumors with EWSR1-CREB1 and EWSR1-ATF1 fusions: the current status. Am J Surg Pathol. 2012;36:e1–e11.
Fang Z, Lin A, Chen J, Zhang X, Liu H, Li H, et al. CREB1 directly activates the transcription of ribonucleotide reductase small subunit M2 and promotes the aggressiveness of human colorectal cancer. Oncotarget. 2016;7:78055–68.
Zhang Y, Zheng D, Zhou T, Song H, Hulsurkar M, Su N, et al. Androgen deprivation promotes neuroendocrine differentiation and angiogenesis through CREB-EZH2-TSP1 pathway in prostate cancers. Nat Comm. 2018;9:4080.
Sunkel B, Wu D, Chen Z, Wang CM, Liu X, Ye Z, et al. Integrative analysis identifies targetable CREB1/FoxA1 transcriptional co-regulation as a predictor of prostate cancer recurrence. Nucleic Acids Res. 2016;44:4105–22.
Rehfuss RP, Walton KM, Loriaux MM, Goodman RH. The cAMP-regulated enhancer-binding protein ATF-1 activates transcription in response to cAMP-dependent protein kinase A. J Biol Chem. 1991;266:18431–4.
Gu T, Zhang Z, Wang J, Guo J, Shen WH, Yin Y. CREB is a novel nuclear target of PTEN phosphatase. Cancer Res. 2011;71:2821–5.
Berger PL, Winn ME, Miranti CK. Miz1, a novel target of ING4, can drive prostate luminal epithelial cell differentiation. Prostate. 2017;77:49–59.
Yan J, Jiang J, Lim CA, Wu Q, Ng HH, Chin KC. BLIMP1 regulates cell growth through repression of p53 transcription. Proc Natl Acad Sci USA. 2007;104:1841–6.
Cizmecioglu O, Warnke S, Arnold M, Duensing S, Hoffmann I. Plk2 regulated centriole duplication is dependent on its localization to the centrioles and a functional polo-box domain. Cell Cycle. 2008;7:3548–55.
Wang J, Han X, Zhang Y. Autoregulatory mechanisms of phosphorylation of checkpoint kinase 1. Cancer Res. 2012;72:3786–94.
Kirschner N, Rosenthal R, Furuse M, Moll I, Fromm M, Brandner JM. Contribution of tight junction proteins to ion, macromolecule, and water barrier in keratinocytes. J Investig Dermatol. 2013;133:1161–9.
Vo N, Goodman RH. CREB-binding protein and p300 in transcriptional regulation. J Biol Chem. 2001;276:13505–8.
Yan R, He L, Li Z, Han X, Liang J, Si W, et al. SCF(JFK) is a bona fide E3 ligase for ING4 and a potent promoter of the angiogenesis and metastasis of breast cancer. Genes Dev. 2015;29:672–85.
Martins G, Calame K. Regulation and functions of Blimp-1 in T and B lymphocytes. Ann. Rev Immunol. 2008;26:133–69.
Magnusdottir E, Kalachikov S, Mizukoshi K, Savitsky D, Ishida-Yamamoto A, Panteleyev AA, et al. Epidermal terminal differentiation depends on B lymphocyte-induced maturation protein-1. Proc Natl Acad Sci USA. 2007;104:14988–93.
Mochizuki M, Lorenz V, Ivanek R, Della Verde G, Gaudiello E, Marsano A. et al. Polo-like kinase 2 is dynamically regulated to coordinate proliferation and early lineage specification downstream of Yes-associated protein 1 in cardiac progenitor cells. J Am Heart Assoc. 2017;6:1–19.
Capaldo CT, Nusrat A. Claudin switching: physiological ppasticity of the tight junction. Sem Cell Dev Biol. 2015;42:22–29.
Garcia-Hernandez V, Quiros M, Nusrat A. Intestinal epithelial claudins: expression and regulation in homeostasis and inflammation. Ann NY Acad Sci. 2017;1397:66–79.
Landeira BS, Santana T, Araujo JAM, Tabet EI, Tannous BA, Schroeder T, et al. Activity-independent effects of CREB on neuronal survival and differentiation during mouse cerebral cortex development. Cereb Cortex. 2018;28:538–48.
Bhat NR, Zhang P, Mohanty SB. p38 MAP kinase regulation of oligodendrocyte differentiation with CREB as a potential target. Neurochem Res. 2007;32:293–302.
Jeong SG, Cho GW. The tubulin deacetylase sirtuin-2 regulates neuronal differentiation through the ERK/CREB signaling pathway. Biochem Biophys Res Comm. 2017;482:182–7.
Zhu Q, Gao J, Tian G, Tang Z, Tan Y. Adrenomedullin promotes the odontogenic differentiation of dental pulp stem cells through CREB/BMP2 signaling pathway. Acta Biochim Biophys Sin. 2017;49:609–16.
