Abstract
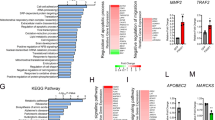
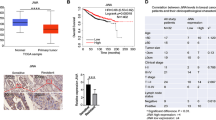
A principal challenge in treating acute myeloid leukemia (AML) is chemotherapy refractory disease. As such, there remains a critical need to identify key regulators of chemotherapy resistance in AML. In this study, we demonstrate that the membrane scaffold, CD82, contributes to the chemoresistant phenotype of AML. Using an RNA-seq approach, we identified the increased expression of the tetraspanin family member, CD82, in response to the chemotherapeutic, daunorubicin. Analysis of the TARGET and BEAT AML databases identifies a correlation between CD82 expression and overall survival of AML patients. Moreover, using a combination of cell lines and patient samples, we find that CD82 overexpression results in significantly reduced cell death in response to chemotherapy. Investigation of the mechanism by which CD82 promotes AML survival in response to chemotherapy identified a crucial role for enhanced protein kinase c alpha (PKCα) signaling and downstream activation of the β1 integrin. In addition, analysis of β1 integrin clustering by super-resolution imaging demonstrates that CD82 expression promotes the formation of dense β1 integrin membrane clusters. Lastly, evaluation of survival signaling following daunorubicin treatment identified robust activation of p38 mitogen-activated protein kinase (MAPK) downstream of PKCα and β1 integrin signaling when CD82 is overexpressed. Together, these data propose a mechanism where CD82 promotes chemoresistance by increasing PKCα activation and downstream activation/clustering of β1 integrin, leading to AML cell survival via activation of p38 MAPK. These observations suggest that the CD82-PKCα signaling axis may be a potential therapeutic target for attenuating chemoresistance signaling in AML.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
American Cancer Society. Cancer facts & figures. Atlanta, GA: American Cancer Society.
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127:2391–405.
Nasir SS, Giri S, Nunnery S, Martin MG. Outcome of adolescents and young adults compared with pediatric patients with acute myeloid and promyelocytic leukemia. Clin Lymphoma Myeloma Leuk. 2017;17:126–32. e1.
Veuger M, Honders W, Spoelder N, Willemze R, Barge R. Effects of the use of multiple cytotoxic drugs on resistance mechanisms in acute myeloid leukemia. Leukemia. 2001;15:503.
Zhang J, Gu Y, Chen BA. Mechanisms of drug resistance in acute myeloid leukemia. Oncotargets Ther. 2019;12:1937–45.
Cloos J, Goemans BF, Hess CJ, van Oostveen JW, Waisfisz Q, Corthals S, et al. Stability and prognostic influence of FLT3 mutations in paired initial and relapsed AML samples. Leukemia. 2006;20:1217–20.
El Fakih R, Rasheed W, Hawsawi Y, Alsermani M, Hassanein M. Targeting FLT3 mutations in acute myeloid leukemia. Cells. 2018;7:E4.
Cook AM, Li L, Ho YW, Lin A, Li L, Stein A, et al. Role of altered growth factor receptor-mediated JAK2 signaling in growth and maintenance of human acute myeloid leukemia stem cells. Blood. 2014;123:2826–37.
McCubrey JA, Steelman LS, Abrams SL, Bertrand FE, Ludwig DE, Basecke J, et al. Targeting survival cascades induced by activation of Ras/Raf/MEK/ERK, PI3K/PTEN/Akt/mTOR and Jak/STAT pathways for effective leukemia therapy. Leukemia. 2008;22:708–22.
Bosman MC, Schuringa JJ, Vellenga E. Constitutive NF-kappaB activation in AML: causes and treatment strategies. Crit Rev Oncol Hematol. 2016;98:35–44.
Davoudi Z, Akbarzadeh A, Rahmatiyamchi M, Movassaghpour AA, Alipour M, Nejati-Koshki K, et al. Molecular target therapy of AKT and NF-kB signaling pathways and multidrug resistance by specific cell penetrating inhibitor peptides in HL-60 cells. Asian Pac J Cancer Prev. 2014;15:4353–8.
Hemler ME. Tetraspanin functions and associated microdomains. Nat Rev Mol Cell Bio. 2005;6:801–11.
