Abstract
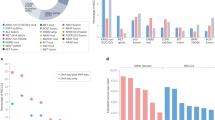
Mutated RAS onco-proteins are key drivers across many cancers. The distribution of somatic RAS mutations varies between cancer entities. Retrospective analyses have associated some RAS mutations with distinct clinical outcomes. However, the clinical impact of the full spectrum of RAS mutations in their disease contextuality remains to be defined. To improve upon this situation, we studied genomically and clinically annotated, prospectively recruited cohorts of patients with RAS-mutated metastatic lung cancer and colorectal cancer. Mutational spectra were compared with predictions derived from analyzing the mutagenic impact at the genome level for each entity. Interestingly, we found concordance of predicted signatures with those actually observed in our patients. Thus, composition of the functionally active RAS mutational subtypes is primarily determined by the mutagenic context. Most RAS mutations seemed dominant oncogenic drivers with entity-dependent clinical outcomes. RAS comutations were enriched in tumors harboring class 2/3 BRAF mutations, highlighting the functional dependency of some mutated BRAF isoforms on RAS. With our dataset, we established a probabilistic model for cross-entity comparison of the prognostic impact of specific RAS mutational subtypes. The resulting prognostic clusters showed largely consistent clinical categorizations in both entities. This suggests mutant subtype-specific functional properties leading to similar clinical effects. A notable exception is KRAS G12C, which imparted an adverse prognosis only in colorectal cancer. Our findings provide a framework for risk stratification of specific RAS mutations across several cancer entities, which is required to guide the analysis of clinical findings in patients treated with direct RAS inhibitors or agents targeting downstream pathways.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Simanshu DK, Nissley DV, McCormick F. RAS proteins and their regulators in human disease. Cell. 2017;170:17–33.
Hunter JC, Manandhar A, Carrasco MA, Gurbani D, Gondi S, Westover KD. Biochemical and structural analysis of common cancer-associated KRAS mutations. Mol Cancer Res. 2015;13:1325–35.
Seeburg PH, Colby WW, Capon DJ, Goeddel DV, Levinson AD. Biological properties of human c-Ha-ras1 genes mutated at codon 12. Nature. 1984;312:71–75.
Guerrero S, Casanova I, Farré L, Mazo A, Capellà G, Mangues R. K-ras codon 12 mutation induces higher level of resistance to apoptosis and predisposition to anchorage-independent growth than codon 13 mutation or proto-oncogene overexpression. Cancer Res. 2000;60:6750–6.
Messner I, Cadeddu G, Huckenbeck W, Knowles HJ, Gabbert HE, Baldus SE, et al. KRAS p.G13D mutations are associated with sensitivity to anti-EGFR antibody treatment in colorectal cancer cell lines. J Cancer Res Clin Oncol. 2013;139:201–9.
Garassino MC, Marabese M, Rusconi P, Rulli E, Martelli O, Farina G, et al. Different types of K-Ras mutations could affect drug sensitivity and tumour behaviour in non-small-cell lung cancer. Ann Oncol. 2011;22:235–7.
Ostrem JM, Peters U, Sos ML, Wells JA, Shokat KM. K-Ras(G12C) inhibitors allosterically control GTP affinity and effector interactions. Nature. 2013;503:548–51.
Lito P, Solomon M, Li L-S, Hansen R, Rosen N. Allele-specific inhibitors inactivate mutant KRAS G12C by a trapping mechanism. Science. 2016;351:604–8.
Patricelli MP, Janes MR, Li L-S, Hansen R, Peters U, Kessler LV, et al. Selective inhibition of oncogenic KRAS output with small molecules targeting the inactive state. Cancer Discov. 2016;6:316–29.
Nan X, Tamgüney TM, Collisson EA, Lin L-J, Pitt C, Galeas J, et al. Ras-GTP dimers activate the mitogen-activated protein kinase (MAPK) pathway. Proc Natl Acad Sci USA. 2015;112:7996–8001.
Ambrogio C, Köhler J, Zhou Z-W, Wang H, Paranal R, Li J, et al. KRAS dimerization impacts MEK inhibitor sensitivity and oncogenic activity of mutant KRAS. Cell. 2018;172:857–868.e15.
Takamochi K, Oh S, Suzuki K. Differences in EGFR and KRAS mutation spectra in lung adenocarcinoma of never and heavy smokers. Oncol Lett. 2013;6:1207–12.
The Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511:543–50.
Prior IA, Lewis PD, Mattos C. A comprehensive survey of Ras mutations in cancer. Cancer Res. 2012;72:2457–67.
McGranahan N, Favero F, de Bruin EC, Birkbak NJ, Szallasi Z, Swanton C. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci Transl Med. 2015;7:283ra54.
