Abstract
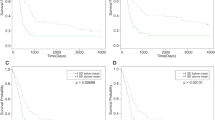
Glioblastomas (GBMs) are the most aggressive primary brain tumors, with an average survival of less than 15 months. Therefore, there is a critical need to develop novel therapeutic strategies for GBM. This study aimed to assess the prognostic value of miR-4516 and investigate its oncogenic functions and the underlying cellular and molecular mechanisms in GBM. To determine the correlation between miR-4516 expression and overall survival of patients with GBM, total RNAs were isolated from 268 FFPE tumor samples, miR expression was assayed (simultaneously) using the nCounter human miRNA v3a assay followed by univariable and multivariable survival analyses. Further, in vitro and in vivo studies were conducted to define the role of miR-4516 in GBM tumorigenesis and the underlying molecular mechanisms. Upon multivariable analysis, miR-4516 was correlated with poor prognosis in GBM patients (HR = 1.49, 95%CI: 1.12–1.99, P = 0.01). Interestingly, the significance of miR-4516 was retained including MGMT methylation status. Overexpression of miR-4516 significantly enhanced cell proliferation and invasion of GBM cells both in vitro and in vivo. While conducting downstream targeting studies, we found that the tumor-promoting function of miR-4516, in part, was mediated by direct targeting of PTPN14 (protein tyrosine phosphatase, non-receptor type 14) which, in turn, regulated the Hippo pathway in GBM. Taken together, our data suggest that miR-4516 represents an independent negative prognostic factor in GBM patients and acts as a novel oncogene in GBM, which regulates the PTPN14/Hippo pathway. Thus, this newly identified miR-4516 may serve as a new potential therapeutic target for GBM treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ostrom QT, Gittleman H, Fulop J, Liu M, Blanda R, Kromer C, et al. CBTRUS Statistical Report: Primary Brain and Central Nervous System Tumors Diagnosed in the United States in 2008-2012. Neuro Oncol. 2015;17(Suppl 4):iv1–iv62.
Losman JA, Kaelin WG Jr. What a difference a hydroxyl makes: mutant IDH, (R)-2-hydroxyglutarate, and cancer. Genes Dev. 2013;27:836–52.
Lenos K, Grawenda AM, Lodder K, Kuijjer ML, Teunisse AF, Repapi E, et al. Alternate splicing of the p53 inhibitor HDMX offers a superior prognostic biomarker than p53 mutation in human cancer. Cancer Res. 2012;72:4074–84.
Zougman A, Hutchins GG, Cairns DA, Verghese E, Perry SL, Jayne DG, et al. Retinoic acid-induced protein 3: identification and characterisation of a novel prognostic colon cancer biomarker. Eur J Cancer. 2013;49:531–9.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–31.
Garzon R, Fabbri M, Cimmino A, Calin GA, Croce CM. MicroRNA expression and function in cancer. Trends Mol Med. 2006;12:580–7.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Shenouda SK, Alahari SK. MicroRNA function in cancer: oncogene or a tumor suppressor? Cancer Metastas Rev. 2009;28:369–78.
Zhang B, Pan X, Cobb GP, Anderson TA. microRNAs as oncogenes and tumor suppressors. Dev Biol. 2007;302:1–12.
Xia H, Yan Y, Hu M, Wang Y, Wang Y, Dai Y, et al. MiR-218 sensitizes glioma cells to apoptosis and inhibits tumorigenicity by regulating ECOP-mediated suppression of NF-kappaB activity. Neuro Oncol. 2013;15:413–22.
Li S, Zeng A, Hu Q, Yan W, Liu Y, You Y. miR-423-5p contributes to a malignant phenotype and temozolomide chemoresistance in glioblastomas. Neuro Oncol. 2017;19:55–65.
Chang W, Liu M, Xu J, Fu H, Zhou B, Yuan T, et al. MiR-377 inhibits the proliferation of pancreatic cancer by targeting Pim-3. Tumour Biol. 2016;37:14813–24.