Hemler ME. Tetraspanin proteins promote multiple cancer stages. Nat Rev Cancer. 2014;14:49–60.
Kwon HY, Bajaj J, Ito T, Blevins A, Konuma T, Weeks J, et al. Tetraspanin 3 is required for the development and propagation of acute myelogenous leukemia. Cell Stem Cell. 2015;17:152–64.
Boyer T, Guihard S, Roumier C, Peyrouze P, Gonzales F, Berthon C, et al. Tetraspanin CD81 is an adverse prognostic marker in acute myeloid leukemia. Oncotarget. 2016;7:62377–85.
Burchert A, Notter M, Menssen HD, Schwartz S, Knauf W, Neubauer A, et al. CD82 (KAI1), a member of the tetraspan family, is expressed on early haemopoietic progenitor cells and up-regulated in distinct human leukaemias. Brit J Haematol. 1999;107:494–504.
Saito-Reis CA, Marjon KD, Pascetti EM, Floren M, Gillette JM. The tetraspanin CD82 regulates bone marrow homing and engraftment of hematopoietic stem and progenitor cells. Mol Biol Cell. 2018;29:2946–58.
Nishioka C, Ikezoe T, Yang J, Yokoyama A. Tetraspanin family member, CD82, regulates expression of EZH2 via inactivation of p38 MAPK signaling in leukemia cells. PLoS ONE. 2015;10:e0125017.
Marjon KD, Termini CM, Karlen KL, Saito-Reis C, Soria CE, Lidke KA, et al. Tetraspanin CD82 regulates bone marrow homing of acute myeloid leukemia by modulating the molecular organization of N-cadherin. Oncogene. 2016;35:4132–40.
Termini CM, Lidke KA, Gillette JM. Tetraspanin CD82 regulates the spatiotemporal dynamics of PKC alpha in acute myeloid leukemia. Sci Rep. 2016;29895.
Kurinna S, Konopleva M, Palla SL, Chen W, Kornblau S, Contractor R, et al. Bcl2 phosphorylation and active PKC alpha are associated with poor survival in AML. Leukemia. 2006;20:1316–9.
Ruvolo PP, Zhou LR, Watt JC, Ruvolo VR, Burks JK, Jiffar T, et al. Targeting pkc-mediated signal transduction pathways using enzastaurin to promote apoptosis in acute myeloid leukemia-derived cell lines and blast cells. J Cell Biochem. 2011;112:1696–707.
Wakui M, Kuriyama K, Miyazaki Y, Hata T, Taniwaki M, Ohtake S, et al. Diagnosis of acute myeloid leukemia according to the WHO classification in the Japan Adult Leukemia Study Group AML-97 protocol. Int J Hematol. 2008;87:144–51.
Walter RB, Othus M, Burnett AK, Lowenberg B, Kantarjian HM, Ossenkoppele GJ, et al. Significance of FAB subclassification of “acute myeloid leukemia, NOS” in the 2008 WHO classification: analysis of 5848 newly diagnosed patients. Blood. 2013;121:2424–31.
Byun JM, Kim YJ, Yoon HJ, Kim SY, Kim HJ, Yoon J, et al. Cytogenetic profiles of 2806 patients with acute myeloid leukemia-a retrospective multicenter nationwide study. Ann Hematol. 2016;95:1223–32.
Okamoto Y, Kudo K, Tabuchi K, Tomizawa D, Taga T, Goto H, et al. Hematopoietic stem-cell transplantation in children with refractory acute myeloid leukemia. Bone Marrow Transpl. 2019;54:1489–98.
Takami M, Katayama K, Noguchi K, Sugimoto Y. Protein kinase C alpha-mediated phosphorylation of PIM-1L promotes the survival and proliferation of acute myeloid leukemia cells. Biochem Biophys Res Commun. 2018;503:1364–71.
Efimova T, Deucher A, Kuroki T, Ohba M, Eckert RL. Novel protein kinase C isoforms regulate human keratinocyte differentiation by activating a p38 delta mitogen-activated protein kinase cascade that targets CCAAT/enhancer-binding protein alpha. J Biol Chem. 2002;277:31753–60.
