Abstract
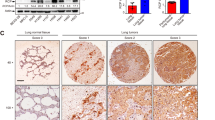
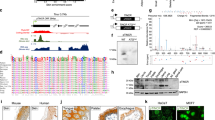
Plakophilin 1 (PKP1) is a member of the arm-repeat (armadillo) and plakophilin gene families and it is an essential component of the desmosomes. Although desmosomes have generally been associated with tumor suppressor functions, we have consistently observed that PKP1 is among the top overexpressed proteins in squamous cell lung cancer. To explore this paradox, we developed in vivo and in vitro functional models of PKP1 gain/loss in squamous cell lung cancer. CRISPR-Cas9 PKP1 knockout severely impaired cell proliferation, but it increased cell dissemination. In addition, PKP1 overexpression increased cell proliferation, cell survival, and in vivo xenograft engraftment. We further investigated the molecular mechanism of the mainly oncogenic function of PKP1 by combining transcriptomics, proteomics, and protein-nucleic acid interaction assays. Interestingly, we found that PKP1 enhances MYC translation in collaboration with the translation initiation complex by binding to the 5′-UTR of MYC mRNA. We propose PKP1 as an oncogene in SqCLC and a novel posttranscriptional regulator of MYC. PKP1 may be a valuable diagnostic biomarker and potential therapeutic target for SqCLC. Importantly, PKP1 inhibition may indirectly target MYC, a primary anticancer target.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The microarray data have been deposited on Gene Expression Omnibus (GEO) under the accession number GSE106770.
Change history
08 February 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41388-019-1147-1
References
Bender E. Epidemiology: the dominant malignancy. Nature. 2014;513:S2. https://doi.org/10.1038/513S2a.
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. https://doi.org/10.3322/caac.21492. Epub 12 Sep 2018.
Molina JR, Yang P, Cassivi SD, Schild SE, Adjei AA. Non-small cell lung cancer: epidemiology, risk factors, treatment, and survivorship. Mayo Clin Proc. 2008;83:584–94. http://www.ncbi.nlm.nih.gov/pubmed/18452692.
Collisson EA, Campbell JD, Brooks AN, Berger AH, Lee W, Chmielecki J. Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511:543–550. https://doi.org/10.1038/nature13385. Epub 9 July 2014.
Kim HR, Shim HS, Chung JH, Lee YJ, Hong YK, Rha SY. Distinct clinical features and outcomes in never-smokers with nonsmall cell lung cancer who harbor EGFR or KRAS mutations or ALK rearrangement. Cancer. 2012;118:729–739. https://doi.org/10.1002/cncr.26311. Epub 30 June 2011.
Angulo B, Suarez-Gauthier A, Lopez-Rios F, Medina PP, Conde E, Tang M. et al. Expression signatures in lung cancer reveal a profile for EGFR-mutant tumours and identify selective PIK3CA overexpression by gene amplification. J Pathol. 2008;214:347–56. https://onlinelibrary.wiley.com/doi/abs/10.1002/path.2267.
Sanchez-Palencia A, Gomez-Morales M, Gomez-Capilla JA, Pedraza V, Boyero L, Rosell R, et al. Gene expression profiling reveals novel biomarkers in nonsmall cell lung cancer. Int J Cancer. 2011;129:355–64. https://onlinelibrary.wiley.com/doi/abs/10.1002/ijc.25704.
Perez-Moreno P, Brambilla E, Thomas R, Soria JC. Squamous cell carcinoma of the lung: molecular subtypes and therapeutic opportunities. Clin Cancer Res. 2012;18:2443–2451. https://doi.org/10.1158/1078-0432.CCR-11-2370. Epub 8 Mar 2012.
Hatzfeld M. Plakophilins: multifunctional proteins or just regulators of desmosomal adhesion? Biochim Biophys Acta. 2007;1773:69–77. http://www.sciencedirect.com/science/article/pii/S0167488906000954.
