Abstract
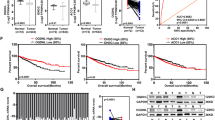
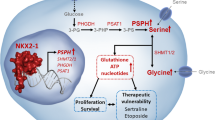
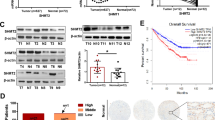
Advanced Bladder Cancer (BLCA) remains a clinical challenge that lacks effective therapeutic measures. Here, we show that distinct, stage-wise metabolic alterations in BLCA are associated with the loss of function of aldehyde oxidase (AOX1). AOX1 associated metabolites have a high predictive value for advanced BLCA and our findings demonstrate that AOX1 is epigenetically silenced during BLCA progression by the methyltransferase activity of EZH2. Knockdown (KD) of AOX1 in normal bladder epithelial cells re-wires the tryptophan-kynurenine pathway resulting in elevated NADP levels which may increase metabolic flux through the pentose phosphate (PPP) pathway, enabling increased nucleotide synthesis, and promoting cell invasion. Inhibition of NADP synthesis rescues the metabolic effects of AOX1 KD. Ectopic AOX1 expression decreases NADP production, PPP flux and nucleotide synthesis, while decreasing invasion in cell line models and suppressing growth in tumor xenografts. Further gain and loss of AOX1 confirm the EZH2-dependent activation, metabolic deregulation, and tumor growth in BLCA. Our findings highlight the therapeutic potential of AOX1 and provide a basis for the development of prognostic markers for advanced BLCA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Change history
17 August 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41388-020-1283-7
References
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–86.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7–30.
Sievert KD, Amend B, Nagele U, Schilling D, Bedke J, Horstmann M, et al. Economic aspects of bladder cancer: what are the benefits and costs? World J Urol. 2009;27:295–300.
Thoma C. Bladder cancer: genomics of noninvasive disease. Nat Rev Urol. 2018;15:1.
Hurst CD, Alder O, Platt FM, Droop A, Stead LF, Burns JE, et al. Genomic subtypes of non-invasive bladder cancer with distinct metabolic profile and female gender bias in KDM6A mutation frequency. Cancer Cell. 2017;32:701–15 e7.
Garattini E, Fratelli M, Terao M. The mammalian aldehyde oxidase gene family. Hum Genom. 2009;4:119–30.
Garattini E, Fratelli M, Terao M. Mammalian aldehyde oxidases: genetics, evolution and biochemistry. Cell Mol Life Sci. 2008;65:1019–48.
Kitamura S, Sugihara K, Ohta S. Drug-metabolizing ability of molybdenum hydroxylases. Drug Metab Pharm. 2006;21:83–98.
Putluri N, Shojaie A, Vasu VT, Vareed SK, Nalluri S, Putluri V, et al. Metabolomic profiling reveals potential markers and bioprocesses altered in bladder cancer progression. Cancer Res. 2011;71:7376–86.
Baylin SB, Herman JG, Graff JR, Vertino PM, Issa JP. Alterations in DNA methylation: a fundamental aspect of neoplasia. Adv Cancer Res. 1998;72:141–96.
Merlo A, Herman JG, Mao L, Lee DJ, Gabrielson E, Burger PC, et al. 5’ CpG island methylation is associated with transcriptional silencing of the tumour suppressor p16/CDKN2/MTS1 in human cancers. Nat Med. 1995;1:686–92.
Esteller M, Tortola S, Toyota M, Capella G, Peinado MA, Baylin SB, et al. Hypermethylation-associated inactivation ofp14(ARF) is independent of p16(INK4a) methylation and p53 mutational status. Cancer Res. 2000;60:129–33.
Di Croce L, Helin K. Transcriptional regulation by Polycomb group proteins. Nat Struct Mol Biol. 2013;20:1147–55.
Sun S, Yu F, Zhang L, Zhou X. EZH2, an on-off valve in signal network of tumor cells. Cell Signal. 2016;28:481–7.