Du W, Liu X, Chen L, Dou Z, Lei X, Chang L, et al. Targeting the SMO oncogene by miR-326 inhibits glioma biological behaviors and stemness. Neuro Oncol. 2015;17:243–53.
Belle L, Ali N, Lonic A, Li X, Paltridge JL, Roslan S, et al. The tyrosine phosphatase PTPN14 (Pez) inhibits metastasis by altering protein trafficking. Sci Signal. 2015;8:ra18.
Laczmanska I, Sasiadek MM. Tyrosine phosphatases as a superfamily of tumor suppressors in colorectal cancer. Acta Biochim Pol. 2011;58:467–70.
Liu X, Yang N, Figel SA, Wilson KE, Morrison CD, Gelman IH, et al. PTPN14 interacts with and negatively regulates the oncogenic function of YAP. Oncogene. 2013;32:1266–73.
Wang LJ, He CC, Sui X, Cai MJ, Zhou CY, Ma JL, et al. MiR-21 promotes intrahepatic cholangiocarcinoma proliferation and growth in vitro and in vivo by targeting PTPN14 and PTEN. Oncotarget. 2015;6:5932–46.
Chowdhari S, Saini N. hsa-miR-4516 mediated downregulation of STAT3/CDK6/UBE2N plays a role in PUVA induced apoptosis in keratinocytes. J Cell Physiol. 2014;229:1630–8.
Michaloglou C, Lehmann W, Martin T, Delaunay C, Hueber A, Barys L, et al. The tyrosine phosphatase PTPN14 is a negative regulator of YAP activity. PLoS ONE. 2013;8:e61916.
Wang W, Huang J, Wang X, Yuan J, Li X, Feng L, et al. PTPN14 is required for the density-dependent control of YAP1. Genes Dev. 2012;26:1959–71.
Pan D. The hippo signaling pathway in development and cancer. Dev Cell. 2010;19:491–505.
Yu FX, Zhao B, Guan KL. Hippo pathway in organ size control, tissue homeostasis, and cancer. Cell. 2015;163:811–28.
Zhao B, Li L, Lei Q, Guan KL. The Hippo-YAP pathway in organ size control and tumorigenesis: an updated version. Genes Dev. 2010;24:862–74.
Varelas X. The Hippo pathway effectors TAZ and YAP in development, homeostasis and disease. Development. 2014;141:1614–26.
Stupp R, Hegi ME, Mason WP, van den Bent MJ, Taphoorn MJ, Janzer RC, et al. Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol. 2009;10:459–66.
McNamara MG, Sahebjam S, Mason WP. Emerging biomarkers in glioblastoma. Cancers. 2013;5:1103–19.
Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100:57–70.
Bell EH, Kirste S, Fleming JL, Stegmaier P, Drendel V, Mo X, et al. A novel miRNA-based predictive model for biochemical failure following post-prostatectomy salvage radiation therapy. PLoS ONE. 2015;10:e0118745.
Borrelli N, Denaro M, Ugolini C, Poma AM, Miccoli M, Vitti P, et al. miRNA expression profiling of 'noninvasive follicular thyroid neoplasms with papillary-like nuclear features' compared with adenomas and infiltrative follicular variants of papillary thyroid carcinomas. Mod Pathol. 2017;30:39–51.
Wilson KE, Li YW, Yang N, Shen H, Orillion AR, Zhang J. PTPN14 forms a complex with Kibra and LATS1 proteins and negatively regulates the YAP oncogenic function. J Biol Chem. 2014;289:23693–700.
Team RC. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2016.
Bady P, Sciuscio D, Diserens AC, Bloch J, van den Bent MJ, Marosi C, et al. MGMT methylation analysis of glioblastoma on the Infinium methylation BeadChip identifies two distinct CpG regions associated with gene silencing and outcome, yielding a prediction model for comparisons across datasets, tumor grades, and CIMP-status. Acta Neuropathol. 2012;124:547–60.
