Abstract
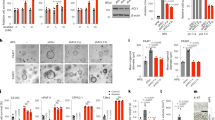
In an effort to understand the underlying mechanisms of lymph node metastasis in oral squamous cell carcinoma (OSCC), through in vivo selection, LN1-1 cells were previously established from OEC-M1 cells and showed enhanced lymphangiogenesis and lymphatic metastasis capabilities. In the current study, we use a stable isotope labeling with amino acids in cell culture (SILAC) and liquid chromatography-tandem mass spectrometry (LC-MS/MS)-based proteomic platform to compare LN1-1 to OEC-M1 cells. Interferon-stimulated gene 15 (ISG15) was found highly expressed in LN1-1 cells. Immunohistochemical analysis and meta-analysis of publicly available microarray datasets revealed that the ISG15 level was increased in human OSCC tissues and associated with poor disease outcome. Knockdown of ISG15 had minimal effects on tumor growth but did decrease tumor lymphangiogenesis and lymphatic metastasis of LN1-1 cells. Consistent with the in vivo assay, ISG15 knockdown did not impair cell growth but diminished cell migration, invasion, and transendothelial migration in vitro. ISG15-induced cell migration was independent of ISGylation and associated with membrane protrusion. Ectopic expression of ISG15 increased Rac1 activity and knockdown of Rac1 impaired ISG15-enhanced migration. Furthermore, Rac1 colocalized with ISG15 to a region of membrane protrusion and ISG15 coimmunoprecipitated with Rac1, especially with the Rac1-GDP form. Importantly, as shown by proximity ligation assays, ISG15 and Rac1 physically interacted with each other. Our results indicated that ISG15 affects cell migration by interacting with Rac1 and regulating Rac1 activity and contributes to lymphatic metastasis in OSCC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Liao CT, Chang JT, Wang HM, Ng SH, Hsueh C, Lee LY, et al. Pretreatment primary tumor SUVmax measured by FDG-PET and pathologic tumor depth predict for poor outcomes in patients with oral cavity squamous cell carcinoma and pathologically positive lymph nodes. Int J Radiat Oncol Biol Phys. 2009;73:764–71.
Ahmad Y, Lamond AI. A perspective on proteomics in cell biology. Trends Cell Biol. 2014;24:257–64.
Walther TC, Mann M. Mass spectrometry-based proteomics in cell biology. J Cell Biol. 2010;190:491–500.
Ong SE, Blagoev B, Kratchmarova I, Kristensen DB, Steen H, Pandey A, et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol Cell Proteom. 2002;1:376–86.
Yen YC, Hsiao JR, Jiang SS, Chang JS, Wang SH, Shen YY, et al. Insulin-like growth factor-independent insulin-like growth factor binding protein 3 promotes cell migration and lymph node metastasis of oral squamous cell carcinoma cells by requirement of integrin beta1. Oncotarget. 2015;6:41837–55.
Zhang X, Liu Y, Gilcrease MZ, Yuan XH, Clayman GL, Adler-Storthz K, et al. A lymph node metastatic mouse model reveals alterations of metastasis-related gene expression in metastatic human oral carcinoma sublines selected from a poorly metastatic parental cell line. Cancer. 2002;95:1663–72.
Reich N, Evans B, Levy D, Fahey D, Knight E Jr, Darnell JE Jr. Interferon-induced transcription of a gene encoding a 15-kDa protein depends on an upstream enhancer element. Proc Natl Acad Sci USA. 1987;84:6394–8.
Durfee LA, Huibregtse JM. The ISG15 conjugation system. Methods Mol Biol. 2012;832:141–9.
Cajee UF, Hull R, Ntwasa M. Modification by ubiquitin-like proteins: significance in apoptosis and autophagy pathways. Int J Mol Sci. 2012;13:11804–31.
Kim KI, Yan M, Malakhova O, Luo JK, Shen MF, Zou W, et al. Ube1L and protein ISGylation are not essential for alpha/beta interferon signaling. Mol Cell Biol. 2006;26:472–9.
Hishiki T, Han Q, Arimoto K, Shimotohno K, Igarashi T, Vasudevan SG, et al. Interferon-mediated ISG15 conjugation restricts dengue virus 2 replication. Biochem Biophys Res Commun. 2014;448:95–100.
Loeb KR, Haas AL. The interferon-inducible 15-kDa ubiquitin homolog conjugates to intracellular proteins. J Biol Chem. 1992;267:7806–13.
