Abstract
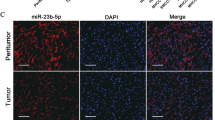
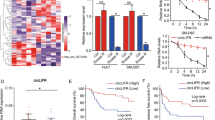
Cumulative evidence suggests that microRNAs (miRNAs) promote gene expression in cancers. However, the pathophysiologic relevance of miRNA-mediated RNA activation in hepatocellular carcinoma (HCC) remains to be established. Our previous miRNA expression profiling in seven-paired HCC specimens revealed miR-93-5p as an HCC-related miRNA. In this study, miR-93-5p expression was assessed in HCC tissues and cell lines by quantitative real-time PCR and fluorescence in situ hybridization. The correlation of miR-93-5p expression with survival and clinicopathological features of HCC was determined by statistical analysis. The function and potential mechanism of miR-93-5p in HCC were further investigated by a series of gain- or loss-of-function experiments in vitro and in vivo. We identified that miR-93-5p, overexpressed in HCC specimens and cell lines, leads to poor outcomes in HCC cases and promotes proliferation, migration, and invasion in HCC cell lines. Mechanistically, rather than decreasing target mRNA levels as expected, miR-93-5p binds to the 3′-untranslated region (UTR) of mitogen-activated protein kinase kinase kinase 2 (MAP3K2) to directly upregulate its expression and downstream p38 and c-Jun N-terminal kinase (JNK) pathway, thereby leading to cell cycle progression in HCC. Notably, we also demonstrated that c-Jun, a downstream effector of the JNK pathway, enhances miR-93-5p transcription by targeting its promoter region. Besides, downregulation of miR-93-5p significantly retarded tumor growth, while overexpression of miR-93-5p accelerated tumor growth in the HCC xenograft mouse model. Altogether, we revealed a miR-93-5p/MAP3K2/c-Jun positive feedback loop to promote HCC progression in vivo and in vitro, representing an RNA-activating role of miR-93-5p in HCC development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Johnson JM, Edwards S, Shoemaker D, Schadt EE. Dark matter in the genome: evidence of widespread transcription detected by microarray tiling experiments. Trends Genet. 2005;21:93–102.
Sadri Nahand J, Bokharaei-Salim F, Salmaninejad A, Nesaei A, Mohajeri F, Moshtzan A, et al. microRNAs: Key players in virus-associated hepatocellular carcinoma. J Cell Physiol. 2019;234:12188–225.
Li N, Miao Y, Shan Y, Liu B, Li Y, Zhao L, et al. MiR-106b and miR-93 regulate cell progression by suppression of PTEN via PI3K/Akt pathway in breast cancer. Cell Death Dis. 2017;8:e2796.
Du L, Zhao Z, Ma X, Hsiao TH, Chen Y, Young E, et al. miR-93-directed downregulation of DAB2 defines a novel oncogenic pathway in lung cancer. Oncogene. 2014;33:4307–15.
Zhu HR, Huang RZ, Yu XN, Shi X, Bilegsaikhan E, Guo HY, et al. Microarray expression profiling of microRNAs reveals potential biomarkers for hepatocellular carcinoma. Tohoku J Exp Med. 2018;245:89–98.
Ohta K, Hoshino H, Wang J, Ono S, Iida Y, Hata K, et al. MicroRNA-93 activates c-Met/PI3K/Akt pathway activity in hepatocellular carcinoma by directly inhibiting PTEN and CDKN1A. Oncotarget. 2015;6:3211–24.
Xue XF, Wang XN, Zhao YB, Hu RK, Qin L. Exosomal miR-93 promotes proliferation and invasion in hepatocellular carcinoma by directly inhibiting TIMP2/TP53INP1/CDKN1A. Biochem Biophys Res Commun. 2018;502:515–21.
Ji CM, Liu H, Yin Q, Li H, Gao H. miR-93 enhances hepatocellular carcinoma invasion and metastasis by EMT via targeting PDCD4. Biotechnol Lett. 2017;39:1621–9.
Chaluvally-Raghavan P, Jeong KJ, Pradeep S, Silva AM, Yu S, Liu W, et al. Direct upregulation of STAT3 by microRNA-551b-3p deregulates growth and metastasis of ovarian cancer. Cell Rep. 2016;15:1493–504.
Xiao M, Li J, Li W, Wang Y, Wu FZ, Xi YP, et al. MicroRNAs activate gene transcription epigenetically as an enhancer trigger. Rna Biol. 2017;14:1326–34.
Orom UA, Nielsen FC, Lund AH. MicroRNA-10a binds the 5′ UTR of ribosomal protein mRNAs and enhances their translation. Mol Cell. 2008;30:460–71.
