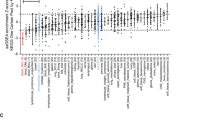
Abstract
Most of the drugs currently prescribed for cancer treatment are riddled with substantial side effects. In order to develop more effective and specific strategies to treat cancer, it is of importance to understand the biology of drug targets, particularly the newly emerging ones. A comprehensive evaluation of these targets will benefit drug development with increased likelihood for success in clinical trials. The folate-mediated one-carbon (1C) metabolism pathway has drawn renewed attention as it is often hyperactivated in cancer and inhibition of this pathway displays promise in developing anticancer treatment with fewer side effects. Here, we systematically review individual enzymes in the 1C pathway and their compartmentalization to mitochondria and cytosol. Based on these insight, we conclude that (1) except the known 1C targets (DHFR, GART, and TYMS), MTHFD2 emerges as good drug target, especially for treating hematopoietic cancers such as CLL, AML, and T-cell lymphoma; (2) SHMT2 and MTHFD1L are potential drug targets; and (3) MTHFD2L and ALDH1L2 should not be considered as drug targets. We highlight MTHFD2 as an excellent therapeutic target and SHMT2 as a complementary target based on structural/biochemical considerations and up-to-date inhibitor development, which underscores the perspectives of their therapeutic potential.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pavlova NN, Thompson CB. The emerging hallmarks of cancer metabolism. Cell Metab. 2016;23:27–47.
Vander Heiden MG. Targeting cancer metabolism: a therapeutic window opens. Nat Rev Drug Discov. 2011;10:671–84.
McGuirk S, Audet-Delage Y, St-Pierre J. Metabolic fitness and plasticity in cancer progression. Trends Cancer. 2020;6:49–61.
Kreuzaler P, Panina Y, Segal J, Yuneva M. Adapt and conquer: metabolic flexibility in cancer growth, invasion and evasion. Mol Metab. 2020;33:83–101.
Amelio I, Cutruzzolá F, Antonov A, Agostini M, Melino G. Serine and glycine metabolism in cancer. Trends Biochem Sci. 2014;39:191–8.
Locasale JW. Serine, glycine and one-carbon units: cancer metabolism in full circle. Nat Rev Cancer. 2013;13:572–83.
Choi Y-K, Park K-G. Targeting glutamine metabolism for cancer treatment. Biomol Ther. 2018;26:19–28.
Fadaka A, Ajiboye B, Ojo O, Adewale O, Olayide I, Emuowhochere R. Biology of glucose metabolization in cancer cells. J Oncological Sci. 2017;3:45–51.
Shiratori R, Furuichi K, Yamaguchi M, Miyazaki N, Aoki H, Chibana H, et al. Glycolytic suppression dramatically changes the intracellular metabolic profile of multiple cancer cell lines in a mitochondrial metabolism-dependent manner. Sci Rep. 2019;9:18699.
Lin J, Xia L, Liang J, Han Y, Wang H, Oyang L, et al. The roles of glucose metabolic reprogramming in chemo- and radio-resistance. J Exp Clin Cancer Res. 2019;38:218.
Farber S, Diamond LK, Mercer RD, Sylvester RF, Wolff JA. Temporary remissions in acute leukemia in children produced by folic acid antagonist, 4-aminopteroyl-glutamic acid (aminopterin). N Engl J Med. 1948;238:787–93.
Chattopadhyay S, Moran RG, Goldman ID. Pemetrexed: biochemical and cellular pharmacology, mechanisms, and clinical applications. Mol Cancer Ther. 2007;6:404–17.
Estrada A, Wright DL, Anderson AC. Antibacterial antifolates: from development through resistance to the next generation. Cold Spring Harb Perspect Med. 2016;6. https://doi.org/10.1101/cshperspect.a028324.
Purcell WT, Ettinger DS. Novel antifolate drugs. Curr Oncol Rep. 2003;5:114–25.
Carver JR, Szalda D, Ky B. Asymptomatic cardiac toxicity in long-term cancer survivors: defining the population and recommendations for surveillance. Semin Oncol. 2013;40:229–38.
Yeh Edward TH, Tong Ann T, Lenihan Daniel J, Wamique YusufS, Joseph Swafford, Christopher Champion, et al. Cardiovascular complications of cancer therapy. Circulation. 2004;109:3122–31.