Jeon YJ, Yoo HM, Chung CH. ISG15 and immune diseases. Biochim Biophys Acta. 2010;1802:485–96.
Burks J, Reed RE, Desai SD. Free ISG15 triggers an antitumor immune response against breast cancer: a new perspective. Oncotarget. 2015;6:7221–31.
Dos Santos PF, Mansur DS. Beyond ISGlylation: functions of free intracellular and extracellular ISG15. J Interferon Cytokine Res. 2017;37:246–53.
Sgorbissa A, Brancolini C. IFNs, ISGylation and cancer: Cui prodest? Cytokine Growth Factor Rev. 2012;23:307–14.
Andersen JB, Hassel BA. The interferon regulated ubiquitin-like protein, ISG15, in tumorigenesis: friend or foe? Cytokine Growth Factor Rev. 2006;17:411–21.
Desai SD, Reed RE, Burks J, Wood LM, Pullikuth AK, Haas AL, et al. ISG15 disrupts cytoskeletal architecture and promotes motility in human breast cancer cells. Exp Biol Med. 2012;237:38–49.
Li C, Wang J, Zhang H, Zhu MG, Chen FF, Hu YF, et al. Interferon-stimulated gene 15 (ISG15) is a trigger for tumorigenesis and metastasis of hepatocellular carcinoma. Oncotarget. 2014;5:8429–41.
Spano D, Heck C, De Antonellis P, Christofori G, Zollo M. Molecular networks that regulate cancer metastasis. Semin Cancer Biol. 2012;22:234–49.
Friedl P, Locker J, Sahai E, Segall JE. Classifying collective cancer cell invasion. Nat Cell Biol. 2012;14:777–83.
Charras G, Sahai E. Physical influences of the extracellular environment on cell migration. Nat Rev Mol Cell Biol. 2014;15:813–24.
Graziano BR, Weiner OD. Self-organization of protrusions and polarity during eukaryotic chemotaxis. Curr Opin Cell Biol. 2014;30:60–7.
Boureux A, Vignal E, Faure S, Fort P. Evolution of the Rho family of ras-like GTPases in eukaryotes. Mol Biol Evol. 2007;24:203–16.
Ueda T, Kikuchi A, Ohga N, Yamamoto J, Takai Y. Purification and characterization from bovine brain cytosol of a novel regulatory protein inhibiting the dissociation of GDP from and the subsequent binding of GTP to rhoBp20, a ras p21-like GTP-binding protein. J Biol Chem. 1990;265:9373–80.
Symons M, Settleman J. Rho family GTPases: more than simple switches. Trends Cell Biol. 2000;10:415–9.
Rhodes DR, Kalyana-Sundaram S, Mahavisno V, Varambally R, Yu J, Briggs BB. et al. Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia. 2007;9:166–80.
Estilo CL, Oc P, Talbot S, Socci ND, Carlson DL, Ghossein R. et al. Oral tongue cancer gene expression profiling: identification of novel potential prognosticators by oligonucleotide microarray analysis. BMC Cancer. 2009;9:11.
Pyeon D, Newton MA, Lambert PF, den Boon JA, Sengupta S, Marsit CJ, et al. Fundamental differences in cell cycle deregulation in human papillomavirus-positive and human papillomavirus-negative head/neck and cervical cancers. Cancer Res. 2007;67:4605–19.
Talbot SG, Estilo C, Maghami E, Sarkaria IS, Pham DK, Oc P, et al. Gene expression profiling allows distinction between primary and metastatic squamous cell carcinomas in the lung. Cancer Res. 2005;65:3063–71.
Ye H, Yu T, Temam S, Ziober BL, Wang J, Schwartz JL, et al. Transcriptomic dissection of tongue squamous cell carcinoma. BMC Genom. 2008;9:69.
Toruner GA, Ulger C, Alkan M, Galante AT, Rinaggio J, Wilk R, et al. Association between gene expression profile and tumor invasion in oral squamous cell carcinoma. Cancer Genet Cytogenet. 2004;154:27–35.
Kuriakose MA, Chen WT, He ZM, Sikora AG, Zhang P, Zhang ZY, et al. Selection and validation of differentially expressed genes in head and neck cancer. Cell Mol Life Sci. 2004;61:1372–83.
Peng CH, Liao CT, Peng SC, Chen YJ, Cheng AJ, Juang JL. et al. A novel molecular signature identified by systems genetics approach predicts prognosis in oral squamous cell carcinoma. PLoS ONE. 2011;6:e23452.