Krunic AL, Garrod DR, Madani S, Buchanan MD, Clark RE. Immunohistochemical staining for desmogleins 1 and 2 in keratinocytic neoplasms with squamous phenotype: actinic keratosis, keratoacanthoma and squamous cell carcinoma of the skin. Br J Cancer. 1998;77:1275–9. https://www.ncbi.nlm.nih.gov/pubmed/9579833.
Dusek RL, Attardi LD. Desmosomes: new perpetrators in tumour suppression. Nat Rev Cancer. 2011;11:317. https://doi.org/10.1038/nrc3051.
Kundu ST, Gosavi P, Khapare N, Patel R, Hosing AS, Maru GB, et al. Plakophilin3 downregulation leads to a decrease in cell adhesion and promotes metastasis. Int J Cancer. 2008;123:2303–14. https://onlinelibrary.wiley.com/doi/abs/10.1002/ijc.23797.
Richardson G, Johnson BE. The biology of lung cancer. Semin Oncol. 1993;20:105–27.
Bhattacharjee A, Richards WG, Staunton J, Li C, Monti S, Vasa P,et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc Natl Acad Sci. 2001;98:13790–5. https://www.pnas.org/content/98/24/13790.
Hou J, Aerts J, den Hamer B, van IJcken W, den Bakker M, Riegman P, et al. Gene expression-based classification of non-small cell lung carcinomas and survival prediction. PLoS ONE. 2010;5:1–12. https://doi.org/10.1371/journal.pone.0010312.
TCGA Research Network. 2018. https://cancergenome.nih.gov/.
Furukawa C, Daigo Y, Ishikawa N, Kato T, Ito T, Tsuchiya E, et al. Plakophilin 3 oncogene as prognostic marker and therapeutic target for lung cancer. Cancer Res. 2005;65:7102–10. http://cancerres.aacrjournals.org/content/65/16/7102.
Schmidt A, Langbein L, Rode M, Prätzel S, Zimbelmann R, Franke W. Plakophilins 1a and 1b: Widespread nuclear proteins recruited in specific epithelial cells as desmosomal plaque components. Cell Tissue Res. 1997;290:481–99.
Clark GJ, Cox AD, Graham SM, Der CJ. Biological assays for Ras transformation. Methods Enzymol. 1995;255:395–412. https://doi.org/10.1016/s0076-6879(95)55042-9.
Liang C-C, Park AY, Guan J-L. In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat Protoc. 2007;2:329. https://doi.org/10.1038/nprot.2007.30.
Hatzfeld M, Haffner C, Schulze K, Vinzens U. The function of plakophilin 1 in desmosome assembly and actin filament organization. J Cell Biol. 2000;149:209–22. https://www.ncbi.nlm.nih.gov/pubmed/10747098.
McGrath JA, McMillan JR, Shemanko CS, Runswick SK, Leigh IM, Lane EB, et al. Mutations in the plakophilin 1 gene result in ectodermal dysplasia/skin fragility syndrome. Nat Genet. 1997;17:240. https://doi.org/10.1038/ng1097-240.
Consortium Gte. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 2013;45:580–5. https://www.ncbi.nlm.nih.gov/pubmed/23715323.
Budczies J, von Winterfeld M, Klauschen F, Bockmayr M, Lennerz JK, Denkert C, et al. The landscape of metastatic progression patterns across major human cancers. Oncotarget. 2015;6:570–83. https://www.ncbi.nlm.nih.gov/pubmed/25402435.
Gomez-Morales M, Camara-Pulido M, Miranda-Leon MT, Sanchez-Palencia A, Boyero L, Gomez-Capilla JA, et al. Differential immunohistochemical localization of desmosomal plaque-related proteins in non-small-cell lung cancer. Histopathology. 2013;63:103–13. https://www.ncbi.nlm.nih.gov/pubmed/23711109.
Alatas ET, Kara A, Kara M, Dogan G, Baysal O. Ectodermal dysplasia-skin fragility syndrome with a new mutation. Indian J Dermatol Venereol Leprol. 2017;83:476–9. https://www.ncbi.nlm.nih.gov/pubmed/28540868e.