Ma J, Shojaie A, Michailidis G. Network-based pathway enrichment analysis with incomplete network information. Bioinformatics. 2016;32:3165–74.
Kim WJ, Kim EJ, Kim SK, Kim YJ, Ha YS, Jeong P, et al. Predictive value of progression-related gene classifier in primary non-muscle invasive bladder cancer. Mol Cancer. 2010;9:3.
Cancer Genome Atlas Research Network Comprehensive molecular characterization of urothelial bladder carcinoma. Nature. 2014;507:315–22.
Sanchez-Carbayo M, Socci ND, Lozano J, Saint F, Cordon-Cardo C. Defining molecular profiles of poor outcome in patients with invasive bladder cancer using oligonucleotide microarrays. J Clin Oncol. 2006;24:778–89.
Choi W, Porten S, Kim S, Willis D, Plimack ER, Hoffman-Censits J, et al. Identification of distinct basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell. 2014;25:152–65.
McCabe MT, Ott HM, Ganji G, Korenchuk S, Thompson C, Van Aller GS, et al. EZH2 inhibition as a therapeutic strategy for lymphoma with EZH2-activating mutations. Nature. 2012;492:108–12.
Vire E, Brenner C, Deplus R, Blanchon L, Fraga M, Didelot C, et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature. 2006;439:871–4.
Kleer CG, Cao Q, Varambally S, Shen R, Ota I, Tomlins SA, et al. EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells. Proc Natl Acad Sci USA. 2003;100:11606–11.
Oster B, Thorsen K, Lamy P, Wojdacz TK, Hansen LL, Birkenkamp-Demtroder K, et al. Identification and validation of highly frequent CpG island hypermethylation in colorectal adenomas and carcinomas. Int J Cancer. 2011;129:2855–66.
Salter M, Pogson CI. The role of tryptophan 2,3-dioxygenase in the hormonal control of tryptophan metabolism in isolated rat liver cells. Effects of glucocorticoids and experimental diabetes. Biochem J. 1985;229:499–504.
D’Amato NC, Rogers TJ, Gordon MA, Greene LI, Cochrane DR, Spoelstra NS, et al. A TDO2-AhR signaling axis facilitates anoikis resistance and metastasis in triple-negative breast cancer. Cancer Res. 2015;75:4651–64.
Yang J, Weinberg RA. Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev Cell. 2008;14:818–29.
De Craene B, Berx G. Regulatory networks defining EMT during cancer initiation and progression. Nat Rev Cancer. 2013;13:97–110.
Deryugina EI, Quigley JP. Chick embryo chorioallantoic membrane model systems to study and visualize human tumor cell metastasis. Histochem Cell Biol. 2008;130:1119–30.
Kompier LC, Lurkin I, van der Aa MN, van Rhijn BW, van der Kwast TH, Zwarthoff EC. FGFR3, HRAS, KRAS, NRAS and PIK3CA mutations in bladder cancer and their potential as biomarkers for surveillance and therapy. PLoS ONE. 2010;5:e13821.
Vantaku V, Dong J, Ambati CR, Perera D, Donepudi SR, Amara CS, et al. Multi-omics integration analysis robustly predicts high-grade patient survival and identifies CPT1B effect on fatty acid metabolism in Bladder Cancer. Clin Cancer Res. 2019;15:3689–701.
Platten M, Wick W, Van den Eynde BJ. Tryptophan catabolism in cancer: beyond IDO and tryptophan depletion. Cancer Res. 2012;72:5435–40.
Ablain J, de The H. Retinoic acid signaling in cancer: the parable of acute promyelocytic leukemia. Int J Cancer. 2014;135:2262–72.
Yang M, Pollard PJ. Succinate: a new epigenetic hacker. Cancer Cell. 2013;23:709–11.
Zhai L, Spranger S, Binder DC, Gritsina G, Lauing KL, Giles FJ, et al. Molecular pathways: targeting IDO1 and other tryptophan dioxygenases for cancer immunotherapy. Clin Cancer Res. 2015;21:5427–33.
