Abstract
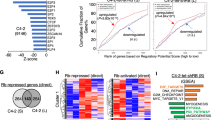
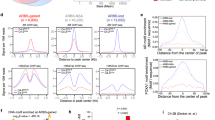
B lymphoma Mo-MLV insertion region 1 (BMI1) has been reported to be an oncoprotein. BMI1 represses tumor suppressors to promote cell proliferation, epithelial-mesenchymal transition (EMT), and cancer progression. Although it is known that the expression of BMI1 is increased in many cancer types, the mechanism of BMI1 upregulation is not yet clear. We performed integrative analysis for 3 sets of prostate cancer (PCa) genomic data, and found that BMI1 and androgen receptor (AR) were positively correlated, suggesting that AR might regulate BMI1. Next, we showed that dihydrotestosterone (DHT) upregulated both mRNA and protein levels of BMI1 and that BMI1 was increased in castration-resistant prostate cancer (CRPC) from both human patients and a mouse xenograph model. We further identified an AR binding site in the promoter/enhancer region of BMI1, and confirmed BMI1 as the direct target of AR using gene-editing technology. We also demonstrated that high expression of BMI1 is critical for the development of castration-resistance. Our data also suggest that BMI1-specific inhibitors could be an effective treatment of CRPC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The next-generation sequencing data haves been deposited into Gene Expression Omnibus (GEO) under accession GSE97831.
References
Song LB, Li J, Liao WT, Feng Y, Yu CP, Hu LJ, et al. The polycomb group protein Bmi-1 represses the tumor suppressor PTEN and induces epithelial-mesenchymal transition in human nasopharyngeal epithelial cells. J Clin Invest. 2009;119:3626–36.
Ren H, Du P, Ge Z, Jin Y, Ding D, Liu X, et al. TWIST1 and BMI1 in cancer metastasis and chemoresistance. J Cancer. 2016;7:1074–80.
Glinsky GV, Berezovska O, Glinskii AB. Microarray analysis identifies a death-from-cancer signature predicting therapy failure in patients with multiple types of cancer. J Clin Invest. 2005;115:1503–21.
van Leenders GJ, Dukers D, Hessels D, van den Kieboom SW, Hulsbergen CA, Witjes JA, et al. Polycomb-group oncogenes EZH2, BMI1, and RING1 are overexpressed in prostate cancer with adverse pathologic and clinical features. Eur Urol. 2007;52:455–63.
Siddique HR, Saleem M. Role of BMI1, a stem cell factor, in cancer recurrence and chemoresistance: preclinical and clinical evidences. Stem Cells. 2012;30:372–8.
Zhu S, Zhao D, Yan L, Jiang W, Kim JS, Gu B, et al. BMI1 regulates androgen receptor in prostate cancer independently of the polycomb repressive complex 1. Nat Commun. 2018;9:500.
Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS, et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18:11–22.
Grasso CS, Wu YM, Robinson DR, Cao X, Dhanasekaran SM, Khan AP, et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature. 2012;487:239–43.
Wang W, Epstein JI. Small cell carcinoma of the prostate. A morphologic and immunohistochemical study of 95 cases. Am J Surg Pathol. 2008;32:65–71.
Grant CE, Bailey TL, Noble WS. FIMO: scanning for occurrences of a given motif. Bioinformatics. 2011;27:1017–8.
Mounir Z, Korn JM, Westerling T, Lin F, Kirby CA, Schirle M et al. ERG signaling in prostate cancer is driven through PRMT5-dependent methylation of the androgen receptor. Elife 2016; 5:pii: e13964.
Takayama K, Suzuki T, Fujimura T, Urano T, Takahashi S, Homma Y, et al. CtBP2 modulates the androgen receptor to promote prostate cancer progression. Cancer Res. 2014;74:6542–53.
Kron KJ, Murison A, Zhou S, Huang V, Yamaguchi TN, Shiah YJ, et al. TMPRSS2-ERG fusion co-opts master transcription factors and activates NOTCH signaling in primary prostate cancer. Nat Genet. 2017;49:1336–45.
Bansal N, Bartucci M, Yusuff S, Davis S, Flaherty K, Huselid E, et al. BMI-1 targeting interferes with patient-derived tumor-initiating cell survival and tumor growth in prostate cancer. Clin Cancer Res. 2016;22:6176–91.
Yong KJ, Basseres DS, Welner RS, Zhang WC, Yang H, Yan B, et al. Targeted BMI1 inhibition impairs tumor growth in lung adenocarcinomas with low CEBPα expression. Sci Transl Med. 2016;8:350ra104.
Kreso A, van Galen P, Pedley NM, Lima-Fernandes E, Frelin C, Davis T, et al. Self-renewal as a therapeutic target in human colorectal cancer. Nat Med. 2014;20:29–36.
Mourgues L, Imbert V, Nebout M, Colosetti P, Neffati Z, Lagadec P, et al. The BMI1 polycomb protein represses cyclin G2-induced autophagy to support proliferation in chronic myeloid leukemia cells. Leukemia. 2015;29:1993–2002.
Nishida Y, Maeda A, Kim MJ, Cao L, Kubota Y, Ishizawa J, et al. The novel BMI-1 inhibitor PTC596 downregulates MCL-1 and induces p53-independent mitochondrial apoptosis in acute myeloid leukemia progenitor cells. Blood Cancer J. 2017;7:e527.
