Abstract
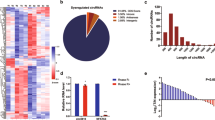
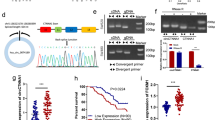
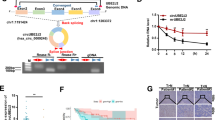
CircRNAs have been reported to exert momentous roles in regulating pathophysiological process and guiding clinical diagnosis and treatment in colorectal cancer (CRC). However, there are still a lot of circRNAs that need to be unearthed. In this study, we evaluated the expression profile of circRNAs in 10 CRC tissues and their corresponding normal-appearing tissues (NATs) by microarray, and identified that hsa_circ_101555 (circ101555) was significantly up-regulated in tumor tissues and closely related to the prognosis of CRC patients. A specific close loop structure of circ101555 was described, which was generated by back-splicing of the host gene CSNK1G1 and showed greater stability than the linear RNA. The results in vitro and in vivo showed that silencing circ101555 expression significantly suppressed cell proliferation, induced apoptosis and impaired the DNA repair capacity of CRC cells, while rescue experiments suggested that down-expression of miR-597-5p could significantly attenuate the biological effects of circ101555 knockdown on CRC cells. Subsequent experiments in vitro, including double fluorescence in situ hybridization (D-FISH) analysis, RIP analysis and biotin-coupled probe pull down assay, confirmed that miR-597-5p was effectively enriched by circ101555, and circ101555 might serve as a sponge of miR-597-5p. Moreover, two putative oncogenes (CDK6 and RPA3) were identified as the miR-597-5p potential targets. Taken together, our results proved that circ101555 might function as a competing endogenous RNA of miR-597-5p to up-regulate CDK6 and RPA3 expression in CRC. Circ101555 could be a useful prognostic indicator in patients with CRC, and silence of circ101555 provided a new attractive therapeutic measure for CRC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA. 2016;66:7–30.
Wilkins T, McMechan D, Talukder A. Colorectal cancer screening and prevention. Am Fam physician. 2018;97:658–65.
Hawk ET, Levin B. Colorectal cancer prevention. J Clin Oncol. 2005;23:378–91.
Pattison AM, Merlino DJ, Blomain ES, Waldman SA. Guanylyl cyclase C signaling axis and colon cancer prevention. World J Gastroenterol. 2016;22:8070–7.
Haddad FG, Eid R, Kourie HR, Barouky E, Ghosn M. Prognostic and predictive biomarkers in nonmetastatic colorectal cancers. Future Oncol. 2018;14:2097–102.
Chen LL. The biogenesis and emerging roles of circular RNAs. Nat Rev Mol Cell Biol. 2016;17:205–11.
Holdt LM, Kohlmaier A, Teupser D. Molecular roles and function of circular RNAs in eukaryotic cells. Cell Mol life Sci. 2018;75:1071–98.
Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12:381–8.
Cocquerelle C, Mascrez B, Hetuin D, Bailleul B. Mis-splicing yields circular RNA molecules. FASEB J. 1993;7:155–60.
Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016;44:2846–58.
Du WW, Yang W, Chen Y, Wu ZK, Foster FS, Yang Z, et al. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur Heart J. 2017;38:1402–12.
Hansen TB, Kjems J, Damgaard CK. Circular RNA and miR-7 in cancer. Cancer Res. 2013;73:5609–12.
Dragomir M, Calin GA. Circular RNAs in cancer - lessons learned from microRNAs. Front Oncol. 2018;8:179.
Zhou R, Wu Y, Wang W, Su W, Liu Y, Wang Y, et al. Circular RNAs (circRNAs) in cancer. Cancer Lett. 2018;425:134–42.
Yang Q, Du WW, Wu N, Yang W, Awan FM, Fang L, et al. A circular RNA promotes tumorigenesis by inducing c-myc nuclear translocation. Cell Death Differ. 2017;24:1609–20.
Yao Z, Luo J, Hu K, Lin J, Huang H, Wang Q, et al. ZKSCAN1 gene and its related circular RNA (circZKSCAN1) both inhibit hepatocellular carcinoma cell growth, migration, and invasion but through different signaling pathways. Mol Oncol. 2017;11:422–37.