Hocevar BA, Morrow DM, Tykocinski ML, Fields AP. Protein kinase C isotypes in human erythroleukemia cell proliferation and differentiation. J Cell Sci. 1992;101:671–9.
Myklebust JH, Smeland EB, Josefsen D, Sioud M. Protein kinase C-alpha isoform is involved in erythropoietin-induced erythroid differentiation of CD34(+) progenitor cells from human bone marrow. Blood. 2000;95:510–8.
Jin JK, Tien PC, Cheng CJ, Song JH, Huang C, Lin SH, et al. Talin1 phosphorylation activates beta1 integrins: a novel mechanism to promote prostate cancer bone metastasis. Oncogene. 2015;34:1811–21.
Ramirez P, Rettig MP, Uy GL, Deych E, Holt MS, Ritchey JK, et al. BIO5192, a small molecule inhibitor of VLA-4, mobilizes hematopoietic stem and progenitor cells. Blood. 2009;114:1340–3.
van Kooyk Y, Figdor CG. Avidity regulation of integrins: the driving force in leukocyte adhesion. Curr Opin Cell Biol. 2000;12:542–7.
Kolanus W, Seed B. Integrins and inside-out signal transduction: converging signals from PKC and PIP3. Curr Opin Cell Biol. 1997;9:725–31.
Martin Ester H-PK, Jorg Sander, Xiaowei Xu. A density-based algorithm for discovering clusters in large spatial databases with noise. In Proceedings of the 2nd International Conference on Knowledge Discovery and Data Mining (KDD-96). Portland, Oregon: AAAI Press; 1996. p. 226–31.
Thornton TM, Rincon M. Non-classical p38 map kinase functions: cell cycle checkpoints and survival. Int J Biol Sci. 2009;5:44–51.
Wood CD, Thornton TM, Sabio G, Davis RA, Rincon M. Nuclear localization of p38 MAPK in response to DNA damage. Int J Biol Sci. 2009;5:428–37.
Gupta J, Igea A, Papaioannou M, Lopez-Casas PP, Llonch E, Hidalgo M, et al. Pharmacological inhibition of p38 MAPK reduces tumor growth in patient-derived xenografts from colon tumors. Oncotarget. 2015;6:8539–51.
Naci D, Aoudjit F. Alpha2beta1 integrin promotes T cell survival and migration through the concomitant activation of ERK/Mcl-1 and p38 MAPK pathways. Cell Signal. 2014;26:2008–15.
Scott MJ, Billiar TR. Beta2-integrin-induced p38 MAPK activation is a key mediator in the CD14/TLR4/MD2-dependent uptake of lipopolysaccharide by hepatocytes. J Biol Chem. 2008;283:29433–46.
Hwang S, Takimoto T, Hemler ME. Integrin-independent support of cancer drug resistance by tetraspanin CD151. Cell Mol Life Sci. 2019;76:1595–604.
Ye M, Wei T, Wang Q, Sun Y, Tang R, Guo L, et al. TSPAN12 promotes chemoresistance and proliferation of SCLC under the regulation of miR-495. Biochem Biophys Res Commun. 2017;486:349–56.
Kohmo S, Kijima T, Otani Y, Mori M, Minami T, Takahashi R, et al. Cell surface tetraspanin CD9 mediates chemoresistance in small cell lung cancer. Cancer Res. 2010;70:8025–35.
Ullah M, Akbar A, Ng NN, Concepcion W, Thakor AS. Mesenchymal stem cells confer chemoresistance in breast cancer via a CD9 dependent mechanism. Oncotarget. 2019;10:3435–50.
Nishioka C, Ikezoe T, Yang J, Nobumoto A, Kataoka S, Tsuda M, et al. CD82 regulates STAT5/IL-10 and supports survival of acute myelogenous leukemia cells. Int J Cancer. 2014;134:55–64.
Ji H, Chen L, Xing Y, Li S, Dai J, Zhao P, et al. CD82 supports survival of childhood acute myeloid leukemia cells via activation of Wnt/beta-catenin signaling pathway. Pediatr Res. 2019;85:1024–31.