Icard P, Poulain L, Lincet H. Understanding the central role of citrate in the metabolism of cancer cells. Biochim Biophys Acta. 2012;1825:111–6.
Ozturk S, Papageorgis P, Wong CK, Lambert AW, Abdolmaleky HM, Thiagalingam A, et al. SDPR functions as a metastasis suppressor in breast cancer by promoting apoptosis. Proc Natl Acad Sci USA. 2016;113:638–43.
Haldrup C, Mundbjerg K, Vestergaard EM, Lamy P, Wild P, Schulz WA, et al. DNA methylation signatures for prediction of biochemical recurrence after radical prostatectomy of clinically localized prostate cancer. J Clin Oncol. 2013;31:3250–8.
Park JS, Choi SB, Chung JW, Kim SW, Kim DW. Classification of serous ovarian tumors based on microarray data using multicategory support vector machines. Conf Proc IEEE Eng Med Biol Soc. 2014;2014:3430–3.
Varambally S, Dhanasekaran SM, Zhou M, Barrette TR, Kumar-Sinha C, Sanda MG, et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature. 2002;419:624–9.
Bachmann IM, Halvorsen OJ, Collett K, Stefansson IM, Straume O, Haukaas SA, et al. EZH2 expression is associated with high proliferation rate and aggressive tumor subgroups in cutaneous melanoma and cancers of the endometrium, prostate, and breast. J Clin Oncol. 2006;24:268–73.
Sudo T, Utsunomiya T, Mimori K, Nagahara H, Ogawa K, Inoue H, et al. Clinicopathological significance of EZH2 mRNA expression in patients with hepatocellular carcinoma. Br J Cancer. 2005;92:1754–8.
Hussain M, Rao M, Humphries AE, Hong JA, Liu F, Yang M, et al. Tobacco smoke induces polycomb-mediated repression of Dickkopf-1 in lung cancer cells. Cancer Res. 2009;69:3570–8.
Yan J, Ng SB, Tay JL, Lin B, Koh TL, Tan J, et al. EZH2 overexpression in natural killer/T-cell lymphoma confers growth advantage independently of histone methyltransferase activity. Blood. 2013;121:4512–20.
Tan JZ, Yan Y, Wang XX, Jiang Y, Xu HE. EZH2: biology, disease, and structure-based drug discovery. Acta Pharmacol Sin. 2014;35:161–74.
The EZH2 Inhibitor Tazemetostat Is Well Tolerated in a Phase I Trial. Cancer Discov. 2018;8:OF15. http://cancerdiscovery.aacrjournals.org/content/early/2018/04/20/2159-8290.CD-RW2018-067, https://doi.org/10.1158/2159-8290.CD-RW2018-067. Accessed 9 Apr 2018.
Kim KH, Roberts CW. Targeting EZH2 in cancer. Nat Med. 2016;22:128–34.
Dudziec E, Goepel JR, Catto JW. Global epigenetic profiling in bladder cancer. Epigenomics. 2011;3:35–45.
Reinert T, Modin C, Castano FM, Lamy P, Wojdacz TK, Hansen LL, et al. Comprehensive genome methylation analysis in bladder cancer: identification and validation of novel methylated genes and application of these as urinary tumor markers. Clin Cancer Res. 2011;17:5582–92.
Wolff EM, Chihara Y, Pan F, Weisenberger DJ, Siegmund KD, Sugano K, et al. Unique DNA methylation patterns distinguish noninvasive and invasive urothelial cancers and establish an epigenetic field defect in premalignant tissue. Cancer Res. 2010;70:8169–78.
Agledal L, Niere M, Ziegler M. The phosphate makes a difference: cellular functions of NADP. Redox Rep. 2010;15:2–10.
Ying W. NAD+/NADH and NADP+/NADPH in cellular functions and cell death: regulation and biological consequences. Antioxid Redox Signal. 2008;10:179–206.