Jamal-Hanjani M, Wilson GA, McGranahan N, Birkbak NJ, Watkins TBK, Veeriah S, et al. Tracking the evolution of non–small-cell lung cancer. N Engl J Med. 2017;376:2109–21.
Ostrow SL, Simon E, Prinz E, Bick T, Shentzer T, Nagawkar SS, et al. Variation in KRAS driver substitution distributions between tumor types is determined by both mutation and natural selection. Sci Rep. 2016;6. https://doi.org/10.1038/srep21927.
Haigis KM. KRAS alleles: the devil is in the detail. Trends Cancer. 2017;3:686–97.
Ding L, Getz G, Wheeler DA, Mardis ER, McLellan MD, Cibulskis K, et al. Somatic mutations affect key pathways in lung adenocarcinoma. Nature. 2008;455:1069–75.
Imielinski M, Berger AH, Hammerman PS, Hernandez B, Pugh TJ, Hodis E, et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell. 2012;150:1107–20.
Jordan EJ, Kim HR, Arcila ME, Barron D, Chakravarty D, Gao J, et al. Prospective comprehensive molecular characterization of lung adenocarcinomas for efficient patient matching to approved and emerging therapies. Cancer Discov. 2017;7:596–609.
Giannakis M, Mu XJ, Shukla SA, Qian ZR, Cohen O, Nishihara R, et al. Genomic correlates of immune-cell infiltrates in colorectal carcinoma. Cell Rep. 2016. https://doi.org/10.1016/j.celrep.2016.03.075.
The Cancer Genome Atlas Research Network. Comprehensive genomic characterization of squamous cell lung cancers. Nature. 2012;489:519–25.
Brannon AR, Vakiani E, Sylvester BE, Scott SN, McDermott G, Shah RH, et al. Comparative sequencing analysis reveals high genomic concordance between matched primary and metastatic colorectal cancer lesions. Genome Biol. 2014;15:454.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401–4.
Pfeifer GP, Denissenko MF, Olivier M, Tretyakova N, Hecht SS, Hainaut P. Tobacco smoke carcinogens, DNA damage and p53 mutations in smoking-associated cancers. Oncogene. 2002;21:7435–51.
Kuong KJ, Loeb LA. APOBEC3B mutagenesis in cancer. Nat Genet. 2013;45:964–5.
Zhao H, Thienpont B, Yesilyurt BT, Moisse M, Reumers J, Coenegrachts L, et al. Mismatch repair deficiency endows tumors with a unique mutation signature and sensitivity to DNA double-strand breaks. eLife. 2014;3:e02725.
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SAJR, Behjati S, Biankin AV, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21.
Castellucci E, He T, Goldstein DY, Halmos B, Chuy J. DNA polymerase ɛ deficiency leading to an ultramutator phenotype: a novel clinically relevant entity. Oncologist. 2017;22:497–502.
Wiesweg M, Eberhardt WEE, Reis H, Ting S, Savvidou N, Skiba C, et al. High prevalence of concomitant oncogene mutations in prospectively identified patients with ROS1-positive metastatic lung cancer. J Thorac Oncol. 2017;12:54–64.
Grasso CS, Wu Y-M, Robinson DR, Cao X, Dhanasekaran SM, Khan AP, et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature. 2012;487:239–43.
Toyooka S, Date H, Uchida A, Kiura K, Takata M. The epidermal growth factor receptor D761Y mutation and effect of tyrosine kinase inhibitor. Clin Cancer Res. 2007;13:3431 (author reply 3431-2).
Kerner GSMA, Schuuring E, Sietsma J, Hiltermann TJN, Pieterman RM, Leede GPJde, et al. Common and rare EGFR and KRAS mutations in a Dutch non-small-cell lung cancer population and their clinical outcome. PLoS ONE. 2013;8:e70346.
Yao Z, Yaeger R, Rodrik-Outmezguine VS, Tao A, Torres NM, Chang MT, et al. Tumours with class 3 BRAF mutants are sensitive to the inhibition of activated RAS. Nature. 2017;548:234–8.
Oh B-Y, Lee R-A, Chung S-S, Kim KH. Epidermal growth factor receptor mutations in colorectal cancer patients. J Korean Soc Coloproctol. 2011;27:127–32.
Deihimi S, Lev A, Slifker M, Shagisultanova E, Xu Q, Jung K, et al. BRCA2, EGFR, and NTRK mutations in mismatch repair-deficient colorectal cancers with MSH2 or MLH1 mutations. Oncotarget. 2017;8:39945–62.
Jones JC, Renfro LA, Al-Shamsi HO, Schrock AB, Rankin A, Zhang BY, et al. Non-V600BRAF mutations define a clinically distinct molecular subtype of metastatic colorectal cancer. J Clin Oncol. 2017;35:2624–30.