Sun S, Li B, Wang Y, Li X, Wang P, Wang F, et al. Clinical significance of the decreased expression of hsa_circ_001242 in oral squamous cell carcinoma. Dis Markers. 2018;2018:6514795.
Li T, Shao Y, Fu L, Xie Y, Zhu L, Sun W, et al. Plasma circular RNA profiling of patients with gastric cancer and their droplet digital RT-PCR detection. J Mol Med. 2018;96:85–96.
Zeng Y, Xu Y, Shu R, Sun L, Tian Y, Shi C, et al. Altered expression profiles of circular RNA in colorectal cancer tissues from patients with lung metastasis. Int J Mol Med. 2017;40:1818–28.
Chen S, Zhang L, Su Y, Zhang X. Screening potential biomarkers for colorectal cancer based on circular RNA chips. Oncol Rep. 2018;39:2499–512.
Jiang W, Zhang X, Chu Q, Lu S, Zhou L, Lu X, et al. The circular RNA profiles of colorectal tumor metastatic cells. Front Genet. 2018;9:34.
Yuan Y, Liu W, Zhang Y, Zhang Y, Sun S. CircRNA circ_0026344 as a prognostic biomarker suppresses colorectal cancer progression via microRNA-21 and microRNA-31. Biochem Biophys Res Commun. 2018;503:870–5.
Esteller M. Non-coding RNAs in human disease. Nat Rev Genet. 2011;12:861–74.
Qu S, Yang X, Li X, Wang J, Gao Y, Shang R, et al. Circular RNA: a new star of noncoding RNAs. Cancer Lett. 2015;365:141–8.
Wu J, Jiang Z, Chen C, Hu Q, Fu Z, Chen J, et al. CircIRAK3 sponges miR-3607 to facilitate breast cancer metastasis. Cancer Lett. 2018;430:179–92.
Yang X, Yuan W, Tao J, Li P, Yang C, Deng X, et al. Identification of circular RNA signature in bladder cancer. J Cancer. 2017;8:3456–63.
Chen G, Shi Y, Zhang Y, Sun J. CircRNA_100782 regulates pancreatic carcinoma proliferation through the IL6-STAT3 pathway. OncoTargets Ther. 2017;10:5783–94.
Xu N, Chen S, Liu Y, Li W, Liu Z, Bian X, et al. Profiles and bioinformatics analysis of differentially expressed circrnas in taxol-resistant non-small cell lung cancer cells. Cell Physiol Biochem. 2018;48:2046–60.
Wang X, Zhang Y, Huang L, Zhang J, Pan F, Li B, et al. Decreased expression of hsa_circ_001988 in colorectal cancer and its clinical significances. Int J Clin Exp Pathol. 2015;8:16020–5.
Zhu M, Xu Y, Chen Y, Yan F. Circular BANP, an upregulated circular RNA that modulates cell proliferation in colorectal cancer. Biomed Pharm. 2017;88:138–44.
Guo JN, Li J, Zhu CL, Feng WT, Shao JX, Wan L, et al. Comprehensive profile of differentially expressed circular RNAs reveals that hsa_circ_0000069 is upregulated and promotes cell proliferation migration, and invasion in colorectal cancer. OncoTargets Ther. 2016;9:7451–8.
Hsiao KY, Lin YC, Gupta SK, Chang N, Yen L, Sun HS, et al. Noncoding effects of circular RNA CCDC66 promote colon cancer growth and metastasis. Cancer Res. 2017;77:2339–50.
Zhang R, Xu J, Zhao J, Wang X. Silencing of hsa_circ_0007534 suppresses proliferation and induces apoptosis in colorectal cancer cells. Eur Rev Med Pharmacol Sci. 2018;22:118–26.
Zeng K, Chen X, Xu M, Liu X, Hu X, Xu T, et al. CircHIPK3 promotes colorectal cancer growth and metastasis by sponging miR-7. Cell Death Dis. 2018;9:417.
Xiong DD, Dang YW, Lin P, Wen DY, He RQ, Luo DZ, et al. A circRNA-miRNA-mRNA network identification for exploring underlying pathogenesis and therapy strategy of hepatocellular carcinoma. J Transl Med. 2018;16:220.
