Abstract
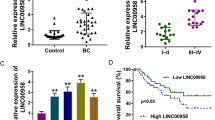
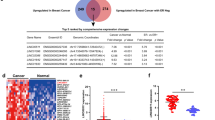
Increasing evidence supports that long noncoding RNAs (lncRNAs) act as master regulators involved in tumorigenesis and development at the N6-methyladenine (m6A) epigenetic modification level. However, the underlying regulatory mechanism in breast cancer (BRCA) remains elusive. Here, we unveil that LINC00942 (LNC942) exerts its functions as an oncogene in promoting METTL14-mediated m6A methylation and regulating the expression and stability of its target genes CXCR4 and CYP1B1 in BRCA initiation and progression. Specifically, LNC942 and METTL14 were significantly upregulated accompanied with the upregulation of m6A levels in BRCA cells and our included BRCA cohorts (n = 150). Functionally, LNC942 elicits potent oncogenic effects on promoting cell proliferation and colony formation and inhibiting cell apoptosis, subsequently elevating METTL14-mediated m6A methylation levels and its associated mRNA stability and protein expression of CXCR4 and CYP1B1 in BRCA cells. Mechanistically, LNC942 directly recruits METTL14 protein by harboring the specific recognize sequence (+176–+265), thereby stabilized the expression of downstream targets of LNC942 including CXCR4 and CYP1B1 through posttranscriptional m6A methylation modification in vitro and in vivo. Therefore, our results uncover a novel LNC942-METTL14-CXCR4/CYP1B1 signaling axis, which provides new targets and crosstalk m6A epigenetic modification mechanism for BRCA prevention and treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The authors declare that all the data supporting the findings in this study are available in this study and its Supplementary materials, or are available from the corresponding author through reasonable request.
Change history
15 February 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41388-022-02194-0
References
Adhikari S, Xiao W, Zhao YL, Yang YG. m(6)A: Signaling for mRNA splicing. RNA Biol. 2016;13:756–9.
Deng X, Su R, Weng H, Huang H, Li Z, Chen J. RNA N(6)-methyladenosine modification in cancers: current status and perspectives. Cell Res. 2018;28:507–17.
Huang H, Weng H, Chen J. m(6)A Modification in coding and non-coding RNAs: roles and therapeutic implications in cancer. Cancer Cell. 2020;37:270–88.
Pan Y, Ma P, Liu Y, Li W, Shu Y. Multiple functions of m(6)A RNA methylation in cancer. J Hematol Oncol. 2018;11:48.
Niu Y, Zhao X, Wu YS, Li MM, Wang XJ, Yang YG. N6-methyl-adenosine (m6A) in RNA: an old modification with a novel epigenetic function. Genom Proteom Bioinforma. 2013;11:8–17.
Sun T, Wu R, Ming L. The role of m6A RNA methylation in cancer. Biomed Pharmacother. 2019;112:108613.
Jin D, Guo J, Wu Y, Du J, Yang L, Wang X, et al. m(6)A mRNA methylation initiated by METTL3 directly promotes YAP translation and increases YAP activity by regulating the MALAT1-miR-1914-3p-YAP axis to induce NSCLC drug resistance and metastasis. J Hematol Oncol. 2019;12:135.
Liu N, Dai Q, Zheng G, He C, Parisien M, Pan T. N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature. 2015;518:560–4.
Zhang S, Zhao BS, Zhou A, Lin K, Zheng S, Lu Z, et al. m(6)A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1 expression and cell proliferation program. Cancer Cell. 2017;31:591–606.e6.
Wang X, Zhang J, Wang Y. Long noncoding RNA GAS5-AS1 suppresses growth and metastasis of cervical cancer by increasing GAS5 stability. Am J Transl Res. 2019;11:4909–21.
Zhang C, Zhang M, Ge S, Huang W, Lin X, Gao J, et al. Reduced m6A modification predicts malignant phenotypes and augmented Wnt/PI3K-Akt signaling in gastric cancer. Cancer Med. 2019;8:4766–81.
Li Z, Weng H, Su R, Weng X, Zuo Z, Li C, et al. FTO plays an oncogenic role in acute myeloid leukemia as a N(6)-methyladenosine RNA demethylase. Cancer Cell. 2017;31:127–41.
Li F, Yi Y, Miao Y, Long W, Long T, Chen S, et al. N(6)-methyladenosine modulates nonsense-mediated mRNA decay in human glioblastoma. Cancer Res. 2019;79:5785–98.
Yao QJ, Sang L, Lin M, Yin X, Dong W, Gong Y, et al. Mettl3-Mettl14 methyltransferase complex regulates the quiescence of adult hematopoietic stem cells. Cell Res. 2018;28:952–4.