Petrelli F, Tomasello G, Borgonovo K, Ghidini M, Turati L, Dallera P, et al. Prognostic survival associated with left-sided vs right-sided colon cancer: a systematic review and meta-analysis. JAMA Oncol. 2017;3:211–9.
Tejpar S, Stintzing S, Ciardiello F, Tabernero J, Cutsem EV, Beier F, et al. Prognostic and predictive relevance of primary tumor location in patients with RAS wild-type metastatic colorectal cancer: retrospective analyses of the CRYSTAL and FIRE-3 trials. JAMA Oncol. 2017;3:194–201.
Arnold D, Lueza B, Douillard J-Y, Peeters M, Lenz H-J, Venook A, et al. Prognostic and predictive value of primary tumour side in patients with RAS wild-type metastatic colorectal cancer treated with chemotherapy and EGFR directed antibodies in six randomized trials. Ann Oncol. 2017;28:1713–29.
Lee GH, Malietzis G, Askari A, Bernardo D, Al-Hassi HO, Clark SK. Is right-sided colon cancer different to left-sided colorectal cancer? – A systematic review. Eur J Surg Oncol. 2015;41:300–8.
Lans H, Vermeulen W. Tissue specific response to DNA damage: C. elegans as role model. DNA Repair (Amst). 2015;32:141–8.
Blanpain C, Mohrin M, Sotiropoulou PA, Passegué E. DNA-damage response in tissue-specific and cancer stem cells. Cell Stem Cell. 2011;8:16–29.
Iyama T, Wilson DM. DNA repair mechanisms in dividing and non-dividing cells. DNA Repair (Amst). 2013;12:620–36.
Vitale I, Manic G, De Maria R, Kroemer G, Galluzzi L. DNA damage in stem cells. Mol Cell. 2017;66:306–19.
Karahalil B, Hogue BA, De Souza-Pinto NC, Bohr VA. Base excision repair capacity in mitochondria and nuclei: tissue-specific variations. FASEB J. 2002;16:1895–902.
Dion V. Tissue specificity in DNA repair: lessons from trinucleotide repeat instability. Trends Genet. 2014;30:220–9.
Yao Z, Torres NM, Tao A, Gao Y, Luo L, Li Q, et al. BRAF mutants evade ERK dependent feedback by different mechanisms that determine their sensitivity to pharmacologic inhibition. Cancer Cell. 2015;28:370–83.
Larki P, Gharib E, Yaghoob Taleghani M, Khorshidi F, Nazemalhosseini-Mojarad E, Asadzadeh Aghdaei H. Coexistence of KRAS and BRAF mutations in colorectal cancer: a case report supporting the concept of tumoral heterogeneity. Cell J. 2017;19:113–7.
Vittal A, Middinti A, Kasi Loknath Kumar A. Are all mutations the same? A rare case report of coexisting mutually exclusive KRAS and BRAF mutations in a patient with metastatic colon adenocarcinoma. Case Rep Oncol Med. 2017;2017. https://doi.org/10.1155/2017/2321052.
Sahin IH, Kazmi SMA, Yorio JT, Bhadkamkar NA, Kee BK, Garrett CR. Rare though not mutually exclusive: a report of three cases of concomitant KRAS and BRAF mutation and a review of the literature. J Cancer. 2013;4:320–2.
Zheng G, Tseng L-H, Chen G, Haley L, Illei P, Gocke CD, et al. Clinical detection and categorization of uncommon and concomitant mutations involving BRAF. BMC Cancer. 2015;15. https://doi.org/10.1186/s12885-015-1811-y.
Kwak MS, Cha JM, Yoon JY, Jeon JW, Shin HP, Chang HJ, et al. Prognostic value of KRAS codon 13 gene mutation for overall survival in colorectal cancer. Medicine (Baltimore). 2017;96. https://doi.org/10.1097/MD.0000000000007882.
Blons H, Emile JF, Le Malicot K, Julié C, Zaanan A, Tabernero J, et al. Prognostic value of KRAS mutations in stage III colon cancer: post hoc analysis of the PETACC8 phase III trial dataset. Ann Oncol. 2014;25:2378–85.
Jones RP, Sutton PA, Evans JP, Clifford R, McAvoy A, Lewis J, et al. Specific mutations in KRAS codon 12 are associated with worse overall survival in patients with advanced and recurrent colorectal cancer. Br J Cancer. 2017;116:923–9.
Cserepes M, Ostoros G, Lohinai Z, Raso E, Barbai T, Timar J, et al. Subtype-specific KRAS mutations in advanced lung adenocarcinoma: a retrospective study of patients treated with platinum-based chemotherapy. Eur J Cancer. 2014;50:1819–28.
