Abstract
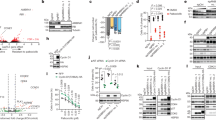
Mdm2 antagonizes the tumor suppressor p53. Targeting the Mdm2-p53 interaction represents an attractive approach for the treatment of cancers with functional p53. Investigating mechanisms underlying Mdm2-p53 regulation is therefore important. The scaffold protein β-arrestin2 (β-arr2) regulates tumor suppressor p53 by counteracting Mdm2. β-arr2 nucleocytoplasmic shuttling displaces Mdm2 from the nucleus to the cytoplasm resulting in enhanced p53 signaling. β-arr2 is constitutively exported from the nucleus, via a nuclear export signal, but mechanisms regulating its nuclear entry are not completely elucidated. β-arr2 can be SUMOylated, but no information is available on how SUMO may regulate β-arr2 nucleocytoplasmic shuttling. While we found β-arr2 SUMOylation to be dispensable for nuclear import, we identified a non-covalent interaction between SUMO and β-arr2, via a SUMO interaction motif (SIM), that is required for β-arr2 cytonuclear trafficking. This SIM promotes association of β-arr2 with the multimolecular RanBP2/RanGAP1-SUMO nucleocytoplasmic transport hub that resides on the cytoplasmic filaments of the nuclear pore complex. Depletion of RanBP2/RanGAP1-SUMO levels result in defective β-arr2 nuclear entry. Mutation of the SIM inhibits β-arr2 nuclear import, its ability to delocalize Mdm2 from the nucleus to the cytoplasm and enhanced p53 signaling in lung and breast tumor cell lines. Thus, a β-arr2 SIM nuclear entry checkpoint, coupled with active β-arr2 nuclear export, regulates its cytonuclear trafficking function to control the Mdm2-p53 signaling axis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Enslen H, Lima-Fernandes E, Scott MG. Arrestins as regulatory hubs in cancer signalling pathways. Handb Exp Pharmacol. 2014;219:405–25.
Laporte SA, Scott MGH. beta-Arrestins: multitask scaffolds orchestrating the where and when in cell signalling. Methods Mol Biol. 2019;1957:9–55.
Peterson YK, Luttrell LM. The diverse roles of arrestin scaffolds in G protein-coupled receptor signaling. Pharmacol Rev. 2017;69:256–97.
Luttrell LM, Roudabush FL, Choy EW, Miller WE, Field ME, Pierce KL, et al. Activation and targeting of extracellular signal-regulated kinases by beta-arrestin scaffolds. Proc Natl Acad Sci USA. 2001;98:2449–54.
Luttrell LM, Wang J, Plouffe B, Smith JS, Yamani L, Kaur S. et al. Manifold roles of beta-arrestins in GPCR signaling elucidated with siRNA and CRISPR/Cas9. Sci Signal. 2018;11:eaat7650.
McDonald PH, Chow CW, Miller WE, Laporte SA, Field ME, Lin FT, et al. Beta-arrestin 2: a receptor-regulated MAPK scaffold for the activation of JNK3. Science. 2000;290:1574–7.
Boularan C, Scott MG, Bourougaa K, Bellal M, Esteve E, Thuret A, et al. beta-arrestin 2 oligomerization controls the Mdm2-dependent inhibition of p53. Proc Natl Acad Sci USA. 2007;104:18061–6.
Shenoy SK, McDonald PH, Kohout TA, Lefkowitz RJ. Regulation of receptor fate by ubiquitination of activated beta 2-adrenergic receptor and beta-arrestin. Science. 2001;294:1307–13.
Wang P, Wu Y, Ge X, Ma L, Pei G. Subcellular localization of beta-arrestins is determined by their intact N domain and the nuclear export signal at the C terminus. J Biol Chem. 2003;278:11648–53.
Javadi A, Deevi RK, Evergren E, Blondel-Tepaz E, Baillie GS, Scott MG. et al. PTEN controls glandular morphogenesis through a juxtamembrane beta-Arrestin1/ARHGAP21 scaffolding complex. Elife. 2017;6:e24578.
Lima-Fernandes E, Enslen H, Camand E, Kotelevets L, Boularan C, Achour L, et al. Distinct functional outputs of PTEN signalling are controlled by dynamic association with beta-arrestins. EMBO J. 2011;30:2557–68.
Lima-Fernandes E, Misticone S, Boularan C, Paradis JS, Enslen H, Roux PP, et al. A biosensor to monitor dynamic regulation and function of tumour suppressor PTEN in living cells. Nat Commun. 2014;5:4431.
Alexander RA, Lot I, Saha K, Abadie G, Lambert M, Decosta E. et al. Beta-arrestins operate an on/off control switch for focal adhesion kinase activity. Cell Mol Life Sci. 2020;77:5259–79.
Scott MG, Le Rouzic E, Perianin A, Pierotti V, Enslen H, Benichou S, et al. Differential nucleocytoplasmic shuttling of beta-arrestins. Characterization of a leucine-rich nuclear export signal in beta-arrestin2. J Biol Chem. 2002;277:37693–701.