Wang Y, Probin V, Zhou D. Cancer therapy-induced residual bone marrow injury-mechanisms of induction and implication for therapy. Curr Cancer Ther Rev. 2006;2:271–9.
Appling DR. Compartmentation of folate-mediated one-carbon metabolism in eukaryotes. FASEB J.1991;5(12):2645–51.
Toriello HV. Folic acid and neural tube defects. Genet Med. 2005;7:283–4.
Kujovich JL. Evaluation of anemia. Obstet Gynecol Clin North Am. 2016;43:247–64.
Fox JT, Stover PJ. Folate-mediated one-carbon metabolism. Vitam Horm. 2008;79:1–44.
Ducker GS, Rabinowitz JD. One-carbon metabolism in health and disease. Cell Metab. 2017;25:27–42.
Agapakis CM, Boyle PM, Silver PA. Natural strategies for the spatial optimization of metabolism in synthetic biology. Nat Chem Biol. 2012;8:527–35.
Kit S. The biosynthesis of free glycine and serine by tumors. Cancer Res. 1955;15:715–8.
Labuschagne CF, van den Broek NJF, Mackay GM, Vousden KH, Maddocks ODK. Serine, but not glycine, supports one-carbon metabolism and proliferation of cancer cells. Cell Rep. 2014;7:1248–58.
Shin M, Bryant JD, Momb J, Appling DR. Mitochondrial MTHFD2L is a dual redox cofactor-specific methylenetetrahydrofolate dehydrogenase/methenyltetrahydrofolate cyclohydrolase expressed in both adult and embryonic tissues. J Biol Chem. 2014;289:15507–17.
Cybulski RL, Fisher RR. Intramitochondrial localization and proposed metabolic significance of serine transhydroxymethylase. Biochemistry. 1976;15:3183–7.
Balut C, vandeVen M, Despa S, Lambrichts I, Ameloot M, Steels P, et al. Measurement of cytosolic and mitochondrial pH in living cells during reversible metabolic inhibition. Kidney Int. 2008;73:226–32.
Boron WF. Regulation of intracellular pH. Adv Physiol Educ. 2004;28:160–79.
Shirmanova MV, Druzhkova IN, Lukina MM, Matlashov ME, Belousov VV, Snopova LB, et al. Intracellular pH imaging in cancer cells in vitro and tumors in vivo using the new genetically encoded sensor SypHer2. Biochim Biophys Acta (BBA) - Gen Subj. 2015;1850:1905–11.
White KA, Grillo-Hill BK, Barber DL. Cancer cell behaviors mediated by dysregulated pH dynamics at a glance. J Cell Sci. 2017;130:663–9.
Hu Y, Li Y. Effect of low pH treatment on cell cycle and cell growth. FASEB J. 2018;32:804.49.
Vazquez A, Tedeschi PM, Bertino JR. Overexpression of the mitochondrial folate and glycine–serine pathway: a new determinant of methotrexate selectivity in tumors. Cancer Res. 2013;73:478–82.
Noguchi K, Konno M, Koseki J, Nishida N, Kawamoto K, Yamada D, et al. The mitochondrial one-carbon metabolic pathway is associated with patient survival in pancreatic cancer. Oncol Lett. 2018;16:1827–34.
Nilsson R, Jain M, Madhusudhan N, Sheppard NG, Strittmatter L, Kampf C, et al. Metabolic enzyme expression highlights a key role for MTHFD2 and the mitochondrial folate pathway in cancer. Nat Commun. 2014;5:3128.
Agarwal S, Behring M, Hale K, Al Diffalha S, Wang K, Manne U, et al. MTHFD1L, a folate cycle enzyme, is involved in progression of colorectal cancer. Transl Oncol. 2019;12:1461–7.
Eich M-L, Rodriguez Pena MDC, Chandrashekar DS, Chaux A, Agarwal S, Gordetsky JB, et al. Expression and role of methylenetetrahydrofolate dehydrogenase 1 like (MTHFD1L) in bladder cancer. Transl Oncol. 2019;12:1416–24.
Lee D, Xu IM-J, Chiu DK-C, Lai RK-H, Tse AP-W, Lan Li L, et al. Folate cycle enzyme MTHFD1L confers metabolic advantages in hepatocellular carcinoma. J Clin Investig. 2017;127:1856–72.
