Abstract
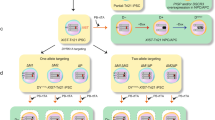
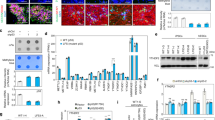
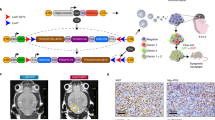
Mutations in the isocitrate dehydrogenase 1 (IDH1) gene have been identified in a number of cancer types, including brain cancer. The Cancer Genome Atlas project has revealed that IDH1 mutations occur in 70–80% of grade II and grade III gliomas. Until recently, most of the functional studies of IDH1 mutations in cellular models have been conducted in overexpression systems with the IDH1 wild type background. In this study, we employed a modified CRISPR/Cas9 genome editing technique called “single base editing”, and efficiently introduced heterozygous IDH1 R132H mutation (IDH1R132H/WT) in human astroglial cells. Global DNA methylation profiling revealed hypermethylation as well as hypomethylation induced by IDH1R132H/WT. Global gene expression analysis identified molecular targets and pathways altered by IDH1R132H/WT, including cell proliferation, extracellular matrix (ECM), and cell migration. Our phenotype analysis indicated that compared with IDH1 wild type cells, IDH1R132H/WT promoted cell migration by upregulating integrin β4 (ITGB4); and significantly inhibited cell proliferation. Using our mutated IDH1 models generated by “single base editing”, we identified novel molecular targets of IDH1R132H/WT, namely Yes-associated protein (YAP) and its downstream signaling pathway Notch, to mediate the cell growth-inhibiting effect of IDH1R132H/WT. In summary, the “single base editing” strategy has successfully created heterozygous IDH1 R132H mutation that recapitulates the naturally occurring IDH1 mutation. Our isogenic cellular systems that differ in a single nucleotide in one allele of the IDH1 gene provide a valuable model for novel discoveries of IDH1R132H/WT-driven biological events.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Parsons DW, Jones S, Zhang XS, Lin JCH, Leary RJ, Angenendt P, et al. An integrated genomic analysis of human glioblastoma Multiforme. Science. 2008;321:1807–12.
Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W, et al. IDH1 and IDH2 mutations in gliomas. N Engl J Med. 2009;360:765–73.
Hartmann C, Meyer J, Balss J, Capper D, Mueller W, Christians A, et al. Type and frequency of IDH1 and IDH2 mutations are related to astrocytic and oligodendroglial differentiation and age: a study of 1,010 diffuse gliomas. Acta Neuropathol. 2009;118:469–74.
Mardis ER, Ding L, Dooling DJ, Larson DE, McLellan MD, Chen K, et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N Engl J Med. 2009;361:1058–66.
Kang MR, Kim MS, Oh JE, Kim YR, Song SY, Seo SI, et al. Mutational analysis of IDH1 codon 132 in glioblastomas and other common cancers. Int J Cancer. 2009;125:353–5.
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, et al. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol. 2016;131:803–20.
Watanabe T, Nobusawa S, Kleihues P, Ohgaki H. IDH1 mutations are early events in the development of astrocytomas and oligodendrogliomas. Am J Pathol. 2009;174:1149–53.
Labussiere M, Idbaih A, Wang XW, Marie Y, Boisselier B, Falet C, et al. All the 1p19q codeleted gliomas are mutated on IDH1 or IDH2. Neurology. 2010;74:1886–90.
Piaskowski S, Bienkowski M, Stoczynska-Fidelus E, Stawski R, Sieruta M, Szybka M, et al. Glioma cells showing IDH1 mutation cannot be propagated in standard cell culture conditions. Br J Cancer. 2011;104:968–70.
Luchman HA, Stechishin OD, Dang NH, Blough MD, Chesnelong C, Kelly JJ, et al. An in vivo patient-derived model of endogenous IDH1-mutant glioma. Neuro Oncol. 2012;14:184–91.
Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering. Cell. 2014;157:1262–78.
Richardson CD, Ray GJ, DeWitt MA, Curie GL, Corn JE. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat Biotechnol. 2016;34:339–44.
Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature. 2016;533:420–4.
Major EO, Miller AE, Mourrain P, Traub RG, de Widt E, Sever J. Establishment of a line of human fetal glial cells that supports JC virus multiplication. Proc Natl Acad Sci USA. 1985;82:1257–61.
