Abstract
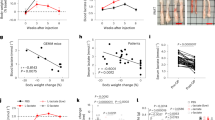
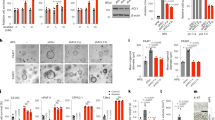
Despite significant progression in the study of hepatocellular carcinoma (HCC), the role of the proteasome in regulating cross talk between mTOR signaling and glycolysis in liver cancer progression is not fully understood. Here, we demonstrate that deficiency of REGγ, a proteasome activator, in mice significantly attenuates DEN-induced liver tumor formation. Ablation of REGγ increases the stability of PP2Ac (protein phosphatase 2 catalytic subunit) in vitro and in vivo, which dephosphorylates PRAS40 (AKT1 substrate 1) and stabilizes the interaction between PRAS40 and Raptor to inactive mTORC1-mediated hyper-glycolytic metabolism. In the DEN-induced animal model and clinical hepato-carcinoma samples, high levels of REGγ in HCC tumor regions contribute to reduced expression of PP2Ac, leading to accumulation of phosphorylated PRAS40 and mTORC1-mediated activation of HIF1α. Interestingly, mTORC1 enhances REGγ activity in HCC, forming a positive feedback regulatory loop. In conclusion, our study identifies REGγ-PP2Ac-PRAS40 axis as a new layer in regulating mTORC1 activity and downstream glycolytic alterations during HCC development, highlighting the REGγ-proteasome as a potential target for personalized HCC therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Laursen L. A preventable cancer. Nature. 2014;516:S2–3.
Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA Cancer J Clin. 2014;64:9–29.
Chiang DY, Villanueva A, Hoshida Y, Peix J, Newell P, Minguez B, et al. Focal gains of VEGFA and molecular classification of hepatocellular carcinoma. Cancer Res. 2008;68:6779–88.
Warburg O, Wind F, Negelein E. The metabolism of tumors in the body. J Gen Physiol. 1927;8:519–30.
Alfarouk KO, Muddathir AK, Shayoub ME. Tumor acidity as evolutionary spite. Cancers (Basel). 2011;3:408–14.
Gatenby RA, Gillies RJ. Why do cancers have high aerobic glycolysis? Nat Rev Cancer. 2004;4:891–9.
Gill KS, Fernandes P, O’Donovan TR, McKenna SL, Doddakula KK, Power DG, et al. Glycolysis inhibition as a cancer treatment and its role in an anti-tumour immune response. Biochim Biophys Acta. 2016;1866:87–105.
Harris AL. Hypoxia–a key regulatory factor in tumour growth. Nat Rev Cancer. 2002;2:38–47.
Karim S, Adams DH, Lalor PF. Hepatic expression and cellular distribution of the glucose transporter family. World J Gastroenterol. 2012;18:6771–81.
Mendez-Blanco C, Fondevila F, Garcia-Palomo A, Gonzalez-Gallego J, Mauriz JL. Sorafenib resistance in hepatocarcinoma: role of hypoxia-inducible factors. Exp Mol Med. 2018;50:134.
Agani F, Jiang BH. Oxygen-independent regulation of HIF-1: novel involvement of PI3K/AKT/mTOR pathway in cancer. Curr Cancer Drug Targets. 2013;13:245–51.
Duvel K, Yecies JL, Menon S, Raman P, Lipovsky AI, Souza AL, et al. Activation of a metabolic gene regulatory network downstream of mTOR complex 1. Mol Cell. 2010;39:171–83.
Han X, Sun S, Zhao M, Cheng X, Chen G, Lin S, et al. Celastrol stimulates hypoxia-inducible factor-1 activity in tumor cells by initiating the ROS/Akt/p70S6K signaling pathway and enhancing hypoxia-inducible factor-1alpha protein synthesis. PLoS ONE. 2014;9:e112470.
Matter MS, Decaens T, Andersen JB, Thorgeirsson SS. Targeting the mTOR pathway in hepatocellular carcinoma: current state and future trends. J Hepatol. 2014;60:855–65.
Kunter I, Erdal E, Nart D, Yilmaz F, Karademir S, Sagol O, et al. Active form of AKT controls cell proliferation and response to apoptosis in hepatocellular carcinoma. Oncol Rep. 2014;31:573–80.
Khemlina G, Ikeda S, Kurzrock R. The biology of hepatocellular carcinoma: implications for genomic and immune therapies. Mol Cancer. 2017;16:149.