Vasudevan S, Tong YC, Steitz JA. Switching from repression to activation: MicroRNAs can up-regulate translation. Science. 2007;318:1931–4.
Delire B, Starkel P. The Ras/MAPK pathway and hepatocarcinoma: pathogenesis and therapeutic implications. Eur J Clin Investig. 2015;45:609–23.
Wagner EF, Nebreda AR. Signal integration by JNK and p38 MAPK pathways in cancer development. Nat Rev Cancer. 2009;9:537–49.
Yu J, Tan QY, Deng B, Fang CS, Qi D, Wang RW. The microRNA-520a-3p inhibits proliferation, apoptosis and metastasis by targeting MAP3K2 in non-small cell lung cancer. Am J Cancer Res. 2015;5:802–11.
Wu J, Li WZ, Huang ML, Wei HL, Wang T, Fan J, et al. Regulation of cancerous progression and epithelial-mesenchymal transition by miR-34c-3p via modulation of MAP3K2 signaling in triple-negative breast cancer cells. Biochem Biophys Res Commun. 2017;483:10–6.
Wang M, Lv G, Jiang C, Xie S, Wang G. miR-302a inhibits human HepG2 and SMMC-7721 cells proliferation and promotes apoptosis by targeting MAP3K2 and PBX3. Sci Rep. 2019;9:2032.
Fang L, Deng Z, Shatseva T, Yang J, Peng C, Du WW, et al. MicroRNA miR-93 promotes tumor growth and angiogenesis by targeting integrin-beta8. Oncogene. 2011;30:806–21.
Li Y, Tan W, Neo TW, Aung MO, Wasser S, Lim SG, et al. Role of the miR-106b-25 microRNA cluster in hepatocellular carcinoma. Cancer Sci. 2009;100:1234–42.
Vasaikar SV, Straub P, Wang J, Zhang B. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res. 2018;46:D956–63.
Niu Y, Jin Y, Deng SC, Deng SJ, Zhu S, Liu Y, et al. MiRNA-646-mediated reciprocal repression between HIF-1alpha and MIIP contributes to tumorigenesis of pancreatic cancer. Oncogene. 2018;37:1743–58.
Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014;42:D92–7.
Yang JH, Li JH, Shao P, Zhou H, Chen YQ, Qu LH. starBase: a database for exploring microRNA-mRNA interaction maps from Argonaute CLIP-Seq and Degradome-Seq data. Nucleic Acids Res. 2011;39:D202–9.
Seki E, Brenner DA, Karin M. A liver full of JNK: signaling in regulation of cell function and disease pathogenesis, and clinical approaches. Gastroenterology. 2012;143:307–20.
Wang CH, Tsao YP, Chen HJ, Chen HL, Wang HW, Chen SL. Transcriptional repression of p21((Waf1/Cip1/Sdi1)) gene by c-jun through Sp1 site. Biochem Biophys Res Commun. 2000;270:303–10.
Gong C, Qu S, Liu B, Pan S, Jiao Y, Nie Y, et al. MiR-106b expression determines the proliferation paradox of TGF-beta in breast cancer cells. Oncogene. 2015;34:84–93.
Chen X, Chen S, Xiu YL, Sun KX, Zong ZH, Zhao Y. RhoC is a major target of microRNA-93-5P in epithelial ovarian carcinoma tumorigenesis and progression. Mol Cancer. 2015;14:11.
Chen C, Zheng Q, Kang W, Yu C. Long non-coding RNA LINC00472 suppresses hepatocellular carcinoma cell proliferation, migration and invasion through miR-93-5p/PDCD4 pathway. Clin Res Hepatol Gastroenterol. 2019;43:436–45.
Petrocca F, Visone R, Onelli MR, Shah MH, Nicoloso MS, de Martino I, et al. E2F1-regulated microRNAs impair TGFbeta-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell. 2008;13:272–86.
Guarnieri AL, Towers CG, Drasin DJ, Oliphant MUJ, Andrysik Z, Hotz TJ, et al. The miR-106b-25 cluster mediates breast tumor initiation through activation of NOTCH1 via direct repression of NEDD4L. Oncogene. 2018;37:3879–93.
Poliseno L, Salmena L, Riccardi L, Fornari A, Song MS, Hobbs RM, et al. Identification of the miR-106b~25 microRNA cluster as a proto-oncogenic PTEN-targeting intron that cooperates with its host gene MCM7 in transformation. Sci Signal. 2010;3:ra29.
Kim EK, Choi EJ. Pathological roles of MAPK signaling pathways in human diseases. Biochim Biophys Acta. 2010;1802:396–405.