Rietscher K, Wolf A, Hause G, Rother A, Keil R, Magin TM, et al. Growth retardation, loss of desmosomal adhesion, and impaired tight junction function identify a unique role of plakophilin 1 in-vivo. J Invest Dermatol. 2016;136:1471–8. https://doi.org/10.1016/j.jid.2016.03.021.
Pirity M, Blanck JK, Schreiber-Agus N. Lessons learned from Myc/Max/Mad knockout mice. Curr Top Microbiol Immunol. 2006;302:205–34. https://www.ncbi.nlm.nih.gov/pubmed/16620030.
Whitfield JR, Beaulieu M-E, Soucek L. Strategies to Inhibit Myc and Their Clinical Applicability. Front cell Dev Biol. 2017;5:10. https://www.ncbi.nlm.nih.gov/pubmed/28280720.
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–308. https://www.ncbi.nlm.nih.gov/pubmed/24157548.
Munoz WA, Lee M, Miller RK, Ahmed Z, Ji H, Link TM. Plakophilin-3 catenin associates with the ETV1/ER81 transcription factor to positively modulate gene activity. PLoS ONE. 2014;9:e86784. https://doi.org/10.1371/journal.pone.0086784.eCollection 2014.
Acknowledgements
We thank the PhD program in Biochemistry and Molecular Biology of the University of Granada and also Montse Sanchez-Céspedes for the critical read of the paper.
Funding
J-M-P and LB were supported by fellowships from Fundación Anticancer San Francisco Javier y Santa Cándida/UGR. AA was supported by a PhD FPU fellowship (FPU17/00067). PP was supported by a PhD “La Caixa Foundation” LCF/BQ/DE15/10360019 Fellowship. IFC was supported by a PhD FPI-fellowship (BES-2013-064596). JCA-P is supported by a Marie Sklodowska Curie action (H2020-MSCA-IF-2018). CB and MIR were supported by Consejería de Sanidad de la Junta de Andalucía (Pl-0245-2017). MEF-V was supported by PAIDI program. Group BIO309. Junta de Andalucía, and Instituto de Salud Carlos III-Fondo de Investigación Sanitaria PI10/00198. PPM laboratory is funded by the Ministry of Economy of Spain (SAF2015-67919-R), Junta de Andalucía (P12-BIO-1655, PAIDI CTS-993), Francisco Cobos Foundation, FERO foundation, International Association for the study of lung cancer (IASLC) and Fundación Científica de la Asociación Española Contra el Cáncer (Lab AECC-2018). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the paper. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
JM-P, LB, MIR, PP, ID-C, CB, IFC, JCA-P performed and validated the experimental data. AA performed most of bioinformatics and statistical analysis. PPM and MEF-V wrote the paper, reviewed and edited the manuscript and provided conceptualization, validation, supervision, expertise and feedback. MEF-V and PPM, participated in the funding acquisition and project administration.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This project has been approved by the institutional ethical committee of the University of Granada.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Martin-Padron, J., Boyero, L., Rodriguez, M.I. et al. Plakophilin 1 enhances MYC translation, promoting squamous cell lung cancer. Oncogene 39, 5479–5493 (2020). https://doi.org/10.1038/s41388-019-1129-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-1129-3
This article is cited by
-
Combined inhibition of histone deacetylase and cytidine deaminase improves epigenetic potency of decitabine in colorectal adenocarcinomas
Clinical Epigenetics (2023)
-
PKP1 and MYC create a feedforward loop linking transcription and translation in squamous cell lung cancer
Cellular Oncology (2022)
-
Protein co-expression network-based profiles revealed from laser-microdissected cancerous cells of lung squamous-cell carcinomas
Scientific Reports (2021)
-
CRISPR/Cas9 library screening uncovered methylated PKP2 as a critical driver of lung cancer radioresistance by stabilizing β-catenin
Oncogene (2021)
-
Identification of PKP 2/3 as potential biomarkers of ovarian cancer based on bioinformatics and experiments
Cancer Cell International (2020)