Yue B, Song C, Yang L, Cui R, Cheng X, Zhang Z, et al. METTL3-mediated N6-methyladenosine modification is critical for epithelial-mesenchymal transition and metastasis of gastric cancer. Mol Cancer 2019;18:142.
Weng H, Huang H, Wu H, Qin X, Zhao BS, Dong L, et al. METTL14 inhibits hematopoietic stem/progenitor differentiation and promotes leukemogenesis via mRNA m(6)A modification. Cell Stem Cell. 2018;22:191–205.e9.
Panneerdoss SA-O, Eedunuri VK, Yadav P, Timilsina S, Rajamanickam S, Viswanadhapalli S, et al. Cross-talk among writers, readers, and erasers of m(6)A regulates cancer growth and progression. Sci Adv. 2018;4:eaar8263.
Lang F, Singh RK, Pei Y, Zhang S, Sun K, Robertson ES. EBV epitranscriptome reprogramming by METTL14 is critical for viral-associated tumorigenesis. PLoS Pathog. 2019;15:e1007796.
Kuppers DA, Arora S, Lim Y, Lim AR, Carter LM, Corrin PD, et al. N(6)-methyladenosine mRNA marking promotes selective translation of regulons required for human erythropoiesis. Nat Commun. 2019;10:4596.
Ma JZ, Yang F, Zhou CC, Liu F, Yuan JH, Wang F, et al. METTL14 suppresses the metastatic potential of hepatocellular carcinoma by modulating N(6)-methyladenosine-dependent primary MicroRNA processing. Hepatology. 2017;65:529–43.
Gu C, Wang Z, Zhou N, Li G, Kou Y, Luo Y, et al. Mettl14 inhibits bladder TIC self-renewal and bladder tumorigenesis through N(6)-methyladenosine of Notch1. Mol Cancer. 2019;18:168.
Cui Q, Shi H, Ye P, Li L, Qu Q, Sun G, et al. m(6)A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep. 2017;18:2622–34.
Gong D, Zhang J, Chen Y, Xu Y, Ma J, Hu G, et al. The m(6)A-suppressed P2RX6 activation promotes renal cancer cells migration and invasion through ATP-induced Ca(2+) influx modulating ERK1/2 phosphorylation and MMP9 signaling pathway. J Exp Clin Cancer Res. 2019;38:233.
Yang X, Zhang S, He C, Xue P, Zhang L, He Z, et al. METTL14 suppresses proliferation and metastasis of colorectal cancer by down-regulating oncogenic long non-coding RNA XIST. Mol Cancer. 2020;19:46.
Liu J, Eckert MA-O, BA-OX Harada, Liu SM, Lu Z, Yu K, et al. m(6)A mRNA methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer. Nat Cell Biol. 2018;20:1074–83.
Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014;42(Database issue):D92–7.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Kashyap D, Kaur H. Cell-free miRNAs as non-invasive biomarkers in breast cancer: Significance in early diagnosis and metastasis prediction. Life Sci. 2020;246:117417.
Anastasiadou E, Jacob LS, Slack FJ. Non-coding RNA networks in cancer. Nat Rev Cancer. 2018;18:5–18.
Matsui M, Corey DR. Non-coding RNAs as drug targets. Nat Rev Drug Discov. 2017;16:167–79.
Zhi H, Li Y, Wang L. Profiling DNA methylation patterns of non-coding RNAs (ncRNAs) in human disease. Adv Exp Med Biol. 2018;1094:49–64.
Zhu KP, Zhang CL, Ma XL, Hu JP, Cai T, Zhang L. Analyzing the interactions of mRNAs and ncRNAs to predict competing endogenous RNA networks in osteosarcoma chemo-resistance. Mol Ther. 2019;27:518–30.
Mego M, Cholujova D, Minarik G, Sedlackova T, Gronesova P, Karaba M, et al. CXCR4-SDF-1 interaction potentially mediates trafficking of circulating tumor cells in primary breast cancer. BMC Cancer. 2016;16:127.
Yang F, Takagaki Y, Yoshitomi Y, Ikeda T, Li J, Kitada M, et al. Inhibition of dipeptidyl peptidase-4 accelerates epithelial-mesenchymal transition and breast cancer metastasis via the CXCL12/CXCR4/mTOR Axis. Cancer Res. 2019;79:735–46.
Golmohammadzadeh G, Mohammadpour A, Ahangar N, Shokrzadeh M. Polymorphisms in Phase I (CYP450) Genes CYP1A1 (rs4646421), CYP1B1 (rs1056836), CYP19A1 (rs749292) and CYP2C8 (rs1058930) and their relation to risk of breast cancer: a case-control study in Mazandaran Province in North of Iran. Open Access Maced J Med Sci. 2019;7:2488–96.
