Abstract
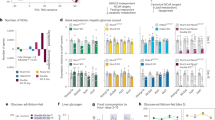
Tumor cells undergo a metabolic shift in order to adapt to the altered microenvironment, although the underlying mechanisms have not been fully explored. HnRNP A1 is involved in the alternative splicing of the pyruvate kinase (PK) mRNA, allowing tumor cells to specifically produce the PKM2 isoform. We found that the acetylation status of hnRNP A1 in hepatocellular carcinoma (HCC) cells was dependent on glucose availability, which affected the PKM2-dependent glycolytic pathway. In the glucose-starved HCC cells, SIRT1 and SIRT6, members of deacetylase sirtuin family, were highly expressed and deacetylated hnRNP A1 after direct binding. We identified four lysine residues in hnRNP A1 that were deacetylated by SIRT1 and SIRT6, resulting in significant inhibition of glycolysis in HCC cells. Deacetylated hnRNP A1 reduced PKM2 and increased PKM1 alternative splicing in HCC cells under normal glucose conditions, thereby reducing the metabolic activity of PK and the non-metabolic PKM2–β-catenin signaling pathway. However, under glucose starvation, the low levels of acetylated hnRNP A1 reduced HCC cell metabolism to adapt to the nutrient deficiency. Taken together, sirtuin-mediated hnRNP A1 deacetylation inhibits HCC cell proliferation and tumorigenesis in a PKM2-dependent manner. These findings point to the metabolic reprogramming induced by hnRNP A1 acetylation in order to adapt to the nutritional status of the tumor microenvironment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Iansante V, Choy PM, Fung SW, Liu Y, Chai JG, Dyson J, et al. PARP14 promotes the Warburg effect in hepatocellular carcinoma by inhibiting JNK1-dependent PKM2 phosphorylation and activation. Nat Commun. 2015;6:7882.
Mazurek S. Pyruvate kinase type M2: a key regulator of the metabolic budget system in tumor cells. Int J Biochem Cell Biol. 2011;43:969–80.
Zhang LF, Lou JT, Lu MH, Gao C, Zhao S, Li B, et al. Suppression of miR-199a maturation by HuR is crucial for hypoxia-induced glycolytic switch in hepatocellular carcinoma. EMBO J. 2015;34:2671–85.
Williams AL, Khadka V, Tang M, Avelar A, Schunke KJ, Menor M, et al. HIF1 mediates a switch in pyruvate kinase isoforms after myocardial infarction. Physiol Genomics. 2018;50:479–94.
Chen H, Hewison M, Hu B, Adams JS. Heterogeneous nuclear ribonucleoprotein (hnRNP) binding to hormone response elements: a cause of vitamin D resistance. Proc Natl Acad Sci USA. 2003;100:6109–14.
Chettouh H, Fartoux L, Aoudjehane L, Wendum D, Claperon A, Chretien Y, et al. Mitogenic insulin receptor-A is overexpressed in human hepatocellular carcinoma due to EGFR-mediated dysregulation of RNA splicing factors. Cancer Res. 2013;73:3974–86.
Park WC, Kim HR, Kang DB, Ryu JS, Choi KH, Lee GO, et al. Comparative expression patterns and diagnostic efficacies of SR splicing factors and HNRNPA1 in gastric and colorectal cancer. BMC Cancer. 2016;16:358.
Kim YJ, Kim BR, Ryu JS, Lee GO, Kim HR, Choi KH, et al. HNRNPA1, a splicing regulator, is an effective target protein for cervical cancer detection: comparison with conventional tumor markers. Int J Gynecol Cancer. 2017;27:326–31.
Roy R, Huang Y, Seckl MJ, Pardo OE. Emerging roles of hnRNPA1 in modulating malignant transformation. Wiley Interdiscip Rev RNA. 2017;8:e1431.
Chen M, Zhang J, Manley JL. Turning on a fuel switch of cancer: hnRNP proteins regulate alternative splicing of pyruvate kinase mRNA. Cancer Res. 2010;70:8977–80.
Huang JZ, Chen M, Chen, Gao XC, Zhu S, Huang H, et al. A peptide encoded by a putative lncRNA HOXB-AS3 suppresses colon cancer growth. Mol Cell. 2017;68:171–84.
Yang W, Zheng Y, Xia Y, Ji H, Chen X, Guo F, et al. ERK1/2-dependent phosphorylation and nuclear translocation of PKM2 promotes the Warburg effect. Nat Cell Biol. 2012;14:1295–304.