Wu M, Wanggou S, Li X, Liu Q, Xie Y. Overexpression of mitochondrial serine hydroxyl-methyltransferase 2 is associated with poor prognosis and promotes cell proliferation and invasion in gliomas. Onco Targets Ther. 2017;10:3781–8.
Tedeschi PM, Vazquez A, Kerrigan JE, Bertino JR. Mitochondrial methylenetetrahydrofolate dehydrogenase (MTHFD2) overexpression is associated with tumor cell proliferation and is a novel target for drug development. Mol Cancer Res. 2015;13:1361–6.
Liu Y, Yin C, Deng M-M, Wang Q, He X-Q, Li M-T, et al. High expression of SHMT2 is correlated with tumor progression and predicts poor prognosis in gastrointestinal tumors. Eur Rev Med Pharm Sci. 2019;23:9379–92.
Di Pietro E, Sirois J, Tremblay ML, MacKenzie RE. Mitochondrial NAD-dependent methylenetetrahydrofolate dehydrogenase-methenyltetrahydrofolate cyclohydrolase is essential for embryonic development. Mol Cell Biol. 2002;22:4158–66.
Stover PJ, Chen LH, Suh JR, Stover DM, Keyomarsi K, Shane B. Molecular cloning, characterization, and regulation of the human mitochondrial serine hydroxymethyltransferase gene. J Biol Chem. 1997;272:1842–8.
Florio R, di Salvo ML, Vivoli M, Contestabile R. Serine hydroxymethyltransferase: a model enzyme for mechanistic, structural, and evolutionary studies. Biochim Biophys Acta. 2011;1814:1489–96.
Giardina G, Brunotti P, Fiascarelli A, Cicalini A, Costa MGS, Buckle AM, et al. How pyridoxal 5′-phosphate differentially regulates human cytosolic and mitochondrial serine hydroxymethyltransferase oligomeric state. FEBS J. 2015;282:1225–41.
Yang X, Wang Z, Li X, Liu B, Liu M, Liu L, et al. SHMT2 Desuccinylation by SIRT5 drives cancer cell proliferation. Cancer Res. 2018;78:372–86.
Morscher RJ, Ducker GS, Li SH-J, Mayer JA, Gitai Z, Sperl W, et al. Mitochondrial translation requires folate-dependent tRNA methylation. Nature. 2018;554:128–32.
Lucas S, Chen G, Aras S, Wang J. Serine catabolism is essential to maintain mitochondrial respiration in mammalian cells. Life Sci Alliance. 2018;1:e201800036.
Anderson DD, Stover PJ. SHMT1 and SHMT2 are functionally redundant in nuclear de novo thymidylate biosynthesis. PLoS ONE. 2009;4:e5839.
Guiducci G, Paone A, Tramonti A, Giardina G, Rinaldo S, Bouzidi A, et al. The moonlighting RNA-binding activity of cytosolic serine hydroxymethyltransferase contributes to control compartmentalization of serine metabolism. Nucleic Acids Res. 2019;47:4240–54.
Anderson DD, Eom JY, Stover PJ. Competition between sumoylation and ubiquitination of serine hydroxymethyltransferase 1 determines its nuclear localization and its accumulation in the nucleus. J Biol Chem. 2012;287:4790–9.
Jain M, Nilsson R, Sharma S, Madhusudhan N, Kitami T, Souza AL, et al. Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation. Science. 2012;336:1040–4.
Lee GY, Haverty PM, Li L, Kljavin NM, Bourgon R, Lee J, et al. Comparative oncogenomics identifies PSMB4 and SHMT2 as potential cancer driver genes. Cancer Res. 2014;74:3114–26.
Tani H, Ohnishi S, Shitara H, Mito T, Yamaguchi M, Yonekawa H, et al. Mice deficient in the Shmt2 gene have mitochondrial respiration defects and are embryonic lethal. Sci Rep. 2018;8:425.
Minton DR, Nam M, McLaughlin DJ, Shin J, Bayraktar EC, Alvarez SW, et al. Serine catabolism by SHMT2 is required for proper mitochondrial translation initiation and maintenance of formylmethionyl-tRNAs. Mol Cell. 2018;69:610–21.e5.
Ning S, Ma S, Saleh AQ, Guo L, Zhao Z, Chen Y. SHMT2 overexpression predicts poor prognosis in intrahepatic cholangiocarcinoma. Gastroenterol Res Pract. 2018;2018:4369253.