Maksimovic J, Gordon L, Oshlack A. SWAN: Subset-quantile within array normalization for illumina infinium HumanMethylation450 BeadChips. Genome Biol. 2012;13:R44.
Fortin JP, Triche TJ Jr, Hansen KD. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinformatics. 2017;33:558–60.
Cui T, Chen Y, Yang L, Knosel T, Huber O, Pacyna-Gengelbach M, et al. The p53 target gene desmocollin 3 acts as a novel tumor suppressor through inhibiting EGFR/ERK pathway in human lung cancer. Carcinogenesis. 2012;33:2326–33.
Han C, Zhao R, Liu X, Srivastava A, Gong L, Mao H, et al. DDB2 suppresses tumorigenicity by limiting the cancer stem cell population in ovarian cancer. Mol Cancer Res. 2014;12:784–94.
Cui T, Srivastava AK, Han C, Yang L, Zhao R, Zou N, et al. XPC inhibits NSCLC cell proliferation and migration by enhancing E-Cadherin expression. Oncotarget. 2015;6:10060–72.
Wong N, Wang X. miRDB: an online resource for microRNA target prediction and functional annotations. Nucleic Acids Res. 2015;43(Database issue):D146–52.
Agarwal V, Bell GW, Nam JW, Bartel DP. Predicting effective microRNA target sites in mammalian mRNAs. eLife. 2015;4:e05005. https://doi.org/10.7554/eLife.05005
Hsu SD, Lin FM, Wu WY, Liang C, Huang WC, Chan WL, et al. miRTarBase: a database curates experimentally validated microRNA-target interactions. Nucleic Acids Res. 2011;39(Database issue):D163–9.
Benjamini Y, Hochberg Y. Controlling the false discovery rate - a practical and powerful approach to multiple testing. J Roy Stat Soc B Met. 1995;57:289–300.
Acknowledgements
We thank the T&P Bohnenn Fund for Neuro-Oncology Research (grant to P.A.R.), the Ohio State University (OSU) Comprehensive Cancer Center Small Animal Imaging Core, and the Ohio State University (OSU) Comprehensive Cancer Center Pathology Core Facility supported in part by grant P30 CA016058, National Cancer Institute, Bethesda, MD.
Funding
This work was supported by National Cancer Institute [R01CA169368 (to A.C.), R01CA11522358 (to A.C.), R01CA1145128 (to A.C.), R01CA108633 (to A.C.), R01CA188228 (to A.C., R.B., K.L., and J.B.), 1RC2CA148190 (to A.C.), and U10CA180850–01 (to A.C.)]; A Brain Tumor Funders Collaborative Grant (to A.C.); Ohio State University Comprehensive Cancer Center Award (to A.C.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Cui, T., Bell, E.H., McElroy, J. et al. miR-4516 predicts poor prognosis and functions as a novel oncogene via targeting PTPN14 in human glioblastoma. Oncogene 38, 2923–2936 (2019). https://doi.org/10.1038/s41388-018-0601-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0601-9
This article is cited by
-
Therapeutic potential of melatonin in targeting molecular pathways of organ fibrosis
Pharmacological Reports (2024)
-
H3K27ac-activated EGFR-AS1 promotes cell growth in cervical cancer through ACTN4-mediated WNT pathway
Biology Direct (2022)
-
MiR-874 Inhibits Cell Proliferation, Migration, and Invasion of Glioma Cells and Correlates with Prognosis of Glioma Patients
NeuroMolecular Medicine (2021)
-
Upregulation of miR-1825 inhibits the progression of glioblastoma by suppressing CDK14 though Wnt/β-catenin signaling pathway
World Journal of Surgical Oncology (2020)
-
Expression and clinical significance of miR-4516 and miR-21-5p in serum of patients with colorectal cancer
BMC Cancer (2020)