Yu AM, Batra N, Tu MJ, Sweeney C. Novel approaches for efficient in vivo fermentation production of noncoding RNAs. Appl Microbiol Biotechnol. 2020;104:1927–37.
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 2012;485:201–6.
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell. 2012;149:1635–46.
Liu N, Parisien M, Dai Q, Zheng G, He C, Pan T, et al. Modification status at single nucleotide resolution in mRNA and long noncoding RNA. RNA. 2013;19:1848–56.
Yang D, Qiao J, Wang G, Lan Y, Li G, Guo X, et al. N6-Methyladenosine modification of lincRNA 1281 is critically required for mESC differentiation potential. Nucleic Acids Res. 2018;46:3906–20.
Ashouri A, Sayin VI, Van den Eynden J, Singh SX, Papagiannakopoulos T, Larsson E. Pan-cancer transcriptomic analysis associates long non-coding RNAs with key mutational driver events. Nat Commun. 2016;7:13197.
Tao Y, Li Y, Huang J, Liang B. Screening LncRNAs related to the prognosis of patients with lung adenocarcinoma by bioinformatics. Int J Lab Med (in Chinese). 2019;40:1541–4.
Patil DP, Chen CK, Pickering BF, Chow A, Jackson C, Guttman M, et al. m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature. 2016;537:369–73.
Ni W, Yao S, Zhou Y, Liu Y, Huang P, Zhou A, et al. Long noncoding RNA GAS5 inhibits progression of colorectal cancer by interacting with and triggering YAP phosphorylation and degradation and is negatively regulated by the m(6)A reader YTHDF3. Mol Cancer. 2019;18:143.
Du H, Zhao Y, He J, Zhang Y, Xi H, Liu M, et al. YTHDF2 destabilizes m(6)A-containing RNA through direct recruitment of the CCR4-NOT deadenylase complex. Nat Commun. 2016;7:12626.
Chen SA-O, Zhou L, Wang Y. ALKBH5-mediated m(6)A demethylation of lncRNA PVT1 plays an oncogenic role in osteosarcoma. Cancer Cell Int. 2020;20:34.
Yan J, Huang X, Zhang X, Chen Z, Ye C, Xiang W, et al. LncRNA LINC00470 promotes the degradation of PTEN mRNA to facilitate malignant behavior in gastric cancer cells. Biochem Biophys Res Commun. 2020;521:887–93.
Acknowledgements
We gratefully appreciate the efforts and contributions of doctors, nurses, and technical staff at the First Hospital of China Medical University, Cancer Hospital of China Medical University.
Funding
This work was supported by grants from the National Natural Science Foundation of China [No. 31828005, 81872905, and 81673475], National Natural Science Foundation of China and Liaoning joint fund key program [No. U1608281], Liaoning Revitalization Talents Program [XLYC1807155], China Postdoctoral Science Foundation (Grant No. 2019M661180), Special Foundation of China postdoctoral science foundation (Grant No. 2019T120225), and Shenyang S&T Projects (19-109-4-09).
Author information
Authors and Affiliations
Contributions
HW, MW, and TS conceived and designed the project. TS, ZW, XW, XD, HG, and WY performed experiments and/or data acquisition and analyses; YW, XH, WQ, SL, DX, YW, QC, YL, YW, and BF contributed technical/reagents materials, analytic tools and/or grant support; HW, MW, and TS prepared, wrote, and/or revision the paper. All authors discussed the results and commented on the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This study was approved by the Ethics Committee of China Medical University. All the animal experiments performed in this study were approved by the Institutional Animal Care and Use Committee of China Medical University.
Informed consent
All of the authors have written informed consent.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Sun, T., Wu, Z., Wang, X. et al. LNC942 promoting METTL14-mediated m6A methylation in breast cancer cell proliferation and progression. Oncogene 39, 5358–5372 (2020). https://doi.org/10.1038/s41388-020-1338-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-1338-9
This article is cited by
-
Emerging role of RNA modification and long noncoding RNA interaction in cancer
Cancer Gene Therapy (2024)
-
Identification of m6A/m5C-related lncRNA signature for prediction of prognosis and immunotherapy efficacy in esophageal squamous cell carcinoma
Scientific Reports (2024)
-
LINC00942 Alleviates NaAsO2-induced Apoptosis by Promoting GSH Synthesis Through Targeting miR-214-5p
Biological Trace Element Research (2024)
-
LINC00942 inhibits ferroptosis and induces the immunosuppression of regulatory T cells by recruiting IGF2BP3/SLC7A11 in hepatocellular carcinoma
Functional & Integrative Genomics (2024)
-
Emerging roles of m6A RNA modification in cancer therapeutic resistance
Experimental Hematology & Oncology (2023)