Wang H, Chong T, Li B-Y, Chen X-S, Zhen W-B. Evaluating the clinical significance of SHMT2 and its co-expressed gene in human kidney cancer. Biol Res. 2020;53:46.
Woo CC, Chen WC, Teo XQ, Radda GK, Teck Hock Lee P. Downregulating serine hydroxymethyltransferase 2 (SHMT2) suppresses tumorigenesis in human hepatocellular carcinoma. Oncotarget. 2016;7:53005–17.
Nonaka H, Nakanishi Y, Kuno S, Ota T, Mochidome K, Saito Y, et al. Design strategy for serine hydroxymethyltransferase probes based on retro-aldol-type reaction. Nat Commun. 2019;10:876.
Snell K, Riches D. Effects of a triazine antifolate (NSC 127755) on serine hydroxymethyltransferase in myeloma cells in culture. Cancer Lett. 1989;44:217–20.
Stover P, Schirch V. 5-Formyltetrahydrofolate polyglutamates are slow tight binding inhibitors of serine hydroxymethyltransferase. J Biol Chem. 1991;266:1543–50.
Marani M, Paone A, Fiascarelli A, Macone A, Gargano M, Rinaldo S, et al. A pyrazolopyran derivative preferentially inhibits the activity of human cytosolic serine hydroxymethyltransferase and induces cell death in lung cancer cells. Oncotarget. 2016;7:4570–83.
Gu Y, Si J, Xiao X, Tian Y, Yang S. miR-92a inhibits proliferation and induces apoptosis by regulating methylenetetrahydrofolate dehydrogenase 2 (MTHFD2) expression in acute myeloid leukemia. Oncol Res. 2017;25:1069–79.
Koseki J, Konno M, Asai A, Colvin H, Kawamoto K, Nishida N, et al. Enzymes of the one-carbon folate metabolism as anticancer targets predicted by survival rate analysis. Sci Rep. 2018;8:303.
Yan Y, Zhang D, Lei T, Zhao C, Han J, Cui J, et al. MicroRNA-33a-5p suppresses colorectal cancer cell growth by inhibiting MTHFD2. Clin Exp Pharm Physiol. 2019;46:928–36.
Ju H-Q, Lu Y-X, Chen D-L, Zuo Z-X, Liu Z-X, Wu Q-N, et al. Modulation of redox homeostasis by inhibition of MTHFD2 in colorectal cancer: mechanisms and therapeutic implications. J Natl Cancer Inst. 2018. https://doi.org/10.1093/jnci/djy160.
Xu T, Zhang K, Shi J, Huang B, Wang X, Qian K, et al. MicroRNA-940 inhibits glioma progression by blocking mitochondrial folate metabolism through targeting of MTHFD2. Am J Cancer Res. 2019;9:250–69.
Nishimura T, Nakata A, Chen X, Nishi K, Meguro-Horike M, Sasaki S, et al. Cancer stem-like properties and gefitinib resistance are dependent on purine synthetic metabolism mediated by the mitochondrial enzyme MTHFD2. Oncogene. 2019;38:2464–81.
Lin H, Huang B, Wang H, Liu X, Hong Y, Qiu S, et al. MTHFD2 overexpression predicts poor prognosis in renal cell carcinoma and is associated with cell proliferation and vimentin-modulated migration and invasion. Cell Physiol Biochem. 2018;51:991–1000.
Asai A, Koseki J, Konno M, Nishimura T, Gotoh N, Satoh T, et al. Drug discovery of anticancer drugs targeting methylenetetrahydrofolate dehydrogenase 2. Heliyon. 2018;4:e01021.
Mejia NR, MacKenzie RE. NAD-dependent methylenetetrahydrofolate dehydrogenase is expressed by immortal cells. J Biol Chem. 1985;260:14616–20.
Mejia NR, MacKenzie RE. NAD-dependent methylenetetrahydrofolate dehydrogenase-methenyltetrahydrofolate cyclohydrolase in transformed cells is a mitochondrial enzyme. Biochem Biophys Res Commun. 1988;155:1–6.
Patel H, Pietro ED, MacKenzie RE. Mammalian fibroblasts lacking mitochondrial NAD+-dependent methylenetetrahydrofolate dehydrogenase-cyclohydrolase are glycine auxotrophs. J Biol Chem. 2003;278:19436–41.