Jin X, Kim LJY, Wu Q, Wallace LC, Prager BC, Sanvoranart T, et al. Targeting glioma stem cells through combined BMI1 and EZH2 inhibition. Nat Med. 2017;23:1352–61.
Kassi E, Moutsatsou P. Glucocorticoid receptor signaling and prostate cancer. Cancer Lett. 2011;302:1–10.
Clyne M. Prostate cancer: androgen deprivation causes EMT in the prostate. Nat Rev Urol. 2011;9:4.
Yoo YA, Roh M, Naseem AF, Lysy B, Desouki MM, Unno K, et al. Bmi1 marks distinct castration-resistant luminal progenitor cells competent for prostate regeneration and tumour initiation. Nat Commun. 2016;7:12943.
Yong KJ, Basseres DS, Welner RS, Zhang WC, Yang H, Yan B, et al. Targeted BMI1 inhibition impairs tumor growth in lung adenocarcinomas with low CEBPalpha expression. Sci Transl Med. 2016;8:350ra104.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Disco. 2012;2:401–4.
Yu Y, Yang O, Fazli L, Rennie PS, Gleave ME, Dong X. Progesterone receptor expression during prostate cancer progression suggests a role of this receptor in stromal cell differentiation. Prostate. 2015;75:1043–50.
Li H, Xie N, Chen R, Verreault M, Fazli L, Gleave ME, et al. UGT2B17 expedites progression of castration-resistant prostate cancers by promoting ligand-independent AR signaling. Cancer Res. 2016;76:6701–11.
Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10:R25.
Chen K, Xi Y, Pan X, Li Z, Kaestner K, Tyler J, et al. DANPOS: dynamic analysis of nucleosome position and occupancy by sequencing. Genome Res. 2013;23:341–51.
Kent WJ, Zweig AS, Barber G, Hinrichs AS, Karolchik D. BigWig and BigBed: enabling browsing of large distributed datasets. Bioinformatics. 2010;26:2204–7.
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, et al. The human genome browser at UCSC. Genome Res. 2002;12:996–1006.
Ji Q, Hao X, Zhang M, Tang W, Yang M, Li L, et al. MicroRNA miR-34 inhibits human pancreatic cancer tumor-initiating cells. PLoS ONE. 2009;4:e6816.
Acknowledgements
We thank Marla Weetall, John Baird, Art Branstrom and PTC Therapeutics for providing PTC596 and valuable inputs. We thank The University of Texas MD Anderson Cancer Center Science Park Next-Generation Sequencing (NGS) Facility for assistance with next-generation sequencing, and the Houston Methodist Comparative Medicine core facility, Jenny Chang, Anthony Kozielski, and Wei Qian for assistance with in vivo work. We thank Johnique Atkins for comments and editing this manuscript.
Funding
This work is supported, in part, by grants from Houston Methodist Research Institute, Prostate Cancer Foundation (13YOUN007 to QC), U.S. Department of Defense (W81XWH-15-1-0639 and W81XWH-17-1-0357 to QC), American Cancer Society (Research Scholar Grant RSG-15-192-01-TBE to QC), and NIH/NCI (R01CA208257 to QC); KC is supported in part by grants from NIH/NIGMS (R01GM125632 to KC) and NIH/NHLBI (R01HL133254 to KC); JY is supported by NIH/NCI (R01CA172384), US Department of Defense (W81XWH-17-1-0405, W81XWH-17-1-0362, and W81XWH-17-1-0578). XD is supported by the Canadian Institute of Health Research (#MOP-137007) and TFRI New Frontier Grant #1062. WZ is supported by the National Natural Science Foundation of China (81572766,81972651 and 31771630), Guangdong Innovative and Entrepreneurial Research Team Program (2016ZT06S029), and Natural Science Foundation of Guangdong Province (2017A030312009); CL is supported by the China Scholarship Council (201706370147). TD is supported by National Natural Science Foundation of China (81770868 and 91742103), and Innovation-driven Project of Central South University (2017CX011). The University of Texas MD Anderson Cancer Center Science Park Next-Generation Sequencing (NGS) Facility is supported by CPRIT grants RP120348 and RP170002.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Zhu, S., Zhao, D., Li, C. et al. BMI1 is directly regulated by androgen receptor to promote castration-resistance in prostate cancer. Oncogene 39, 17–29 (2020). https://doi.org/10.1038/s41388-019-0966-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0966-4
This article is cited by
-
BMI1 promotes osteosarcoma proliferation and metastasis by repressing the transcription of SIK1
Cancer Cell International (2022)
-
Mechanisms of Polycomb group protein function in cancer
Cell Research (2022)
-
Progression of prostate cancer reprograms MYC-mediated lipid metabolism via lysine methyltransferase 2A
Discover Oncology (2022)
-
LncRNA MAFG-AS1 Promotes Lung Adenocarcinoma Cell Migration and Invasion by Targeting miR-3196 and Regulating SOX12 Expression
Molecular Biotechnology (2022)
-
miR-96-5p, miR-134-5p, miR-181b-5p and miR-200b-3p heterogenous expression in sites of prostate cancer versus benign prostate hyperplasia—archival samples study
Histochemistry and Cell Biology (2021)