Nishioka C, Ikezoe T, Takeuchi A, Nobumoto A, Tsuda M, Yokoyama A. The novel function of CD82 and its impact on BCL2L12 via AKT/STAT5 signal pathway in acute myelogenous leukemia cells. Leukemia. 2015;29:2296–306.
Bonardi F, Fusetti F, Deelen P, van Gosliga D, Vellenga E, Schuringa JJ. A proteomics and transcriptomics approach to identify leukemic stem cell (LSC) markers. Mol Cell Proteom. 2013;12:626–37.
Tonoli H, Barrett JC. CD82 metastasis suppressor gene: a potential target for new therapeutics? Trends Mol Med. 2005;11:563–70.
Miranti CK. Controlling cell surface dynamics and signaling: how CD82/KAI1 suppresses metastasis. Cell Signal 2009;21:196–211.
Tsai YC, Weissman AM. Dissecting the diverse functions of the metastasis suppressor CD82/KAI1. FEBS Lett. 2011;585:3166–73.
Nishioka C, Ikezoe T, Furihata M, Yang J, Serada S, Naka T, et al. CD34+/CD38 acute myelogenous leukemia cells aberrantly express CD82 which regulates adhesion and survival of leukemia stem cells. Int J Cancer. 2013;132:2006–19.
Brockstein B, Samuels B, Humerickhouse R, Arietta R, Fishkin P, Wade J. et al. Phase II studies of bryostatin-1 in patients with advanced sarcoma and advanced head and neck cancer. Invest New Drugs. 2001;19:249–54.
Varterasian ML, Mohammad RM, Shurafa MS, Hulburd K, Pemberton PA, Rodriguez DH, et al. Phase II trial of bryostatin 1 in patients with relapsed low-grade non-Hodgkin’s lymphoma and chronic lymphocytic leukemia. Clin Cancer Res. 2000;6:825–8.
Crump M, Leppa S, Fayad L, Lee JJ, Di Rocco A, Ogura M, et al. Randomized, double-blind, phase III trial of enzastaurin versus placebo in patients achieving remission after first-line therapy for high-risk diffuse large b-cell lymphoma. J Clin Oncol. 2016;34:2484–92.
Aoudjit F, Vuori K. Integrin signaling in cancer cell survival and chemoresistance. Chemother Res Pract. 2012;2012:283181.
Naci D, El Azreq MA, Chetoui N, Lauden L, Sigaux F, Charron D, et al. alpha2beta1 integrin promotes chemoresistance against doxorubicin in cancer cells through extracellular signal-regulated kinase (ERK). J Biol Chem. 2012;287:17065–76.
Nair MG, Desai K, Prabhu JS, Hari PS, Remacle J, Sridhar TS. beta3 integrin promotes chemoresistance to epirubicin in MDA-MB-231 through repression of the pro-apoptotic protein, BAD. Exp Cell Res. 2016;346:137–45.
Sansing HA, Sarkeshik A, Yates JR, Patel V, Gutkind JS, Yamada KM, et al. Integrin alphabeta1, alphavbeta, alpha6beta effectors p130Cas, Src and talin regulate carcinoma invasion and chemoresistance. Biochem Biophys Res Commun. 2011;406:171–6.
Seguin L, Desgrosellier JS, Weis SM, Cheresh DA. Integrins and cancer: regulators of cancer stemness, metastasis, and drug resistance. Trends Cell Biol. 2015;25:234–40.
Berrazouane S, Boisvert M, Salti S, Mourad W, Al-Daccak R, Barabe F, et al. Beta1 integrin blockade overcomes doxorubicin resistance in human T-cell acute lymphoblastic leukemia. Cell Death Dis. 2019;10:357.
Zhang XA, Bontrager AL, Hemler ME. Transmembrane-4 superfamily proteins associate with activated protein kinase C (PKC) and link PKC to specific beta(1) integrins. J Biol Chem. 2001;276:25005–13.
Termini CM, Cotter ML, Marjon KD, Buranda T, Lidke K, Gillette JM. Regulation of VLA-4 mediated hematopoietic stem/progenitor cell adhesion by CD82. Mol Biol Cell. 2012;23:1545–1697.