Yang W, Xia Y, Ji H, Zheng Y, Liang J, Huang W, et al. Nuclear PKM2 regulates beta-catenin transactivation upon EGFR activation. Nature. 2011;480:118–22.
Dayton TL, Gocheva V, Miller KM, Bhutkar A, Lewis CA, Bronson RT, et al. Isoform-specific deletion of PKM2 constrains tumor initiation in a mouse model of soft tissue sarcoma. Cancer Metab. 2018;6:6.
Xu Q, Liu LZ, Yin Y, He J, Li Q, Qian X, et al. Regulatory circuit of PKM2/NF-kappaB/miR-148a/152-modulated tumor angiogenesis and cancer progression. Oncogene. 2015;34:5482–93.
Chen J, Chan AW, To KF, Chen W, Zhang Z, Ren J, et al. SIRT2 overexpression in hepatocellular carcinoma mediates epithelial to mesenchymal transition by protein kinase B/glycogen synthase kinase-3beta/beta-catenin signaling. Hepatology. 2013;57:2287–98.
Zhang ZY, Hong D, Nam SH, Kim JM, Paik YH, Joh JW, et al. SIRT1 regulates oncogenesis via a mutant p53-dependent pathway in hepatocellular carcinoma. J Hepatol. 2015;62:121–30.
Ong ALC, Ramasamy TS. Role of Sirtuin1-p53 regulatory axis in aging, cancer and cellular reprogramming. Ageing Res Rev. 2018;43:64–80.
Agerholm M, Dall M, Jensen BAH, Prats C, Madsen S, Basse AL, et al. Perturbations of NAD(+) salvage systems impact mitochondrial function and energy homeostasis in mouse myoblasts and intact skeletal muscle. Am J Physiol Endocrinol Metab. 2018;314:E377–e95.
Park SH, Ozden O, Liu G, Song HY, Zhu Y, Yan Y, et al. SIRT2-mediated deacetylation and tetramerization of pyruvate kinase directs glycolysis and tumor growth. Cancer Res. 2016;76:3802–12.
Bhardwaj A, Das S. SIRT6 deacetylates PKM2 to suppress its nuclear localization and oncogenic functions. Proc Natl Acad Sci USA. 2016;113:E538–47.
Zhu Y, Yan Y, Principe DR, Zou X, Vassilopoulos A. Gius D. SIRT3 and SIRT4 are mitochondrial tumor suppressor proteins that connect mitochondrial metabolism and carcinogenesis. Cancer Metab. 2014;2:15.
Chang C, Hua S, Zhang D, Wang Y, Shen Q, Bo L, et al. AMPK-dependent phosphorylation of GAPDH triggers Sirt1 activation and is necessary for autophagy upon glucose starvation. Mol Cell. 2015;60:930–40.
Parenti MD, Grozio A, Bauer I, Galeno L, Damonte P, Millo E, et al. Discovery of novel and selective SIRT6 inhibitors. J Med Chem. 2014;57:4796–804.
Clower CV, Chatterjee D, Wang Z, Cantley LC, Vander Heiden MG, Krainer AR. The alternative splicing repressors hnRNP A1/A2 and PTB influence pyruvate kinase isoform expression and cell metabolism. Proc Natl Acad Sci USA. 2010;107:1894–9.
Sun Y, Luo M, Chang G, Ren W, Wu K, Li X, et al. Phosphorylation of Ser6 in hnRNPA1 by S6K2 regulates glucose metabolism and cell growth in colorectal cancer. Oncol Lett. 2017;14:7323–31.
Luan W, Wang Y, Chen X, Shi Y, Wang J, Zhang J, et al. PKM2 promotes glucose metabolism and cell growth in gliomas through a mechanism involving a let-7a/c-Myc/hnRNPA1 feedback loop. Oncotarget. 2015;6:13006–18.
Gu Z, Xia J, Xu H, Frech I, Tricot G, Zhan F. NEK2 promotes aerobic glycolysis in multiple myeloma through regulating splicing of pyruvate kinase. J Hematol Oncol. 2017;10:17.
David CJ, Chen M, Assanah M, Canoll P, Manley JL. HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature. 2010;463:364–8.
Ning X, Qi H, Li R, Li Y, Jin Y, McNutt MA, et al. Discovery of novel naphthoquinone derivatives as inhibitors of the tumor cell specific M2 isoform of pyruvate kinase. Eur J Med Chem. 2017;138:343–52.
Mohagheghi F, Prudencio M, Stuani C, Cook C, Jansen-West K, Dickson DW, et al. TDP-43 functions within a network of hnRNP proteins to inhibit the production of a truncated human SORT1 receptor. Hum Mol Genet. 2016;25:534–45.
