Abstract
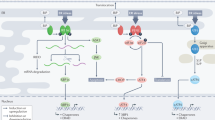
Cancer cells exploit many of the cellular adaptive responses to support their survival needs. One such critical pathway in eukaryotic cells is the unfolded protein response (UPR) that is important in normal physiology as well as disease states, including cancer. Since UPR can serve as a lever between survival and death, regulated control of its activity is critical for tumor formation and growth although the underlying mechanisms are poorly understood. Here we show that one of the main transcriptional effectors of UPR, activating transcription factor 4 (ATF4), is essential for prostate cancer (PCa) growth and survival. Using systemic unbiased gene expression and proteomic analyses, we identified a novel direct ATF4 target gene, family with sequence similarity 129 member A (FAM129A), which is critical in mediating ATF4 effects on prostate tumorigenesis. Interestingly, FAM129A regulated both PERK and eIF2α in a feedback loop that differentially channeled the UPR output. ATF4 and FAM129A protein expression is increased in patient PCa samples compared with benign prostate. Importantly, in vivo therapeutic silencing of ATF4-FAM129A axis profoundly inhibited tumor growth in a preclinical PCa model. These data support that one of the canonical UPR branches, through ATF4 and its target gene FAM129A, is required for PCa growth and thus may serve as a novel therapeutic target.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Supplementary information is available at Oncogene’s website. The microarray data discussed in this publication have been deposited in NCBI’s Gene Expression Omnibus [36] and are accessible through GEO Series accession number GSE125826. The mass spectrometry proteomics data discussed in this publication have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the PRIDE partner repository (ID: PXD012858) [37].
References
Clarke HJ, Chambers JE, Liniker E, Marciniak SJ. Endoplasmic reticulum stress in malignancy. Cancer Cell. 2014;25:563–73.
Wang M, Kaufman RJ. The impact of the endoplasmic reticulum protein-folding environment on cancer development. Nat Rev Cancer. 2014;14:581–97.
Chevet E, Hetz C, Samali A. Endoplasmic reticulum stress-activated cell reprogramming in oncogenesis. Cancer Discov. 2015;5:586–97.
Ameri K, Harris AL. Activating transcription factor 4. Int J Biochem Cell Biol. 2008;40:14–21.
Wortel IMN, van der Meer LT, Kilberg MS, van Leeuwen FN. Surviving stress: modulation of ATF4-mediated stress responses in normal and malignant cells. Trends Endocrinol Metab. 2017;28:794–806.
Han J, Back SH, Hur J, Lin YH, Gildersleeve R, Shan J, et al. ER-stress-induced transcriptional regulation increases protein synthesis leading to cell death. Nat Cell Biol. 2013;15:481–90.
Attard G, Parker C, Eeles RA, Schröder F, Tomlins SA, Tannock I, et al. Prostate cancer. Lancet. 2016;387:70–82.
Storm M, Sheng X, Arnoldussen YJ, Saatcioglu F. Prostate cancer and the unfolded protein response. Oncotarget. 2016;7:54051–66.
Sheng X, Arnoldussen YJ, Storm M, Tesikova M, Nenseth HZ, Zhao S, et al. Divergent androgen regulation of unfolded protein response pathways drives prostate cancer. EMBO Mol Med. 2015;7:788–801.
Ye J, Kumanova M, Hart LS, Sloane K, Zhang H, De Panis DN, et al. The GCN2-ATF4 pathway is critical for tumour cell survival and proliferation in response to nutrient deprivation. EMBO J. 2010;29:2082–96.
Jin Y, Qu S, Tesikova M, Wang L, Kristian A, Maelandsmo GM, et al. Molecular circuit involving KLK4 integrates androgen and mTOR signaling in prostate cancer. Proc Natl Acad Sci USA. 2013;110:E2572–2581.
Mills IG. Maintaining and reprogramming genomic androgen receptor activity in prostate cancer. Nat Rev Cancer. 2014;14:187–98.