Kim JH, Kim K, Kim I, Seong S, Lee KB, Kim N. BCAP promotes osteoclast differentiation through regulation of the p38-dependent CREB signaling pathway. Bone. 2018;107:188–95.
Rahman F, Bordignon B, Culerrier R, Peiretti F, Spicuglia S, Djabali M. et al. Ascorbic acid drives the differentiation of mesoderm-derived embryonic stem cells. Involvement of p38 MAPK/CREB and SVCT2 transporter. Mol Nutrit Food Res. 2017;61:1–10.
Cheng JC, Kinjo K, Judelson DR, Chang J, Wu WS, Schmid I, et al. CREB is a critical regulator of normal hematopoiesis and leukemogenesis. Blood. 2008;111:1182–92.
Steven A, Seliger B. Control of CREB expression in tumors: from molecular mechanisms and signal transduction pathways to therapeutic target. Oncotarget. 2016;7:35454–65.
Chhabra A, Fernando H, Watkins G, Mansel RE, Jiang WG. Expression of transcription factor CREB1 in human breast cancer and its correlation with prognosis. Oncol Rep. 2007;18:953–8.
Melnikova VO, Dobroff AS, Zigler M, Villares GJ, Braeuer RR, Wang H, et al. CREB inhibits AP-2α expression to regulate the malignant phenotype of melanoma. PloS ONE. 2010;5:e12452.
Garcia GE, Nicole A, Bhaskaran S, Gupta A, Kyprianou N, Kumar AP. Akt-and CREB-mediated prostate cancer cell proliferation inhibition by Nexrutine, a Phellodendron amurense extract. Neoplasia. 2006;8:523–33.
Doyon Y, Cayrou C, Ullah M, Landry AJ, Cote V, Selleck W, et al. ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol Cell. 2006;21:51–64.
Lalonde ME, Avvakumov N, Glass KC, Joncas FH, Saksouk N, Holliday M, et al. Exchange of associated factors directs a switch in HBO1 acetyltransferase histone tail specificity. Genes Dev. 2013;27:2009–24.
Ruan K, Yamamoto TG, Asakawa H, Chikashige Y, Kimura H, Masukata H, et al. Histone H4 acetylation required for chromatin decompaction during DNA replication. Sci Rep. 2015;5:12720.
Palacios A, Moreno A, Oliveira BL, Rivera T, Prieto J, Garcia P, et al. The dimeric structure and the bivalent recognition of H3K4me3 by the tumor suppressor ING4 suggests a mechanism for enhanced targeting of the HBO1 complex to chromatin. J Mol Biol. 2010;396:1117–27.
Edick MJ, Tesfay L, Lamb LE, Knudsen BS, Miranti CK. Inhibition of integrin-mediated crosstalk with epidermal growth factor receptor/Erk or Src signaling pathways in autophagic prostate epithelial cells induces caspase-independent death. Mol Biol Cell. 2007;18:2481–90.
Gmyrek GA, Walburg M, Webb CP, Yu HM, You X, Vaughan ED, et al. Normal and malignant prostate epithelial cells differ in their response to hepatocyte growth factor/scatter factor. Am J Pathol. 2001;159:579–90.
Frank SB, Schulz VV, Miranti CK. A streamlined method for the design and cloning of shRNAs into an optimized Dox-inducible lentiviral vector. BMC Biotechnol. 2017;17:24.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–8.
Acknowledgements
We wish to thank the bioinformatics and pathology cores at the Van Andel Institute and University of Arizona (BBSR, TACMSR), and the experimental mouse core (EMSR) at University of Arizona for all their help with this study. These studies were supported by funding from the Department of Defense, W81XWH-14-1-0479 (MJW, PLB, CKM) and W81XWH-17-1-0570 (SBF, KB, LT, CKM) and funding from the Van Andel Research Institute. Cores at the University of Arizona were supported by funds from NIH/NCI P30 CA023074.
Funding
MJW, PLB, KB, SBF, TL, SSG CKM – 2 grants from DOD; GH, MW – VAI.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Watson, M.J., Berger, P.L., Banerjee, K. et al. Aberrant CREB1 activation in prostate cancer disrupts normal prostate luminal cell differentiation. Oncogene 40, 3260–3272 (2021). https://doi.org/10.1038/s41388-021-01772-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01772-y
This article is cited by
-
miR-125b-5p upregulation by TRIM28 induces cisplatin resistance in non-small cell lung cancer through CREB1 inhibition
BMC Pulmonary Medicine (2022)
-
CRISPR/Cas9-mediated deletion of Interleukin-30 suppresses IGF1 and CXCL5 and boosts SOCS3 reducing prostate cancer growth and mortality
Journal of Hematology & Oncology (2022)
-
CircEZH2/miR-133b/IGF2BP2 aggravates colorectal cancer progression via enhancing the stability of m6A-modified CREB1 mRNA
Molecular Cancer (2022)