Yu C, Yang L, Cai M, Zhou F, Xiao S, Li Y, et al. Down-regulation of MTHFD2 inhibits NSCLC progression by suppressing cycle-related genes. J Cell Mol Med. 2020;24:1568–77.
Gustafsson R, Jemth A-S, Gustafsson NMS, Färnegårdh K, Loseva O, Wiita E, et al. Crystal structure of the emerging cancer target MTHFD2 in complex with a substrate-based inhibitor. Cancer Res. 2017;77:937–48.
Fu C, Sikandar A, Donner J, Zaburannyi N, Herrmann J, Reck M, et al. The natural product carolacton inhibits folate-dependent C1 metabolism by targeting FolD/MTHFD. Nat Commun. 2017;8:1529.
Kawai J, Ota M, Ohki H, Toki T, Suzuki M, Shimada T, et al. Structure-based design and synthesis of an isozyme-selective MTHFD2 inhibitor with a tricyclic coumarin scaffold. ACS Med Chem Lett. 2019;10:893–8.
Kawai J, Toki T, Ota M, Inoue H, Takata Y, Asahi T, et al. Discovery of a potent, selective, and orally available MTHFD2 inhibitor (DS18561882) with in vivo antitumor activity. J Med Chem. 2019. https://doi.org/10.1021/acs.jmedchem.9b01113.
Rios-Orlandi EM, MacKenzie RE. The activities of the NAD-dependent methylenetetrahydrofolate dehydrogenase-methenyltetrahydrofolate cyclohydrolase from ascites tumor cells are kinetically independent. J Biol Chem. 1988;263:4662–7.
Shin M, Momb J, Appling DR. Human mitochondrial MTHFD2 is a dual redox cofactor-specific methylenetetrahydrofolate dehydrogenase/methenyltetrahydrofolate cyclohydrolase. Cancer Metab. 2017;5:11.
Gustafsson Sheppard N, Jarl L, Mahadessian D, Strittmatter L, Schmidt A, Madhusudan N, et al. The folate-coupled enzyme MTHFD2 is a nuclear protein and promotes cell proliferation. Sci Rep. 2015;5:15029.
Koufaris C, Nilsson R. Protein interaction and functional data indicate MTHFD2 involvement in RNA processing and translation. Cancer Metab. 2018;6:12.
Kg P, Re M. NAD+-dependent methylenetetrahydrofolate dehydrogenase-cyclohydrolase: detection of the mRNA in normal murine tissues and transcriptional regulation of the gene in cell lines. Biochim Biophys Acta. 1993;1171:281–7.
Bolusani S, Young BA, Cole NA, Tibbetts AS, Momb J, Bryant JD, et al. Mammalian MTHFD2L encodes a mitochondrial methylenetetrahydrofolate dehydrogenase isozyme expressed in adult tissues. J Biol Chem. 2011;286:5166–74.
Nilsson R, Nicolaidou V, Koufaris C. Mitochondrial MTHFD isozymes display distinct expression, regulation, and association with cancer. Gene. 2019;716:144032.
Prasannan P, Pike S, Peng K, Shane B, Appling DR. Human mitochondrial C1-tetrahydrofolate synthase: gene structure, tissue distribution of the mRNA, and immunolocalization in Chinese hamster ovary calls. J Biol Chem. 2003;278:43178–87.
Momb J, Lewandowski JP, Bryant JD, Fitch R, Surman DR, Vokes SA, et al. Deletion of Mthfd1l causes embryonic lethality and neural tube and craniofacial defects in mice. PNAS. 2013;110:549–54.
Krupenko NI, Dubard ME, Strickland KC, Moxley KM, Oleinik NV, Krupenko SA. ALDH1L2 is the mitochondrial homolog of 10-formyltetrahydrofolate dehydrogenase. J Biol Chem. 2010;285:23056–63.
Sarret C, Ashkavand Z, Paules E, Dorboz I, Pediaditakis P, Sumner S, et al. Deleterious mutations in ALDH1L2 suggest a novel cause for neuro-ichthyotic syndrome. NPJ Genom Med. 2019;4:17.
Schober AF, Mathis AD, Ingle C, Park JO, Chen L, Rabinowitz JD, et al. A two-enzyme adaptive unit within bacterial folate metabolism. Cell Rep. 2019;27:3359–70.e7.