Prendergast GC. Cancer: why tumours eat tryptophan. Nature. 2011;478:192–4.
Chen Y, Guillemin GJ. Kynurenine pathway metabolites in humans: disease and healthy States. Int J Tryptophan Res. 2009;2:1–19.
Chalkiadaki A, Guarente L. The multifaceted functions of sirtuins in cancer. Nat Rev Cancer. 2015;15:608–24.
Beaconsfield P, Ginsburg J, Jeacock MK. Glucose metabolism via the pentose phosphate pathway relative to nucleic acid and protein synthesis in the human placenta. Dev Med Child Neurol. 1964;6:469–74.
Davidson B, Abeler VM, Forsund M, Holth A, Yang Y, Kobayashi Y, et al. Gene expression signatures of primary and metastatic uterine leiomyosarcoma. Hum Pathol. 2014;45:691–700.
Opitz CA, Litzenburger UM, Sahm F, Ott M, Tritschler I, Trump S, et al. An endogenous tumour-promoting ligand of the human aryl hydrocarbon receptor. Nature. 2011;478:197–203.
Minn AJ, Gupta GP, Siegel PM, Bos PD, Shu W, Giri DD, et al. Genes that mediate breast cancer metastasis to lung. Nature. 2005;436:518–24.
Bos PD, Zhang XH, Nadal C, Shu W, Gomis RR, Nguyen DX, et al. Genes that mediate breast cancer metastasis to the brain. Nature. 2009;459:1005–9.
Jin F, Thaiparambil J, Donepudi SR, Vantaku V, Piyarathna DWB, Maity S, et al. Tobacco-specific carcinogens induce hypermethylation, DNA adducts, and DNA damage in bladder cancer. Cancer Prev Res. 2017;10:588–97.
Piyarathna DWB, Rajendiran TM, Putluri V, Vantaku V, Soni T, von Rundstedt FC, et al. Distinct lipidomic landscapes associated with clinical stages of urothelial cancer of the bladder. Eur Urol Focus. 2018;4:907–915.
Terunuma A, Putluri N, Mishra P, Mathe EA, Dorsey TH, Yi M, et al. MYC-driven accumulation of 2-hydroxyglutarate is associated with breast cancer prognosis. J Clin Invest. 2014;124:398–412.
Putluri N, Maity S, Kommagani R, Creighton CJ, Putluri V, Chen F, et al. Pathway-centric integrative analysis identifies RRM2 as a prognostic marker in breast cancer associated with poor survival and tamoxifen resistance. Neoplasia. 2014;16:390–402.
Putluri N, Shojaie A, Vasu VT, Nalluri S, Vareed SK, Putluri V, et al. Metabolomic profiling reveals a role for androgen in activating amino acid metabolism and methylation in prostate cancer cells. PLoS ONE. 2011;6:e21417.
Bhowmik SK, Ramirez-Pena E, Arnold JM, Putluri V, Sphyris N, Michailidis G, et al. EMT-induced metabolite signature identifies poor clinical outcome. Oncotarget. 2015;6:42651–60.
Estecio MR, Yan PS, Ibrahim AE, Tellez CS, Shen L, Huang TH, et al. High-throughput methylation profiling by MCA coupled to CpG island microarray. Genome Res. 2007;17:1529–36.
Blecher-Gonen R, Barnett-Itzhaki Z, Jaitin D, Amann-Zalcenstein D, Lara-Astiaso D, Amit I. High-throughput chromatin immunoprecipitation for genome-wide mapping of in vivo protein-DNA interactions and epigenomic states. Nat Protoc. 2013;8:539–54.
O’Reilly MS, Boehm T, Shing Y, Fukai N, Vasios G, Lane WS, et al. Endostatin: an endogenous inhibitor of angiogenesis and tumor growth. Cell. 1997;88:277–85.
Li M, Pathak RR, Lopez-Rivera E, Friedman SL, Aguirre-Ghiso JA, Sikora AG. The in ovo chick chorioallantoic membrane (CAM) assay as an efficient xenograft model of hepatocellular carcinoma. J Vis Exp. 2015.