Zhong Y, Du Y, Yang X, Mo Y, Fan C, Xiong F, et al. Circular RNAs function as ceRNAs to regulate and control human cancer progression. Mol Cancer. 2018;17:79.
Chen L, Zhang S, Wu J, Cui J, Zhong L, Zeng L, et al. circRNA_100290 plays a role in oral cancer by functioning as a sponge of the miR-29 family. Oncogene. 2017;36:4551–61.
Zhang X, Luo P, Jing W, Zhou H, Liang C, Tu J. circSMAD2 inhibits the epithelial-mesenchymal transition by targeting miR-629 in hepatocellular carcinoma. OncoTargets Ther. 2018;11:2853–63.
Zhang XL, Xu LL, Wang F. Hsa_circ_0020397 regulates colorectal cancer cell viability, apoptosis and invasion by promoting the expression of the miR-138 targets TERT and PD-L1. Cell Biol Int. 2017;41:1056–64.
Pek M, Yatim S, Chen Y, Li J, Gong M, Jiang X, et al. Oncogenic KRAS-associated gene signature defines co-targeting of CDK4/6 and MEK as a viable therapeutic strategy in colorectal cancer. Oncogene. 2017;36:4975–86.
Sobecki M, Mrouj K, Colinge J, Gerbe F, Jay P, Krasinska L, et al. Cell-cycle regulation accounts for variability in Ki-67 expression levels. Cancer Res. 2017;77:2722–34.
Dahai Y, Sanyuan S, Hong L, Di Z, Chong Z. A relationship between replication protein A and occurrence and prognosis of esophageal carcinoma. Cell Biochem Biophys. 2013;67:175–80.
Chen CC, Juan CW, Chen KY, Chang YC, Lee JC, Chang MC. Upregulation of RPA2 promotes NF-kappaB activation in breast cancer by relieving the antagonistic function of menin on NF-kappaB-regulated transcription. Carcinogenesis. 2017;38:196–206.
Kanakis D, Levidou G, Gakiopoulou H, Eftichiadis C, Thymara I, Fragkou P, et al. Replication protein A: a reliable biologic marker of prognostic and therapeutic value in human astrocytic tumors. Hum Pathol. 2011;42:1545–53.
Xiao W, Zheng J, Zhou B, Pan L. Replication protein A 3 is associated with hepatocellular carcinoma tumorigenesis and poor patient survival. Dig Dis. 2018;36:26–32.
Givalos N, Gakiopoulou H, Skliri M, Bousboukea K, Konstantinidou AE, Korkolopoulou P, et al. Replication protein A is an independent prognostic indicator with potential therapeutic implications in colon cancer. Mod Pathol. 2007;20:159–66.
Acknowledgements
This study was supported by grants from the Special Subject of Diagnosis Treatment of Key Clinical Diseases of Suzhou City Sci-tech Bureau (LCZX201401), the Project of Invigorating Health Care through Science, Technology and Education, Jiangsu Provincial Medical Youth Talent (QNRC2016723) and Suzhou Gusu Medical Youth Talent (GSWS2019032).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, Z., Ren, R., Wan, D. et al. Hsa_circ_101555 functions as a competing endogenous RNA of miR-597-5p to promote colorectal cancer progression. Oncogene 38, 6017–6034 (2019). https://doi.org/10.1038/s41388-019-0857-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0857-8
This article is cited by
-
Cirscan: a shiny application to identify differentially active sponge mechanisms and visualize circRNA–miRNA–mRNA networks
BMC Bioinformatics (2024)
-
Identification of expression profiles and prognostic value of RFCs in colorectal cancer
Scientific Reports (2024)
-
Integrated whole transcriptome profiling revealed a convoluted circular RNA-based competing endogenous RNAs regulatory network in colorectal cancer
Scientific Reports (2024)
-
Circ_0005615 Regulates the Progression of Colorectal Cancer Through the miR-873-5p/FOSL2 Signaling Pathway
Biochemical Genetics (2023)
-
Circ_0000395 Promoted CRC Progression via Elevating MYH9 Expression by Sequestering miR-432-5p
Biochemical Genetics (2023)