Lee CH, Wong TS, Chan JY, Lu SC, Lin P, Cheng AJ, et al. Epigenetic regulation of the X-linked tumour suppressors BEX1 and LDOC1 in oral squamous cell carcinoma. J Pathol. 2013;230:298–309.
Gardel ML, Schneider IC, Aratyn-Schaus Y, Waterman CM. Mechanical integration of actin and adhesion dynamics in cell migration. Annu Rev Cell Dev Biol. 2009;26:315–33.
Laljee RP, Muddaiah S, Salagundi B, Cariappa PM, Indra AS, Sanjay V, et al. Interferon stimulated gene-ISG15 is a potential diagnostic biomarker in oral squamous cell carcinomas. Asian Pac J Cancer Prev. 2013;14:1147–50.
Zhang Q, He Y, Nie M, Cai W. Roles of miR-138 and ISG15 in oral squamous cell carcinoma. Exp Ther Med. 2017;14:2329–34.
Hermann MR, Jakobson M, Colo GP, Rognoni E, Jakobson M, Kupatt C, et al. Integrins synergise to induce expression of the MRTF-A-SRF target gene ISG15 for promoting cancer cell invasion. J Cell Sci. 2016;129:1391–403.
Sainz B, Martin B, Tatari M, Heeschen C, Guerra S. ISG15 is a critical microenvironmental factor for pancreatic cancer stem cells. Cancer Res. 2014;74:7309–20.
Zhang X, Bogunovic D, Payelle-Brogard B, Francois-Newton V, Speer SD, Yuan C, et al. Human intracellular ISG15 prevents interferon-alpha/beta over-amplification and auto-inflammation. Nature. 2015;517:89–93.
Yamaguchi H, Condeelis J. Regulation of the actin cytoskeleton in cancer cell migration and invasion. Biochim Biophys Acta. 2007;1773:642–52.
Um K, Niu S, Duman JG, Cheng JX, Tu YK, Schwechter B, et al. Dynamic control of excitatory synapse development by a Rac1 GEF/GAP regulatory complex. Dev Cell. 2014;29:701–15.
Castillo-Lluva S, Tatham MH, Jones RC, Jaffray EG, Edmondson RD, Hay RT, et al. SUMOylation of the GTPase Rac1 is required for optimal cell migration. Nat Cell Biol. 2010;12:1078–U70.
Castillo-Lluva S, Tan CT, Daugaard M, Sorensen PHB, Malliri A. The tumour suppressor HACE1 controls cell migration by regulating Rac1 degradation. Oncogene. 2013;32:1735–42.
Yen YC, Shiah SG, Chu HC, Hsu YM, Hsiao JR, Chang JY, et al. Reciprocal regulation of microRNA-99a and insulin-like growth factor I receptor signaling in oral squamous cell carcinoma cells. Mol Cancer. 2014;13:6.
Wong TY, Chen YH, Liu SH, Solis MA, Yu CH, Chang CH, et al. Differential proteomic analysis of human placenta-derived mesenchymal stem cells cultured on normal tissue culture surface and hyaluronan-coated surface. Stem Cells Int. 2016;2016:2809192.
Wang SH, Liou GG, Liu SH, Chang JS, Hsiao JR, Yen YC, et al. Laminin gamma2-enriched extracellular vesicles of oral squamous cell carcinoma cells enhance in vitro lymphangiogenesis via intergrin alpha3-dependent uptake by lymphatic endothelial cells. Int J Cancer. 2018.
Tyanova S, Mann M, Cox J. MaxQuant for in-depth analysis of large SILAC datasets. Methods Mol Biol. 2014;1188:351–64.
Cox J, Neuhauser N, Michalski A, Scheltema RA, Olsen JV, Mann M. Andromeda: a peptide search engine integrated into the MaxQuant environment. J Proteome Res. 2011;10:1794–805.
Apweiler R, Bairoch A, Wu CH, Barker WC, Boeckmann B, Ferro S, et al. UniProt: the Universal Protein knowledgebase. Nucleic Acids Res. 2004;32(Database issue):D115–9.
Chen YW, Paliwal S, Draheim K, Grossman SR, Lewis BC. p19Arf inhibits the invasion of hepatocellular carcinoma cells by binding to C-terminal binding protein. Cancer Res. 2008;68:476–82.