Shan J, Zhang F, Sharkey J, Tang TA, Ord T, Kilberg MS. The C/ebp-Atf response element (CARE) location reveals two distinct Atf4-dependent, elongation-mediated mechanisms for transcriptional induction of aminoacyl-tRNA synthetase genes in response to amino acid limitation. Nucleic Acids Res. 2016;44:9719–32.
Kilberg MS, Balasubramanian M, Fu L, Shan J. The transcription factor network associated with the amino acid response in mammalian cells. Adv Nutr. 2012;3:295–306.
Yu K, Mo D, Wu M, Chen H, Chen L, Li M, et al. Activating transcription factor 4 regulates adipocyte differentiation via altering the coordinate expression of CCATT/enhancer binding protein beta and peroxisome proliferator-activated receptor gamma. FEBS J. 2014;281:2399–409.
Cohen DM, Won KJ, Nguyen N, Lazar MA, Chen CS, Steger DJ. ATF4 licenses C/EBPbeta activity in human mesenchymal stem cells primed for adipogenesis. Elife. 2015;4:e06821.
Sun GD, Kobayashi T, Abe M, Tada N, Adachi H, Shiota A, et al. The endoplasmic reticulum stress-inducible protein Niban regulates eIF2alpha and S6K1/4E-BP1 phosphorylation. Biochem Biophys Res Commun. 2007;360:181–7.
Shaw GL, Whitaker H, Corcoran M, Dunning MJ, Luxton H, Kay J, et al. The early effects of rapid androgen deprivation on human prostate cancer. Eur Urol. 2016;70:214–8.
Massie CE, Lynch A, Ramos-Montoya A, Boren J, Stark R, Fazli L, et al. The androgen receptor fuels prostate cancer by regulating central metabolism and biosynthesis. EMBO J. 2011;30:2719–33.
Thomas BC, Kay JD, Menon S, Vowler SL, Dawson SN, Bucklow LJ, et al. Whole blood mRNA in prostate cancer reveals a four-gene androgen regulated panel. Endocr Relat Cancer. 2016;23:797–812.
Halterman MW, De Jesus C, Rempe DA, Schor NF, Federoff HJ. Loss of c/EBP-beta activity promotes the adaptive to apoptotic switch in hypoxic cortical neurons. Mol Cell Neurosci. 2008;38:125–37.
Ji H, Ding Z, Hawke D, Xing D, Jiang BH, Mills GB, et al. AKT-dependent phosphorylation of Niban regulates nucleophosmin- and MDM2-mediated p53 stability and cell apoptosis. EMBO Rep. 2012;13:554–60.
Adachi H, Majima S, Kon S, Kobayashi T, Kajino K, Mitani H, et al. Niban gene is commonly expressed in the renal tumors: a new candidate marker for renal carcinogenesis. Oncogene. 2004;23:3495–3500.
Ito S, Fujii H, Matsumoto T, Abe M, Ikeda K, Hino O. Frequent expression of Niban in head and neck squamous cell carcinoma and squamous dysplasia. Head Neck. 2010;32:96–103.
Kannangai R, Diehl AM, Sicklick J, Rojkind M, Thomas D, Torbenson M. Hepatic angiomyolipoma and hepatic stellate cells share a similar gene expression profile. Hum Pathol. 2005;36:341–7.
Pakos-Zebrucka K, Koryga I, Mnich K, Ljujic M, Samali A, Gorman AM. The integrated stress response. EMBO Rep. 2016;17:1374–95.
Harding HP, Zhang Y, Zeng H, Novoa I, Lu PD, Calfon M, et al. An integrated stress response regulates amino acid metabolism and resistance to oxidative stress. Mol Cell. 2003;11:619–33.
Yuki R, Aoyama K, Kubota S, Yamaguchi N, Kubota S, Hasegawa H, et al. Overexpression of zinc-finger protein 777 (ZNF777) inhibits proliferation at low cell density through down-regulation of FAM129A. J Cell Biochem. 2015;116:954–68.
Evstafieva AG, Kovaleva IE, Shoshinova MS, Budanov AV, Chumakov PM. Implication of KRT16, FAM129A and HKDC1 genes as ATF4 regulated components of the integrated stress response. PLoS ONE. 2018;13:e0191107.