Ferenczy MW, Johnson KR, Steinberg SM, Marshall LJ, Monaco MC, Beschloss AM, et al. Clonal immortalized human glial cell lines support varying levels of JC virus infection due to differences in cellular gene expression. J Neuroimmune Pharmacol. 2013;8:1303–19.
Bargonetti J, Reynisdottir I, Friedman PN, Prives C. Site-specific binding of wild-type p53 to cellular DNA is inhibited by SV40 T antigen and mutant p53. Genes Dev. 1992;6:1886–98.
Duncan CG, Barwick BG, Jin G, Rago C, Kapoor-Vazirani P, Powell DR, et al. A heterozygous IDH1R132H/WT mutation induces genome-wide alterations in DNA methylation. Genome Res. 2012;22:2339–55.
Turcan S, Rohle D, Goenka A, Walsh LA, Fang F, Yilmaz E, et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature. 2012;483:479–U137.
Flavahan WA, Drier Y, Liau BB, Gillespie SM, Venteicher AS, Stemmer-Rachamimov AO, et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature. 2016;529:110–4.
Gumbiner BM, Kim NG. The Hippo-YAP signaling pathway and contact inhibition of growth. J Cell Sci. 2014;127:709–17.
Licciardello MP, Mullner MK, Durnberger G, Kerzendorfer C, Boidol B, Trefzer C, et al. NOTCH1 activation in breast cancer confers sensitivity to inhibition of SUMOylation. Oncogene. 2015;34:3780–90.
Konsavage WM Jr., Kyler SL, Rennoll SA, Jin G, Yochum GS. Wnt/beta-catenin signaling regulates Yes-associated protein (YAP) gene expression in colorectal carcinoma cells. J Biol Chem. 2012;287:11730–9.
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell. 2010;17:98–110.
Ma S, Jiang B, Deng W, Gu ZK, Wu FZ, Li T, et al. D-2-hydroxyglutarate is essential for maintaining oncogenic property of mutant IDH-containing cancer cells but dispensable for cell growth. Oncotarget. 2015;6:8606–20.
Sulkowski PL, Corso CD, Robinson ND, Scanlon SE, Purshouse KR, Bai H, et al. 2-Hydroxyglutarate produced by neomorphic IDH mutations suppresses homologous recombination and induces PARP inhibitor sensitivity. Sci Transl Med. 2017; 9:eaal2463.
Wan J, Su Y, Song Q, Tung B, Oyinlade O, Liu S, et al. Methylated cis-regulatory elements mediate KLF4-dependent gene transactivation and cell migration. eLife. 2017;6:e20068.
Lynch MD, Smith AJ, De Gobbi M, Flenley M, Hughes JR, Vernimmen D, et al. An interspecies analysis reveals a key role for unmethylated CpG dinucleotides in vertebrate Polycomb complex recruitment. EMBO J. 2012;31:317–29.
Sasaki M, Knobbe CB, Itsumi M, Elia AJ, Harris IS, Chio IIC, et al. D-2-hydroxyglutarate produced by mutant IDH1 perturbs collagen maturation and basement membrane function. Gene Dev. 2012;26:2038–49.
Cui D, Ren J, Shi J, Feng L, Wang K, Zeng T, et al. R132H mutation in IDH1 gene reduces proliferation, cell survival and invasion of human glioma by downregulating Wnt/beta-catenin signaling. Int J Biochem Cell Biol. 2016;73:72–81.
Fu Y, Zheng Y, Li K, Huang R, Zheng S, An N, et al. Mutations in isocitrate dehydrogenase 2 accelerate glioma cell migration via matrix metalloproteinase-2 and 9. Biotechnol Lett. 2012;34:441–6.
Sabit H, Nakada M, Furuta T, Watanabe T, Hayashi Y, Sato H, et al. Characterizing invading glioma cells based on IDH1-R132H and Ki-67 immunofluorescence. Brain Tumor Pathol. 2014;31:242–6.
Miroshnikova YA, Mouw JK, Barnes JM, Pickup MW, Lakins JN, Kim Y, et al. Tissue mechanics promote IDH1-dependent HIF1alpha-tenascin C feedback to regulate glioblastoma aggression. Nat Cell Biol. 2016;18:1336–45.