Anderson DD, Quintero CM, Stover PJ. Identification of a de novo thymidylate biosynthesis pathway in mammalian mitochondria. PNAS. 2011;108:15163–8.
Organista-Nava J, Gómez-Gómez Y, Illades-Aguiar B, Rivera-Ramírez AB, Saavedra-Herrera MV, Leyva-Vázquez MA. Overexpression of dihydrofolate reductase is a factor of poor survival in acute lymphoblastic leukemia. Oncol Lett. 2018;15:8405–11.
Raimondi MV, Randazzo O, La Franca M, Barone G, Vignoni E, Rossi D, et al. DHFR inhibitors: reading the past for discovering novel anticancer agents. Molecules. 2019;24:1140.
Raimondi MV, Randazzo O, La Franca M, Barone G, Vignoni E, Rossi D. et al. DHFR Inhibitors: Reading the Past for Discovering Novel Anticancer Agents. Molecules. 2019;24:1140
Schweitzer BI, Dicker AP, Bertino JR. Dihydrofolate reductase as a therapeutic target. The FASEB J. 1990;4:2441–52.
Patel SN, Kain KC. Atovaquone/proguanil for the prophylaxis and treatment of malaria. Expert Rev Anti Infect Ther. 2005;3:849–61.
Wróbel A, Arciszewska K, Maliszewski D, Drozdowska D. Trimethoprim and other nonclassical antifolates an excellent template for searching modifications of dihydrofolate reductase enzyme inhibitors. J Antibiotics. 2020;73:5–27.
Srinivasan B, Skolnick J. Insights into the slow-onset tight-binding inhibition of Escherichia coli dihydrofolate reductase: detailed mechanistic characterization of pyrrolo [3,2-f] quinazoline-1,3-diamine and its derivatives as novel tight-binding inhibitors. FEBS J. 2015;282:1922–38.
Rajagopalan PTR, Zhang Z, McCourt L, Dwyer M, Benkovic SJ, Hammes GG. Interaction of dihydrofolate reductase with methotrexate: ensemble and single-molecule kinetics. Proc Natl Acad Sci USA. 2002;99:13481–6.
Kremer JM. Toward a better understanding of methotrexate. Arthritis Rheum. 2004;50:1370–82.
Yuthavong Y, Tarnchompoo B, Vilaivan T, Chitnumsub P, Kamchonwongpaisan S, Charman SA, et al. Malarial dihydrofolate reductase as a paradigm for drug development against a resistance-compromised target. PNAS. 2012;109:16823–8.
Agarwal PK, Billeter SR, Rajagopalan PTR, Benkovic SJ, Hammes-Schiffer S. Network of coupled promoting motions in enzyme catalysis. PNAS. 2002;99:2794–9.
Cario H, Smith DEC, Blom H, Blau N, Bode H, Holzmann K, et al. Dihydrofolate reductase deficiency due to a homozygous DHFR mutation causes megaloblastic anemia and cerebral folate deficiency leading to severe neurologic disease. Am J Hum Genet. 2011;88:226–31.
Kotoula V, Krikelis D, Karavasilis V, Koletsa T, Eleftheraki AG, Televantou D, et al. Expression of DNA repair and replication genes in non-small cell lung cancer (NSCLC): a role for thymidylate synthetase (TYMS). BMC Cancer. 2012;12:342.
Burdelski C, Strauss C, Tsourlakis MC, Kluth M, Hube-Magg C, Melling N, et al. Overexpression of thymidylate synthase (TYMS) is associated with aggressive tumor features and early PSA recurrence in prostate cancer. Oncotarget. 2015;6:8377–87.
Calvert AH, Alison DL, Harland SJ, Robinson BA, Jackman AL, Jones TR, et al. A phase I evaluation of the quinazoline antifolate thymidylate synthase inhibitor, N10-propargyl-5,8-dideazafolic acid, CB3717. J Clin Oncol. 1986;4:1245–52.
Webber S, Bartlett CA, Boritzki TJ, Hillard JA, Howland EF, Johnston AL, et al. AG337, a novel lipophilic thymidylate synthase inhibitor: in vitro and in vivo preclinical studies. Cancer Chemother Pharm. 1996;37:509–17.