Bhat M, Sonenberg N, Gores GJ. The mTOR pathway in hepatic malignancies. Hepatology. 2013;58:810–8.
Zhang Y, Kwok-Shing NgP, Kucherlapati M, Chen F, Liu Y, Tsang YH. et al. A pan-cancer proteogenomic atlas of PI3K/AKT/mTOR pathway alterations. Cancer Cell. 2017;31:820–32. e3.
Li X, Lonard DM, Jung SY, Malovannaya A, Feng Q, Qin J, et al. The SRC-3/AIB1 coactivator is degraded in a ubiquitin- and ATP-independent manner by the REGgamma proteasome. Cell 2006;124:381–92.
Chen X, Barton LF, Chi Y, Clurman BE, Roberts JM. Ubiquitin-independent degradation of cell-cycle inhibitors by the REGgamma proteasome. Mol Cell. 2007;26:843–52.
Li X, Amazit L, Long W, Lonard DM, Monaco JJ, O’Malley BW. Ubiquitin- and ATP-independent proteolytic turnover of p21 by the REGgamma-proteasome pathway. Mol Cell. 2007;26:831–42.
Yao L, Zhou L, Xuan Y, Zhang P, Wang X, Wang T, et al. The proteasome activator REGgamma counteracts immunoproteasome expression and autoimmunity. J Autoimmun. 2019;103:102282.
Xiong S, Zheng Y, Jiang P, Liu R, Liu X, Qian J, et al. PA28gamma emerges as a novel functional target of tumour suppressor microRNA-7 in non-small-cell lung cancer. Br J Cancer. 2014;110:353–62.
Wang Q, Gao X, Yu T, Yuan L, Dai J, Wang W, et al. REGgamma controls hippo signaling and reciprocal NF-kappaB-YAP regulation to promote colon cancer. Clin Cancer Res. 2018;24:2015–25.
Guo J, Hao J, Jiang H, Jin J, Wu H, Jin Z, et al. Proteasome activator subunit 3 promotes pancreatic cancer growth via c-Myc-glycolysis signaling axis. Cancer Lett. 2017;386:161–7.
Hindupur SK, Colombi M, Fuhs SR, Matter MS, Guri Y, Adam K, et al. The protein histidine phosphatase LHPP is a tumour suppressor. Nature. 2018;555:678–82.
Alqahtani A, Khan Z, Alloghbi A, Said Ahmed TS, Ashraf M, Hammouda DM. Hepatocellular carcinoma: molecular mechanisms and targeted therapies. Medicina (Kaunas). 2019;55:526.
Obre E, Rossignol R. Emerging concepts in bioenergetics and cancer research: metabolic flexibility, coupling, symbiosis, switch, oxidative tumors, metabolic remodeling, signaling and bioenergetic therapy. Int J Biochem Cell Biol. 2015;59:167–81.
Munoz-Sanchez J, Chanez-Cardenas ME. The use of cobalt chloride as a chemical hypoxia model. J Appl Toxicol. 2019;39:556–70.
Ding M, Bruick RK, Yu Y. Secreted IGFBP5 mediates mTORC1-dependent feedback inhibition of IGF-1 signalling. Nat Cell Biol. 2016;18:319–27.
Chong ZZ. Targeting PRAS40 for multiple diseases. Drug Disco Today. 2016;21:1222–31.
Sun W, Yu Y, Dotti G, Shen T, Tan X, Savoldo B, et al. PPM1A and PPM1B act as IKKbeta phosphatases to terminate TNFalpha-induced IKKbeta-NF-kappaB activation. Cell Signal. 2009;21:95–102.
Gao X, Wang Q, Wang Y, Liu J, Liu S, Liu J, et al. The REGgamma inhibitor NIP30 increases sensitivity to chemotherapy in p53-deficient tumor cells. Nat Commun. 2020;11:3904.
Mi W, Ye Q, Liu S, She QB. AKT inhibition overcomes rapamycin resistance by enhancing the repressive function of PRAS40 on mTORC1/4E-BP1 axis. Oncotarget. 2015;6:13962–77.
Manning BD, Cantley LC. Rheb fills a GAP between TSC and TOR. Trends Biochem Sci. 2003;28:573–6.