Acknowledgements
This research was fully supported by American Cancer Society (ACS) Award 127430-RSG-15-105-01-CNE (N.P.), NIH/NCI R01CA220297 (N.P), and NIH/NCI R01CA216426 (N.P.), partially supported by the following grants: NIH 1RO1CA133458-01 (A.S.K.), and NIH U01 CA167234, Komen CCR award to S.M.K. (CCR16380599) as well as funds from Alkek Center for Molecular Discovery (A.S.K.). This project was also supported by the Agilent Technologies Center of Excellence in Mass Spectrometry at Baylor College of Medicine, Metabolomics Core, Human Tissue Acquisition and Pathology at Baylor College of Medicine with funding from the NIH (P30 CA125123), CPRIT Proteomics and Metabolomics Core Facility (D.P.E.), (RP170005), and Dan L. Duncan Cancer Center. Imaging for this project was supported by the Integrated Microscopy Core at Baylor College of Medicine with funding from NIH (DK56338, CA125123, and 1S10OD020151-01), CPRIT (RP150578), the Dan L. Duncan Comprehensive Cancer Center, and the John S. Dunn Gulf Coast Consortium for Chemical Genomics. CAM assay was supported by the Patient-Derived Xenograft and Advanced in vivo Models Core Facility at Baylor College of Medicine with funding from the Cancer Prevention and Research Institute of Texas (CPRIT) grant #170691. Imaging for this project was supported by the Integrated Microscopy Core at Baylor College of Medicine with funding from NIH (DK56338, and CA125123), CPRIT (RP150578, RP170719), the Dan L. Duncan Comprehensive Cancer Center, and the John S. Dunn Gulf Coast Consortium for Chemical Genomics. Research reported in this study was supported by the National Cancer Institute of the National Institutes of Health under award number 5 P30 CA142543 09.
Author information
Authors and Affiliations
Contributions
V.V., V.P., A.S. and N.P., conceived the project and wrote the manuscript with editorial input from all of the authors. V.V., V.P., D.A.B., S.M.K., Q.C. and N.P. designed the experiments. V.V., V.P., D.A.B., V.D., S.M.K., S.R.D., J.D., A.K.J., F.S. and B.K. performed the experiments. S.M., J.M., J.A., K.R. and C.C. assisted with the dataset analysis. A.S. and N.P. assisted with mass spectroscopy measurements and functional study. F.V.R. and Y.L. provided clinical specimens. F.V.R. and Y.L. provided clinical input on data interpretation. A.G.S. and H.V. performed the OVIVO experiments.
Corresponding author
Ethics declarations
Conflict of interest
S.M.K. is stake holder of NeoZenome Therapeutics Inc.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Vantaku, V., Putluri, V., Bader, D.A. et al. Epigenetic loss of AOX1 expression via EZH2 leads to metabolic deregulations and promotes bladder cancer progression. Oncogene 39, 6265–6285 (2020). https://doi.org/10.1038/s41388-019-0902-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0902-7
This article is cited by
-
Tumor microenvironment deconvolution identifies cell-type-independent aberrant DNA methylation and gene expression in prostate cancer
Clinical Epigenetics (2024)
-
Integrative analysis reveals associations between oral microbiota dysbiosis and host genetic and epigenetic aberrations in oral cavity squamous cell carcinoma
npj Biofilms and Microbiomes (2024)
-
Comprehensive transcriptomic profiling and mutational landscape of primary gastric linitis plastica
Gastric Cancer (2023)
-
LINC01234 Sponging of the miR-513a-5p/AOX1 Axis is Upregulated in Osteoporosis and Regulates Osteogenic Differentiation of Bone Marrow Mesenchymal Stem Cells
Molecular Biotechnology (2023)
-
The long and short non-coding RNAs modulating EZH2 signaling in cancer
Journal of Hematology & Oncology (2022)