Duch DS, Banks S, Dev IK, Dickerson SH, Ferone R, Heath LS, et al. Biochemical and cellular pharmacology of 1843U89, a novel benzoquinazoline inhibitor of thymidylate synthase. Cancer Res. 1993;53:810–8.
Chu E, Callender MA, Farrell MP, Schmitz JC. Thymidylate synthase inhibitors as anticancer agents: from bench to bedside. Cancer Chemother Pharm. 2003;52:80–9.
Peters GJ, Backus HHJ, Freemantle S, van Triest B, Codacci-Pisanelli G, van der Wilt CL, et al. Induction of thymidylate synthase as a 5-fluorouracil resistance mechanism. Biochim Biophys Acta. 2002;1587:194–205.
Chen D, Jansson A, Sim D, Larsson A, Nordlund P. Structural analyses of human thymidylate synthase reveal a site that may control conformational switching between active and inactive states. J Biol Chem. 2017;292:13449–58.
Lovelace LL, Johnson SR, Gibson LM, Bell BJ, Berger SH, Lebioda L. Variants of human thymidylate synthase with loop 181-197 stabilized in the inactive conformation. Protein Sci. 2009;18:1628–36.
Huang X, Gibson LM, Bell BJ, Lovelace LL, Peña MMO, Berger FG, et al. Replacement of Val3 in human thymidylate synthase affects its kinetic properties and intracellular stability. Biochemistry. 2010;49:2475–82.
Peters EJ, Kraja AT, Lin SJ, Yen-Revollo JL, Marsh S, Province MA, et al. Association of thymidylate synthase variants with 5-fluorouracil cytotoxicity. Pharmacogenet Genom. 2009;19:399–401.
Brodsky G, Barnes T, Bleskan J, Becker L, Cox M, Patterson D. The human GARS-AIRS-GART gene encodes two proteins which are differentially expressed during human brain development and temporally overexpressed in cerebellum of individuals with Down syndrome. Hum Mol Genet. 1997;6:2043–50.
Deis SM, Doshi A, Hou Z, Matherly LH, Gangjee A, Dann CE. Structural and enzymatic analysis of tumor-targeted antifolates that inhibit glycinamide ribonucleotide formyltransferase. Biochemistry. 2016;55:4574–82.
Bronder JL, Moran RG. Antifolates targeting purine synthesis allow entry of tumor cells into S phase regardless of p53 function. Cancer Res. 2002;62:5236–41.
Robert F, Garrett C, Dinwoodie WR, Sullivan DM, Bishop M, Amantea M, et al. Results of 2 phase I studies of intravenous (iv) pelitrexol (AG2037), a glycinamide ribonucleotide formyltransferase (GARFT) inhibitor, in patients (pts) with solid tumors. J Clin Onocol. 2004;22:3075.
Shih C, Chen VJ, Gossett LS, Gates SB, MacKellar WC, Habeck LL. et al. LY231514, a pyrrolo[2,3-d]pyrimidine-based antifolate that inhibits multiple folate-requiring enzymes. Cancer Res. 1997;57:1116–23.
Welin M, Grossmann JG, Flodin S, Nyman T, Stenmark P, Trésaugues L, et al. Structural studies of tri-functional human GART. Nucleic Acids Res. 2010;38:7308–19.
Burda P, Kuster A, Hjalmarson O, Suormala T, Bürer C, Lutz S, et al. Characterization and review of MTHFD1 deficiency: four new patients, cellular delineation and response to folic and folinic acid treatment. J Inherit Metab Dis. 2015;38:863–72.
Sdelci S, Rendeiro AF, Rathert P, You W, Lin J-MG, Ringler A, et al. MTHFD1 interaction with BRD4 links folate metabolism to transcriptional regulation. Nat Genet. 2019;51:990–8.
New Antifolates in Clinical Development. Cancer Network. https://www.cancernetwork.com/view/new-antifolates-clinical-development.
Pikman Y, Puissant A, Alexe G, Furman A, Chen LM, Frumm SM, et al. Targeting MTHFD2 in acute myeloid leukemia. J Exp Med. 2016;213:1285–306.
Zeller KI, Jegga AG, Aronow BJ, O’Donnell KA, Dang CV. An integrated database of genes responsive to the Myc oncogenic transcription factor: identification of direct genomic targets. Genome Biol. 2003;4:R69.