Shi J, Zuo H, Ni L, Xia L, Zhao L, Gong M, et al. An IDH1 mutation inhibits growth of glioma cells via GSH depletion and ROS generation. Neurol Sci. 2014;35:839–45.
Wang G, Sai K, Gong F, Yang Q, Chen F, Lin J. Mutation of isocitrate dehydrogenase 1 induces glioma cell proliferation via nuclear factor-kappaB activation in a hypoxia-inducible factor 1-alpha dependent manner. Mol Med Rep. 2014;9:1799–805.
Moroishi T, Hansen CG, Guan KL. The emerging roles of YAP and TAZ in cancer. Nat Rev Cancer. 2015;15:73–79.
Tschaharganeh DF, Chen X, Latzko P, Malz M, Gaida MM, Felix K, et al. Yes-associated protein up-regulates Jagged-1 and activates the Notch pathway in human hepatocellular carcinoma. Gastroenterology. 2013;144:1530–42. e1512.
Xia S, Lal B, Tung B, Wang S, Goodwin CR, Laterra J, et al. Tumor microenvironment tenascin-C promotes glioblastoma invasion and negatively regulates tumor proliferation. Neuro Oncol. 2016;18:507–517.
Goodwin CR, Lal B, Zhou X, Ho S, Xia S, Taeger A, et al. Cyr61 mediates hepatocyte growth factor-dependent tumor cell growth, migration, and Akt activation. Cancer Res. 2010;70:2932–41.
Reznik TE, Sang Y, Ma Y, Abounader R, Rosen EM, Xia S, et al. Transcription-dependent epidermal growth factor receptor activation by hepatocyte growth factor. Mol Cancer Res. 2008;6:139–50.
Assenov Y, Muller F, Lutsik P, Walter J, Lengauer T, Bock C. Comprehensive analysis of DNA methylation data with RnBeads. Nat Methods. 2014;11:1138–40.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Anders S, Pyl PT, Huber W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–9.
Anders S, Huber W. Differential expression analysis for sequence count data. Genome Biol. 2010;11:R106.
Shah SR, David JM, Tippens ND, Mohyeldin A, Martinez-Gutierrez JC, Ganaha S, et al. Brachyury-YAP Regulatory Axis Drives Stemness and Growth in Cancer. Cell Rep. 2017;21:495–507.
Acknowledgements
We thank Dr. G. Major from NIH/NINDS for SVG cells, and Dr. Mingyao Ying from Kennedy Krieger Institute for YAP constructs. We thank Dr. John Laterra and Dr. Hernando Lopez for comments on this work. This work was supported by grants from NIH R01NS091165 (S.X.), NIH EY024580 (J.Q.), NIH EY023188 (J.Q.), and NIH GM111514 (H.Z. and J.Q.), Ford Foundation pre-doctoral fellowship program (O.O.) and NIH T32 GM007445 (O.O.).
Author contributions:
Shang Wei, O.O., L.K., and Q. X.: conducted experiments, data collection, figure preparation, and manuscript writing; J.W.: bioinformatics analysis, figure preparation, and manuscript writing; D.M., Shuyan Wang, B.L., S. L., H.Z., and Y.L.: conducted experiments and data collection; S.R.S., A.Q.H., and S.L.: sample collection and manuscript writing; L.C.: experimental design and manuscript writing; J.Q.: bioinformatics analysis and financial support; S.X.: experiment design, financial support, figure preparation, and manuscript writing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Wei, S., Wang, J., Oyinlade, O. et al. Heterozygous IDH1R132H/WT created by “single base editing” inhibits human astroglial cell growth by downregulating YAP. Oncogene 37, 5160–5174 (2018). https://doi.org/10.1038/s41388-018-0334-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0334-9
This article is cited by
-
The development of a hiPSC-based platform to identify tissue-dependencies of IDH1 R132H
Cell Death Discovery (2023)
-
Advances in the Immunotherapeutic Potential of Isocitrate Dehydrogenase Mutations in Glioma
Neuroscience Bulletin (2022)
-
Current biomarker-associated procedures of cancer modeling-a reference in the context of IDH1 mutant glioma
Cell Death & Disease (2020)
-
A microfluidic cell-migration assay for the prediction of progression-free survival and recurrence time of patients with glioblastoma
Nature Biomedical Engineering (2020)