Wyatt D, Malik R, Vesecky AC, Marchese A. Small ubiquitin-like modifier modification of arrestin-3 regulates receptor trafficking. J Biol Chem. 2011;286:3884–93.
Xiao N, Li H, Mei W, Cheng J. SUMOylation attenuates human beta-arrestin 2 inhibition of IL-1R/TRAF6 signaling. J Biol Chem. 2015;290:1927–35.
Flotho A, Melchior F. Sumoylation: a regulatory protein modification in health and disease. Annu Rev Biochem. 2013;82:357–85.
Geiss-Friedlander R, Melchior F. Concepts in sumoylation: a decade on. Nat Rev Mol Cell Biol. 2007;8:947–56.
Hay RT. SUMO: a history of modification. Mol Cell. 2005;18:1–12.
Rodriguez MS, Dargemont C, Hay RT. SUMO-1 conjugation in vivo requires both a consensus modification motif and nuclear targeting. J Biol Chem. 2001;276:12654–9.
Guo D, Li M, Zhang Y, Yang P, Eckenrode S, Hopkins D, et al. A functional variant of SUMO4, a new I kappa B alpha modifier, is associated with type 1 diabetes. Nat Genet. 2004;36:837–41.
Kerscher O. SUMO junction-what’s your function? New insights through SUMO-interacting motifs. EMBO Rep. 2007;8:550–5.
Jakobs A, Koehnke J, Himstedt F, Funk M, Korn B, Gaestel M, et al. Ubc9 fusion-directed SUMOylation (UFDS): a method to analyze function of protein SUMOylation. Nat Methods. 2007;4:245–50.
Beauclair G, Bridier-Nahmias A, Zagury JF, Saib A, Zamborlini A. JASSA: a comprehensive tool for prediction of SUMOylation sites and SIMs. Bioinformatics. 2015;31:3483–91.
Kudo N, Matsumori N, Taoka H, Fujiwara D, Schreiner EP, Wolff B, et al. Leptomycin B inactivates CRM1/exportin 1 by covalent modification at a cysteine residue in the central conserved region. Proc Natl Acad Sci USA. 1999;96:9112–7.
Namkung Y, Le Gouill C, Lukashova V, Kobayashi H, Hogue M, Khoury E, et al. Monitoring G protein-coupled receptor and beta-arrestin trafficking in live cells using enhanced bystander BRET. Nat Commun. 2016;7:12178.
Sakin V, Richter SM, Hsiao HH, Urlaub H, Melchior F. Sumoylation of the GTPase Ran by the RanBP2 SUMO E3 Ligase Complex. J Biol Chem. 2015;290:23589–602.
Walde S, Thakar K, Hutten S, Spillner C, Nath A, Rothbauer U, et al. The nucleoporin Nup358/RanBP2 promotes nuclear import in a cargo- and transport receptor-specific manner. Traffic. 2012;13:218–33.
Mahajan R, Delphin C, Guan T, Gerace L, Melchior F. A small ubiquitin-related polypeptide involved in targeting RanGAP1 to nuclear pore complex protein RanBP2. Cell. 1997;88:97–107.
Matunis MJ, Wu J, Blobel G. SUMO-1 modification and its role in targeting the Ran GTPase-activating protein, RanGAP1, to the nuclear pore complex. J Cell Biol. 1998;140:499–509.
Wu J, Matunis MJ, Kraemer D, Blobel G, Coutavas E. Nup358, a cytoplasmically exposed nucleoporin with peptide repeats, Ran-GTP binding sites, zinc fingers, a cyclophilin A homologous domain, and a leucine-rich region. J Biol Chem. 1995;270:14209–13.
Yokoyama N, Hayashi N, Seki T, Pante N, Ohba T, Nishii K, et al. A giant nucleopore protein that binds Ran/TC4. Nature. 1995;376:184–8.
Hashizume C, Kobayashi A, Wong RW. Down-modulation of nucleoporin RanBP2/Nup358 impaired chromosomal alignment and induced mitotic catastrophe. Cell Death Dis. 2013;4:e854.
Monteith JA, Mellert H, Sammons MA, Kuswanto LA, Sykes SM, Resnick-Silverman L, et al. A rare DNA contact mutation in cancer confers p53 gain-of-function and tumor cell survival via TNFAIP8 induction. Mol Oncol. 2016;10:1207–20.
Gonzalez-Santamaria J, Campagna M, Garcia MA, Marcos-Villar L, Gonzalez D, Gallego P, et al. Regulation of vaccinia virus E3 protein by small ubiquitin-like modifier proteins. J Virol. 2011;85:12890–900.
Li R, Wang L, Liao G, Guzzo CM, Matunis MJ, Zhu H, et al. SUMO binding by the Epstein-Barr virus protein kinase BGLF4 is crucial for BGLF4 function. J Virol. 2012;86:5412–21.
Brandariz-Nunez A, Roa A, Valle-Casuso JC, Biris N, Ivanov D, Diaz-Griffero F. Contribution of SUMO-interacting motifs and SUMOylation to the antiretroviral properties of TRIM5alpha. Virology. 2013;435:463–71.