Shepherd FA, Domerg C, Hainaut P, Jänne PA, Pignon J-P, Graziano S, et al. Pooled analysis of the prognostic and predictive effects of KRAS mutation status and KRAS mutation subtype in early-stage resected non–small-cell lung cancer in four trials of adjuvant chemotherapy. J Clin Oncol. 2013;31:2173–81.
Metro G, Chiari R, Duranti S, Siggillino A, Fischer MJ, Giannarelli D, et al. Impact of specific mutant KRAS on clinical outcome of EGFR-TKI-treated advanced non-small cell lung cancer patients with an EGFR wild type genotype. Lung Cancer. 2012;78:81–86.
Yu HA, Sima CS, Shen R, Kass S, Gainor J, Shaw A, et al. Prognostic impact of KRAS mutation subtypes in 677 patients with metastatic lung adenocarcinomas. J Thorac Oncol. 2015;10:431–7.
Tejpar S, Celik I, Schlichting M, Sartorius U, Bokemeyer C, Van Cutsem E. Association of KRAS G13D tumor mutations with outcome in patients with metastatic colorectal cancer treated with first-line chemotherapy with or without cetuximab. J Clin Oncol. 2012;30:3570–7.
Wiesweg M, Ting S, Reis H, Worm K, Kasper S, Tewes M, et al. Feasibility of preemptive biomarker profiling for personalised early clinical drug development at a Comprehensive Cancer Center. Eur J Cancer. 2013;49:3076–82.
Wiesweg M, Reis H, Köster T, Goetz M, Worm K, Herold T, et al. Phosphorylation of p70 ribosomal protein S6 kinase β-1 is an independent prognostic parameter in metastatic colorectal cancer. Clin Colorectal Cancer. 2018;17:e331–52.
Bose R, Kavuri SM, Searleman AC, Shen W, Shen D, Koboldt DC, et al. Activating HER2 mutations in HER2 gene amplification negative breast cancer. Cancer Discov. 2013;3:224–37.
Xing K, Zhou X, Zhao X, Sun S, Luo Z, Wang H, et al. A novel point mutation in exon 20 of EGFR showed sensitivity to erlotinib. Med Oncol. 2014;31:36.
Acknowledgements
We thank the patients and their families for participation in the Precision Oncology Program of the West German Cancer Center. The invaluable support of the staff of all departments and institutes of the University Hospital Essen and Ruhrlandklinik participating in the WTZ Precision Oncology Program is gratefully acknowledged.
Funding
This work was supported by the Oncology Center of Excellence Program of the Deutsche Krebshilfe (grant number 110534); the German Federal and North Rhine-Westphalian State governments via the German Cancer Consortium (DKTK); Novartis Pharma AG (research grant); the Medical Faculty of the University Duisburg-Essen (IFORES fellowship to M.W.). The funding sources had no influence on the analysis and interpretation of data, and contents of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
We declare the following potential conflicts of interest: Prof. Dr. M. Schuler: Consultancy: AstraZeneca, Boehringer Ingelheim, Bristol-Myers Squibb, Celgene, Institut für Qualität und Wirtschaftlichkeit im Gesundheitswesen (IQWiG), Lilly, Novartis; Honoraria for CME presentations: Alexion, Boehringer Ingelheim, Celgene, GlaxoSmithKline, Lilly, Novartis; Research funding to institution: Boehringer Ingelheim, Bristol Myers-Squibb, Novartis; Other: Universität Duisburg-Essen (Patents). Prof. Dr. S. Kasper: Consultancy, Honoraria and Travel Support: Roche, Bristol-Myers Squibb, Merck Sharp & Dohme. All remaining authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Wiesweg, M., Kasper, S., Worm, K. et al. Impact of RAS mutation subtype on clinical outcome—a cross-entity comparison of patients with advanced non-small cell lung cancer and colorectal cancer. Oncogene 38, 2953–2966 (2019). https://doi.org/10.1038/s41388-018-0634-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0634-0
This article is cited by
-
Targeting KRASG12C in Non-Small-Cell Lung Cancer: Current Standards and Developments
Drugs (2024)
-
RAS oncogenic activity predicts response to chemotherapy and outcome in lung adenocarcinoma
Nature Communications (2022)
-
Targeting Mutated KRAS Genes to Treat Solid Tumours
Molecular Diagnosis & Therapy (2022)
-
Common and mutation specific phenotypes of KRAS and BRAF mutations in colorectal cancer cells revealed by integrative -omics analysis
Journal of Experimental & Clinical Cancer Research (2021)
-
Biomarker-guided therapy for colorectal cancer: strength in complexity
Nature Reviews Clinical Oncology (2020)