Wang C, Tang K, Li Z, Chen Z, Xu H, Ye Z. Targeted p21(WAF1/CIP1) activation by miR-1236 inhibits cell proliferation and correlates with favorable survival in renal cell carcinoma. Urol Oncol. 2016;34:59.e23–34.
Huang V, Place RF, Portnoy V, Wang J, Qi Z, Jia Z, et al. Upregulation of Cyclin B1 by miRNA and its implications in cancer. Nucleic Acids Res. 2012;40:1695–707.
Pichon X, Wilson LA, Stoneley M, Bastide A, King HA, Somers J, et al. RNA binding protein/RNA element interactions and the control of translation. Curr Protein Pept Sci. 2012;13:294–304.
Fallmann J, Sedlyarov V, Tanzer A, Kovarik P, Hofacker IL. AREsite2: an enhanced database for the comprehensive investigation of AU/GU/U-rich elements. Nucleic Acids Res. 2016;44:D90–5.
Sun Q, Yang Z, Li P, Wang X, Sun L, Wang S, et al. A novel miRNA identified in GRSF1 complex drives the metastasis via the PIK3R3/AKT/NF-kappaB and TIMP3/MMP9 pathways in cervical cancer cells. Cell Death Dis. 2019;10:636.
Yang Z, Sun Q, Guo J, Wang S, Song G, Liu W, et al. GRSF1-mediated MIR-G-1 promotes malignant behavior and nuclear autophagy by directly upregulating TMED5 and LMNB1 in cervical cancer cells. Autophagy. 2019;15:668–85.
Das M, Jiang F, Sluss HK, Zhang C, Shokat KM, Flavell RA, et al. Suppression of p53-dependent senescence by the JNK signal transduction pathway. Proc Natl Acad Sci USA. 2007;104:15759–64.
Schwabe RF, Bradham CA, Uehara T, Hatano E, Bennett BL, Schoonhoven R, et al. c-Jun-N-terminal kinase drives cyclin D1 expression and proliferation during liver regeneration. Hepatology. 2003;37:824–32.
He Y, Yu B. MicroRNA-93 promotes cell proliferation by directly targeting P21 in osteosarcoma cells. Exp Ther Med. 2017;13:2003–11.
Wong CC, Wong CM, Tung EK, Au SL, Lee JM, Poon RT, et al. The microRNA miR-139 suppresses metastasis and progression of hepatocellular carcinoma by down-regulating Rho-kinase 2. Gastroenterology. 2011;140:322–31.
Liang CC, Park AY, Guan JL. In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat Protoc. 2007;2:329–33.
Yu XN, Zhang GC, Sun JL, Zhu HR, Shi X, Song GQ, et al. Enhanced mLST8 expression correlates with tumor progression in hepatocellular carcinoma. Ann Surg Oncol. 2020;27:1546–57.
Yu XN, Deng Y, Zhang GC, Liu J, Liu TT, Dong L, et al. Sorafenib-conjugated zinc phthalocyanine-based nanocapsule for trimodal therapy in an orthotopic hepatocellular carcinoma xenograft mouse model. ACS Appl Mater Interfaces. 2020;12:17193–206.
Acknowledgements
This work was partly supported by the National Natural Science Foundation of China (grant numbers 81402273, 81672720, 81672334, and 81700550), and by the National Clinical Key Special Subject of China.
Author information
Authors and Affiliations
Contributions
XS, TTL, and XNY data curation, methodology, and writing original draft; AB, HRZ, HYG, GCZ, and EB data curation, formal analysis, and writing review; JLS and GQS investigation, resources, and writing review; SQW and LD investigation, resources, visualization, and writing review; MO conceptualization, and writing review and editing; XZS and JMZ conceptualization, funding acquisition, supervision, and writing review and editing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Shi, X., Liu, TT., Yu, XN. et al. microRNA-93-5p promotes hepatocellular carcinoma progression via a microRNA-93-5p/MAP3K2/c-Jun positive feedback circuit. Oncogene 39, 5768–5781 (2020). https://doi.org/10.1038/s41388-020-01401-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-01401-0
This article is cited by
-
MiRNA-93: a novel signature in human disorders and drug resistance
Cell Communication and Signaling (2023)
-
Non-coding RNA methylation modifications in hepatocellular carcinoma: interactions and potential implications
Cell Communication and Signaling (2023)
-
Bladder Cancer Progression Is Suppressed Through the Heart and Neural Crest Derivatives Expressed 2-Antisense RNA 1/microRNA-93-5p/Defective in Cullin Neddylation 1 Domain Containing 3 Axis
Applied Biochemistry and Biotechnology (2023)
-
Circular RNA MTCL1 promotes advanced laryngeal squamous cell carcinoma progression by inhibiting C1QBP ubiquitin degradation and mediating beta-catenin activation
Molecular Cancer (2022)