Bachegowda L, Morrone K, Winski SL, Mantzaris I, Bartenstein M, Ramachandra N, et al. Pexmetinib: a novel dual inhibitor of Tie2 and p38 MAPK with efficacy in preclinical models of myelodysplastic syndromes and acute myeloid leukemia. Cancer Res. 2016;76:4841–9.
Garcia-Manero G, Khoury HJ, Jabbour E, Lancet J, Winski SL, Cable L, et al. A phase I study of oral ARRY-614, a p38 MAPK/Tie2 dual inhibitor, in patients with low or intermediate-1 risk myelodysplastic syndromes. Clin Cancer Res. 2015;21:985–94.
Beckwith KA, Byrd JC, Muthusamy N. Tetraspanins as therapeutic targets in hematological malignancy: a concise review. Front Physiol. 2015;6:91.
Brown RB, Madrid NJ, Suzuki H, Ness SA. Optimized approach for Ion Proton RNA sequencing reveals details of RNA splicing and editing features of the transcriptome. PLoS ONE. 2017;12:e0176675.
Brayer KJ, Frerich CA, Kang H, Ness SA. Recurrent fusions in MYB and MYBL1 define a common, transcription factor-driven oncogenic pathway in salivary gland adenoid cystic carcinoma. Cancer Discov. 2016;6:176–87.
Frerich CA, Brayer KJ, Painter BM, Kang H, Mitani Y, El-Naggar AK, et al. Transcriptomes define distinct subgroups of salivary gland adenoid cystic carcinoma with different driver mutations and outcomes. Oncotarget. 2018;9:7341–58.
Anders S, Pyl PT, Huber W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–9.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
Varet H, Brillet-Gueguen L, Coppee JY, Dillies MA. SARTools: a DESeq2- and EdgeR-based R pipeline for comprehensive differential analysis of RNA-Seq data. PLoS ONE. 2016;11:e0157022.
Saldanha AJ. Java treeview-extensible visualization of microarray data. Bioinformatics. 2004;20:3246–8.
Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun. 2019;10:1523.
Tyner JW, Tognon CE, Bottomly D, Wilmot B, Kurtz SE, Druker BJ, et al. Functional genomic landscape of acute myeloid leukaemia. Nature. 2018;562:526–31.
Valley CC, Liu S, Lidke DS, Lidke KA. Sequential superresolution imaging of multiple targets using a single fluorophore. PLoS ONE. 2015;10:e0123941.
Huang F, Schwartz SL, Byars JM, Lidke KA. Simultaneous multiple-emitter fitting for single molecule super-resolution imaging. Biomed Opt Express. 2011;2:1377–93.
Acknowledgements
This work was supported by fellowships: F31CA232480 to MF, F31HL124977 to CMT, and T32HL007736 to KDM, and an NHLBI investigator grant (RO1HL122483 to JMG), and an American Cancer Society Research Scholar Grant (130675 to JMG). This work was partially supported by the UNM Spatiotemporal Modeling Center (P50GM085273) and the UNM Comprehensive Cancer Center Support Grant (P30CA118100) through the following shared resources: Analytical and Translational Genomics, Human Tissue Repository, Fluorescence Microscopy, Flow Cytometry and Biostatistics. We also acknowledge Victoria Balise for critical manuscript review and the experimental assistance of Christian Doyle.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Floren, M., Restrepo Cruz, S., Termini, C.M. et al. Tetraspanin CD82 drives acute myeloid leukemia chemoresistance by modulating protein kinase C alpha and β1 integrin activation. Oncogene 39, 3910–3925 (2020). https://doi.org/10.1038/s41388-020-1261-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-1261-0
This article is cited by
-
The versatile roles of testrapanins in cancer from intracellular signaling to cell–cell communication: cell membrane proteins without ligands
Cell & Bioscience (2023)
-
Single-cell RNA sequencing distinctly characterizes the wide heterogeneity in pediatric mixed phenotype acute leukemia
Genome Medicine (2023)
-
Transcriptionally imprinted glycomic signatures of acute myeloid leukemia
Cell & Bioscience (2023)
-
Identification of Hub Genes Associated with Resistance to Prednisolone in Acute Lymphoblastic Leukemia Based on Weighted Gene Co-expression Network Analysis
Molecular Biotechnology (2023)