Quirós PM, Prado MA, Zamboni N, D’Amico D, Williams RW, Finley D, et al. Multi-omics analysis identifies ATF4 as a key regulator of the mitochondrial stress response in mammals. J Cell Biol. 2017;216:2027–45.
Marshall CJ. Specificity of receptor tyrosine kinase signaling: transient versus sustained extracellular signal-regulated kinase activation. Cell. 1995;80:179–85.
Sheng X, Nenseth HZ, Qu S, Kuzu OF, Frahnow T, Simon L, et al. IRE1α-XBP1s pathway promotes prostate cancer by activating c-MYC signaling. Nat Commun. 2019;10:323.
Brijwani N, Jain M, Dhandapani M, Zahed F, Mukhopadhyay P, Biswas M, et al. Rationally co-targeting divergent pathways in KRAS wild-type colorectal cancers by CANscript technology reveals tumor dependence on Notch and Erbb2. Sci Rep. 2017;7:1502.
Nevedomskaya E, Baumgart SJ, Haendler B. Recent advances in prostate cancer treatment and drug discovery. Int J Mol Sci. 2018;19, 1359–84.
Jin Y, Wang L, Qu S, Sheng X, Kristian A, Maelandsmo GM, et al. STAMP2 increases oxidative stress and is critical for prostate cancer. EMBO Mol Med. 2015;7:315–31.
Edgar R, Domrachev M, Lash AE. Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002;30:207–10.
Vizcaíno JA, Côté RG, Csordas A, Dianes JA, Fabregat A, Foster JM, et al. The PRoteomics IDEntifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res. 2013;41:D1063–1069.
Acknowledgements
We would like to thank Veronica F. Blihovde, Xia Sheng and Deane Stryker for experimental help and members of the FS laboratory for helpful discussions. We would like to thank Jeffrey Petro for excellent technical support and Drs Eva Corey and Colm Morrissey of University of Washington for providing LuCaP PDX lines. This work was supported by Norwegian Research Council grant 193337, Norwegian Cancer Society grant 419204, and Health South East Norway grant 36024. The microarray gene expression service was provided by the Genomics Core Facility (GCF), Norwegian University of Science and Technology (NTNU). GCF is funded by the Faculty of Medicine at NTNU and Central Norway Regional Health Authority. The LC-MS analyses were performed by the proteomics core facility at the Department of Biosciences, University of Oslo.
Author information
Authors and Affiliations
Contributions
NP, ML, and FS designed the study and experiments. NP and ML performed and analyzed most of the experiments. MT conducted ChIP experiments. HZN performed IP-western experiments. EA and JS performed colocalization studies. YiJ conducted nanoliposomal siRNA delivery experiments. DK contributed to growth assays. PB and AU conducted PDX-derived organoid experiments. LF and PR provided tissue microarrays of PCa, performed IHC analyses, and their interpretation. BO, NK, and HMM provided nanoliposomal siRNAs. YaJ performed IHC analyses on consecutive slides and helped with supervision. OFK did statistical analysis of IHC data. MP scored TMAs. HD provided expertize in IHC interpretation. FS supervised the project. NP, ML, and FS wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Pällmann, N., Livgård, M., Tesikova, M. et al. Regulation of the unfolded protein response through ATF4 and FAM129A in prostate cancer. Oncogene 38, 6301–6318 (2019). https://doi.org/10.1038/s41388-019-0879-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0879-2
This article is cited by
-
THEM6 is a prognostic biomarker for breast cancer and is associated with immune infiltration
Scientific Reports (2023)
-
The endoplasmic reticulum stress response in prostate cancer
Nature Reviews Urology (2022)
-
Dynamic alteration in miRNA and mRNA expression profiles at different stages of chronic arsenic exposure-induced carcinogenesis in a human cell culture model of skin cancer
Archives of Toxicology (2021)
-
DENR promotes translation reinitiation via ribosome recycling to drive expression of oncogenes including ATF4
Nature Communications (2020)
-
Alterations in niban gene expression as a response to stress conditions in 3T3-L1 adipocytes
Molecular Biology Reports (2020)