Roda D, Castillo J, Telechea-Fernandez M, Gil A, Lopez-Rodas G, Franco L. et al. EGF-induced acetylation of heterogeneous nuclear ribonucleoproteins is dependent on KRAS mutational status in colorectal cancer cells. PLoS ONE 2015;10:e0130543
Xuan Y, Wang J, Ban L, Lu JJ, Yi C, Li Z, et al. hnRNPA2/B1 activates cyclooxygenase-2 and promotes tumor growth in human lung cancers. Mol Oncol. 2016;10:610–24.
Pina JM, Reynaga JM, Truong AAM, Keppetipola NM. Post-translational modifications in polypyrimidine tract binding proteins PTBP1 and PTBP2. Biochemistry. 2018;57:3873–82.
Pun KK, Young RT, Wang C, Tam CF, Ho PW. The use of glucagon challenge tests in the diagnostic evaluation of hypoglycemia due to hepatoma and insulinoma. J Clin Endocrinol Metab. 1988;67:546–50.
Li H, Xu M, Lee J, He C, Xie Z. Leucine supplementation increases SIRT1 expression and prevents mitochondrial dysfunction and metabolic disorders in high-fat diet-induced obese mice. Am J Physiol Endocrinol Metab. 2012;303:E1234–44.
Li L, Yoshitomi H, Wei Y, Qin L, Zhou J, Xu T. et al. Tang-Nai-Kang alleviates pre-diabetes and metabolic disorders and induces a gene expression switch toward fatty acid oxidation in SHR.Cg-Leprcp/NDmcr rats. PLoS ONE 2015;10:e0122024
Ampferl R, Rodemann HP, Mayer C, Hofling TTA, Dittmann K. Glucose starvation impairs DNA repair in tumour cells selectively by blocking histone acetylation. Radiother Oncol. 2018;126:465–70.
Fang X, Lu G, Ha K, Lin H, Du Y, Zuo Q, et al. Acetylation of TIP60 at K104 is essential for metabolic stress-induced apoptosis in cells of hepatocellular cancer. Exp Cell Res. 2018;362:279–86.
Lv L, Li D, Zhao D, Lin R, Chu Y, Zhang H, et al. Acetylation targets the M2 isoform of pyruvate kinase for degradation through chaperone-mediated autophagy and promotes tumor growth. Mol Cell. 2011;42:719–30.
Xiong Y, Lei QY, Zhao S, Guan KL. Regulation of glycolysis and gluconeogenesis by acetylation of PKM and PEPCK. Cold Spring Harb Symp Quant Biol. 2011;76:285–9.
Chen Y, Huang Q, Liu W, Zhu Q, Cui CP, Xu L, et al. Mutually exclusive acetylation and ubiquitylation of the splicing factor SRSF5 control tumor growth. Nat Commun. 2018;9:2464.
Lee JY, Kapur M, Li M, Choi MC, Choi S, Kim HJ, et al. MFN1 deacetylation activates adaptive mitochondrial fusion and protects metabolically challenged mitochondria. J Cell Sci. 2014;127:4954–63.
Kashima T, Rao N, David CJ, Manley JL. hnRNP A1 functions with specificity in repression of SMN2 exon 7 splicing. Hum Mol Genet. 2007;16:3149–59.
Acknowledgements
This work was supported by research grants from the National Natural Science Foundation of China (Nos. 81530053, 81830052, 81771858, 81602415, 81670573, 81572719, 81874202, and 81803581) and Shanghai Municipal Education Commission (Class II Plateau Disciplinary Construction Program for Medical Technology of SUMHS, 2018–2020).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Yang, H., Zhu, R., Zhao, X. et al. Sirtuin-mediated deacetylation of hnRNP A1 suppresses glycolysis and growth in hepatocellular carcinoma. Oncogene 38, 4915–4931 (2019). https://doi.org/10.1038/s41388-019-0764-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0764-z
This article is cited by
-
hnRNP A1 dysfunction alters RNA splicing and drives neurodegeneration in multiple sclerosis (MS)
Nature Communications (2024)
-
The Role of PKM2 in Multiple Signaling Pathways Related to Neurological Diseases
Molecular Neurobiology (2023)
-
Decoding the role of aberrant RNA alternative splicing in hepatocellular carcinoma: a comprehensive review
Journal of Cancer Research and Clinical Oncology (2023)
-
A comprehensive understanding of hnRNP A1 role in cancer: new perspectives on binding with noncoding RNA
Cancer Gene Therapy (2022)
-
The sirtuin family in health and disease
Signal Transduction and Targeted Therapy (2022)