Wang PY, Hsu PI, Wu DC, Chen TC, Jarman AP, Powell LM. et al. SUMOs mediate the nuclear transfer of p38 and p-p38 during helicobacter pylori infection. Int J Mol Sci. 2018;19:2482
Lin DY, Huang YS, Jeng JC, Kuo HY, Chang CC, Chao TT, et al. Role of SUMO-interacting motif in Daxx SUMO modification, subnuclear localization, and repression of sumoylated transcription factors. Mol Cell. 2006;24:341–54.
Werner A, Flotho A, Melchior F. The RanBP2/RanGAP1*SUMO1/Ubc9 complex is a multisubunit SUMO E3 ligase. Mol Cell. 2012;46:287–98.
Hutten S, Flotho A, Melchior F, Kehlenbach RH. The Nup358-RanGAP complex is required for efficient importin alpha/beta-dependent nuclear import. Mol Biol Cell. 2008;19:2300–10.
Hutten S, Walde S, Spillner C, Hauber J, Kehlenbach RH. The nuclear pore component Nup358 promotes transportin-dependent nuclear import. J Cell Sci. 2009;122:1100–10.
Zhang X, Min X, Wang S, Sun N, Kim KM. Mdm2-mediated ubiquitination of beta-arrestin2 in the nucleus occurs in a Gbetagamma- and clathrin-dependent manner. Biochem Pharm. 2020;178:114049.
Singh R, Brewer MK, Mashburn CB, Lou D, Bondada V, Graham B, et al. Calpain 5 is highly expressed in the central nervous system (CNS), carries dual nuclear localization signals, and is associated with nuclear promyelocytic leukemia protein bodies. J Biol Chem. 2014;289:19383–94.
Bolger GB, Baillie GS, Li X, Lynch MJ, Herzyk P, Mohamed A, et al. Scanning peptide array analyses identify overlapping binding sites for the signalling scaffold proteins, beta-arrestin and RACK1, in cAMP-specific phosphodiesterase PDE4D5. Biochem J. 2006;398:23–36.
Li X, Vadrevu S, Dunlop A, Day J, Advant N, Troeger J, et al. Selective SUMO modification of cAMP-specific phosphodiesterase-4D5 (PDE4D5) regulates the functional consequences of phosphorylation by PKA and ERK. Biochem J. 2010;428:55–65.
Blondel-Tepaz E, Guilbert T, Scott MGH. Methods to investigate the nucleocytoplasmic shuttling properties of beta-arrestins. Methods Mol Biol. 2019;1957:251–69.
Paradis JS, Ly S, Blondel-Tepaz E, Galan JA, Beautrait A, Scott MG, et al. Receptor sequestration in response to beta-arrestin-2 phosphorylation by ERK1/2 governs steady-state levels of GPCR cell-surface expression. Proc Natl Acad Sci USA. 2015;112:E5160–8.
Acknowledgements
We thank Dr. A. Benmerah for helpful discussion, Dr. J. Liotard for excellent technical assistance, and the Institut Cochin Imaging (IMAG’IC) and Sequencing platforms (GENOM’IC). The Institut Cochin lab is part of the Who am I? laboratory of excellence (grant ANR-11-LABX-0071), funded by the “Investments for the Future” program operated by The French National Research Agency (grant ANR-11-IDEX-0005-01). This work was funded by grants from the Fondation ARC pour la Recherche sur le Cancer (“Projet ARC” to MGHS), Ligue contre le Cancer (to MGHS), Royal Society (“International Joint Project Scheme” to MGHS and GSB), France Canada Research Fund (to MGHS and SA), CNRS, and INSERM. The work in the laboratory of M.B. was supported by a CIHR Foundation (FDN148431) grant. M.B. holds the Canada Research Chair in Signal transduction and Molecular Pharmacology. EBT was funded by MESR and Fondation ARC pour la Recherche sur le Cancer doctoral fellowships.
Author information
Authors and Affiliations
Contributions
EBT, ML, BS, JSP, GSB, HE, MB, SA, SM, and MGHS designed research. EBT, ML, JSP, BS, MK, KS, JF, AP, and MGHS performed research. EBT, ML, BS, JSP, CA, ELF, AZ, LG, GSB, HE, MB, SA, SM, and MGHS analyzed data. MGHS supervised the project. EBT and MGHS wrote the paper, which was subsequently reviewed by all other authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Blondel-Tepaz, E., Leverve, M., Sokrat, B. et al. The RanBP2/RanGAP1-SUMO complex gates β-arrestin2 nuclear entry to regulate the Mdm2-p53 signaling axis. Oncogene 40, 2243–2257 (2021). https://doi.org/10.1038/s41388-021-01704-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01704-w
This article is cited by
-
METTL3-mediated RanGAP1 promotes colorectal cancer progression through the MAPK pathway by recruiting YTHDF1
Cancer Gene Therapy (2024)
-
Comprehensive genomic analysis of primary malignant melanoma of the esophagus reveals similar genetic patterns compared with epithelium-associated melanomas
Modern Pathology (2022)