Lin ZS, Chu HC, Yen YC, Lewis BC, Chen YW. Kruppel-like factor 4, a tumor suppressor in hepatocellular carcinoma cells reverts epithelial mesenchymal transition by suppressing slug expression. PLoS ONE. 2012;7:e43593.
Chen RH, Du Y, Han P, Wang HB, Liang FY, Feng GK, et al. ISG15 predicts poor prognosis and promotes cancer stem cell phenotype in nasopharyngeal carcinoma. Oncotarget. 2016;7:16910–22.
Yeh YH, Yang YC, Hsieh MY, Yeh YC, Li TK. A negative feedback of the HIF-1alpha pathway via interferon-stimulated gene 15 and ISGylation. Clin Cancer Res. 2013;19:5927–39.
Festing MF, Altman DG. Guidelines for the design and statistical analysis of experiments using laboratory animals. ILAR J. 2002;43:244–58.
Chen YW, Klimstra DS, Mongeau ME, Tatem JL, Boyartchuk V, Lewis BC. Loss of p53 and Ink4a/Arf cooperate in a cell autonomous fashion to induce metastasis of hepatocellular carcinoma cells. Cancer Res. 2007;67:7589–96.
Chen YW, Chu HC, Ze-Shiang L, Shiah WJ, Chou CP, Klimstra DS, et al. p16 Stimulates CDC42-dependent migration of hepatocellular carcinoma cells. PLoS ONE. 2013;8:e69389.
Acknowledgements
The authors would like to appreciate Drs. Lu-Hai Wang (China Medical University, Taichung, Taiwan) and Tze-Sing Huang (National Health Research Institutes, Miaoli, Taiwan) for providing technical supports in GTPase assay and PLA and Dr. Tsai-Kun Li (National Taiwan University, Taipei, Taiwan) for sharing the ISG15 and ISG15AA plasmids. The authors thank Dr. Szu-Heng Liu (National Health Research Institutes, Miaoli, Taiwan) and the protein core in Institute of Biological Chemistry and Institute of Molecular Biology, Academia Sinica (Taipei, Taiwan) for technical support in SILAC. The authors thank Dr. Yu-Shuen Wang (National Chiao Tung University, Hsinchu, Taiwan) for assistance on program implementation. The authors thank Dr. Shih-Sheng Jiang (National Health Research Institutes, Miaoli, Taiwan) and Taiwan Bioinformatics Institute Core Facility (National Core Facility Program for Biotechnology, MOST 105-2319-B-400-002) for assistances on using Oncomine and data analysis. The histological evaluation was assisted by the Pathology Core Laboratory, National Health Research Institutes, Taiwan. RNAi reagents were purchased from the National RNAi Core Facility at the Institute of Molecular Biology/Genomic Research Center, Academia Sinica, Taiwan.
Funding
This study was supported by grants NHRI CA-107-PP-04 and MOST 106-2314-B-400-026-MY3 from National Health Research Institutes and Ministry of Science and Technology, Taiwan.
Author information
Authors and Affiliations
Contributions
Y-LC performed the work of molecular biology, biochemistry, and cell biology; W-LW performed the proteomic analysis; C-WJ contributed to manuscript preparation; Y-CY, S-HW, and F-YT contributed to the animal work and statistical analysis; Y-YS contributed to the histopathological examination of clinical samples; Y-LC and Y-WC contributed to experimental designs, data analysis, and manuscript writing; all authors contributed to edit the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, YL., Wu, WL., Jang, CW. et al. Interferon-stimulated gene 15 modulates cell migration by interacting with Rac1 and contributes to lymph node metastasis of oral squamous cell carcinoma cells. Oncogene 38, 4480–4495 (2019). https://doi.org/10.1038/s41388-019-0731-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0731-8
This article is cited by
-
SIRT1 ISGylation accelerates tumor progression by unleashing SIRT1 from the inactive state to promote its deacetylase activity
Experimental & Molecular Medicine (2024)
-
Genome-wide DNA methylation profiling of HPV-negative leukoplakia and gingivobuccal complex cancers
Clinical Epigenetics (2023)
-
The diverse repertoire of ISG15: more intricate than initially thought
Experimental & Molecular Medicine (2022)
-
Identification of novel key genes associated with the metastasis of prostate cancer based on bioinformatics prediction and validation
Cancer Cell International (2021)
-
The prognostic significance of interferon-stimulated gene 15 (ISG15) in invasive breast cancer
Breast Cancer Research and Treatment (2021)