Vazquez A, Markert EK, Oltvai ZN. Serine biosynthesis with one carbon catabolism and the glycine cleavage system represents a novel pathway for ATP generation. PLoS ONE. 2011;6:e25881.
Dang CV, Reddy EP, Shokat KM, Soucek L. Drugging the ‘undruggable’ cancer targets. Nat Rev Cancer. 2017;17:502–8.
Sneader W. Drug discovery: a history. UK: John Wiley & Sons; 2005. https://books.google.se/books?id=jglFsz5EJR8C&printsec=frontcover&source=gbs_ge_summary_r&cad=0#v=onepage&q&f=false.
Wang W, Zhou H, Liu L. Side effects of methotrexate therapy for rheumatoid arthritis: a systematic review. Eur J Med Chem. 2018;158:502–16.
Zachariae H. Methotrexate side-effects. Br J Dermatol. 1990;122:127–33.
Albrecht K, Müller-Ladner U. Side effects and management of side effects of methotrexate in rheumatoid arthritis. Clin Exp Rheumatol. 2010;28:S95–101.
Chan ESL, Cronstein BN. Methotrexate—how does it really work? Nat Rev Rheumatol. 2010;6:175–8.
Bertino JR, Göker E, Gorlick R, Li WW, Banerjee D. Resistance mechanisms to methotrexate in tumors. Stem Cells. 1996;14:5–9.
Nagura E. [Methotrexate and its drug resistance]. Gan Kagaku Ryoho. 1988;15:2882–7.
Wang J, Li G. Mechanisms of methotrexate resistance in osteosarcoma cell lines and strategies for overcoming this resistance. Oncol Lett. 2015;9:940–4.
Wojtuszkiewicz A, Peters GJ, van Woerden NL, Dubbelman B, Escherich G, Schmiegelow K, et al. Methotrexate resistance in relation to treatment outcome in childhood acute lymphoblastic leukemia. J Hematol Oncol. 2015;8:61.
Frantz M-C, Wipf P. Mitochondria as a target in treatment. Environ Mol Mutagen. 2010;51:462–75.
Medwig-Kinney TN, Smith JJ, Palmisano NJ, Tank S, Zhang W, Matus DQ. A developmental gene regulatory network for C. elegans anchor cell invasion. Development. 2020;147. https://doi.org/10.1242/dev.185850.
Ducker GS, Chen L, Morscher RJ, Ghergurovich JM, Esposito M, Teng X, et al. Reversal of cytosolic one-carbon flux compensates for loss of the mitochondrial folate pathway. Cell Metab. 2016;23:1140–53.
Toyama BH, Hetzer MW. Protein homeostasis: live long, won’t prosper. Nat Rev Mol Cell Biol. 2013;14:55–61.
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P. From RNA to protein. In: Molecular biology of the cell. 4th ed. New York: Garland Science; 2002. https://www.ncbi.nlm.nih.gov/books/NBK26829/.
Acknowledgements
LNZ was supported by an A*STAR International Fellowship (AIF) from Singapore. This work is supported by the IngaBritt och Arne Lundbergs Forskningsstiftelse LU2020-0013; PK is supported by the Faculty of Medicine, Lund University; the Swedish Foundation for Strategic Research Dnr IRC15-0067, and Swedish Research Council, Strategic Research Area EXODIAB, Dnr 2009–1039. LNZ would like to thank Prof. Chew Lock Yue (NTU) and Prof. Ulf Ryde (LU) for comments on the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhao, L.N., Björklund, M., Caldez, M.J. et al. Therapeutic targeting of the mitochondrial one-carbon pathway: perspectives, pitfalls, and potential. Oncogene 40, 2339–2354 (2021). https://doi.org/10.1038/s41388-021-01695-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01695-8
This article is cited by
-
MTHFD2 in healthy and cancer cells: Canonical and non-canonical functions
npj Metabolic Health and Disease (2024)
-
SHMT2 promotes papillary thyroid cancer metastasis through epigenetic activation of AKT signaling
Cell Death & Disease (2024)
-
Targeting serine-glycine-one-carbon metabolism as a vulnerability in cancers
Biomarker Research (2023)
-
Identification of the molecular subtypes and construction of risk models in neuroblastoma
Scientific Reports (2023)
-
Proteogenomic analysis of acute myeloid leukemia associates relapsed disease with reprogrammed energy metabolism both in adults and children
Leukemia (2023)