He J, Cui L, Zeng Y, Wang G, Zhou P, Yang Y, et al. REGgamma is associated with multiple oncogenic pathways in human cancers. BMC Cancer. 2012;12:75.
Moncsek A, Gruner M, Meyer H, Lehmann A, Kloetzel PM, Stohwasser R. Evidence for anti-apoptotic roles of proteasome activator 28gamma via inhibiting caspase activity. Apoptosis 2015;20:1211–28.
Zannini L, Lecis D, Buscemi G, Carlessi L, Gasparini P, Fontanella E, et al. REGgamma proteasome activator is involved in the maintenance of chromosomal stability. Cell Cycle. 2008;7:504–12.
DeBerardinis RJ, Lum JJ, Hatzivassiliou G, Thompson CB. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab. 2008;7:11–20.
Janssens V, Goris J. Protein phosphatase 2A: a highly regulated family of serine/threonine phosphatases implicated in cell growth and signalling. Biochem J. 2001;353:417–39.
Lin D, Wu J. Hypoxia inducible factor in hepatocellular carcinoma: a therapeutic target. World J Gastroenterol. 2015;21:12171–8.
Li S, Jiang C, Pan J, Wang X, Jin J, Zhao L, et al. Regulation of c-Myc protein stability by proteasome activator REGgamma. Cell Death Differ. 2015;22:1000–11.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
European Association For The Study Of The L, European Organisation For R, and Treatment Of C. EASL-EORTC clinical practice guidelines: management of hepatocellular carcinoma. J Hepatol. 2012;56:908–43.
Dhanasekaran R, Venkatesh SK, Torbenson MS, Roberts LR. Clinical implications of basic research in hepatocellular carcinoma. J Hepatol. 2016;64:736–45.
Zhu AX, Kudo M, Assenat E, Cattan S, Kang YK, Lim HY, et al. Effect of everolimus on survival in advanced hepatocellular carcinoma after failure of sorafenib: the EVOLVE-1 randomized clinical trial. JAMA. 2014;312:57–67.
Miyamoto H, Moriishi K, Moriya K, Murata S, Tanaka K, Suzuki T, et al. Involvement of the PA28gamma-dependent pathway in insulin resistance induced by hepatitis C virus core protein. J Virol. 2007;81:1727–35.
Moriishi K, Mochizuki R, Moriya K, Miyamoto H, Mori Y, Abe T, et al. Critical role of PA28gamma in hepatitis C virus-associated steatogenesis and hepatocarcinogenesis. Proc Natl Acad Sci USA. 2007;104:1661–6.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31730017, 31670882, 81672883), the Science and Technology Commission of Shanghai Municipality (19JC1411900, 16ZR1410000, 16QA1401500). We thank Dr. Wei Liu from Zhejiang University for providing TSC2−/− cell line. We also thank the ECNU Multifunctional Platform for Innovation (011) for keeping and raising mice.
Author information
Authors and Affiliations
Contributions
X.T.L., B.H.Z., J.B.L., and L.F.Y. designed research. L.F.Y. and Y.X. performed the molecular and cell biology, metabolic, data analysis and animal experiments, respectively. X.L.M., T.Y.M., and X.Q.M. were involved in the molecular and cell biology study. H.Y.Z., T.Z.W., W.S.S., and H.B.W. contributed to the animal work. B.Y. contributed to metabolomics analysis. J.R.X., J.J.L., and C.G. provided clinical samples. L.L. and P.Z. contributed to material support. X.T.L., B.H.Z., R.M., and L.F.Y. wrote the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yao, L., Xuan, Y., Zhang, H. et al. Reciprocal REGγ-mTORC1 regulation promotes glycolytic metabolism in hepatocellular carcinoma. Oncogene 40, 677–692 (2021). https://doi.org/10.1038/s41388-020-01558-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-01558-8
This article is cited by
-
B3galt5 deficiency attenuates hepatocellular carcinoma by suppressing mTOR/p70s6k-mediated glycolysis
Cellular and Molecular Life Sciences (2023)
-
Identification of a BRAF/PA28γ/MEK1 signaling axis and its role in epithelial-mesenchymal transition in oral submucous fibrosis
Cell Death & Disease (2022)
-
RETRACTED ARTICLE: LNCAROD enhances hepatocellular carcinoma malignancy by activating glycolysis through induction of pyruvate kinase isoform PKM2
Journal of Experimental & Clinical Cancer Research (2021)