Abstract
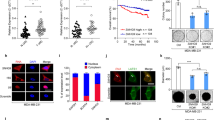
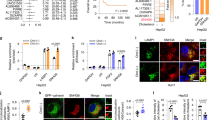
Breast cancer is a leading cause of cancer-related death in women. Small open reading frame (sORF)-encoded proteins or microproteins constitute a new class of molecules often transcribed from presumed long non-coding RNA transcripts (lncRNAs). The translation of some of these sORFs has been confirmed, but their cellular function and importance remains largely unknown. Here, we report the identification and characterization of a novel microprotein of 10 kDa, which we named Cancer-Associated Small Integral Membrane Open reading frame 1 (CASIMO1). CASIMO1 RNA is overexpressed predominantly in hormone receptor-positive breast tumors. Its knockdown leads to decreased proliferation in multiple breast cancer cell lines. Its loss disturbs the organization of the actin cytoskeleton, leads to inhibition of cell motility, and causes a G0/G1 cell cycle arrest. The proliferation phenotype upon overexpression is observed only with CASIMO1 protein expression, but not with a non-translatable mutant attributing the effects to the sORF-derived protein rather than a lncRNA function. CASIMO1 microprotein interacts with squalene epoxidase (SQLE), a key enzyme in cholesterol synthesis and a known oncogene in breast cancer. Overexpression of CASIMO1 leads to SQLE protein accumulation without affecting its RNA levels and increased lipid droplet clustering, while knockdown of CASIMO1 decreased SQLE protein abundance and ERK phosphorylation downstream of SQLE. Importantly, SQLE knockdown mimicked the CASIMO1 knockdown phenotype and in turn SQLE overexpression fully rescued the effect of CASIMO1 knockdown. These findings establish CASIMO1 as the first functional microprotein that plays a role in carcinogenesis and is implicated in the cell lipid homeostasis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–386.
Skibinski A, Kuperwasser C. The origin of breast tumor heterogeneity. Oncogene. 2015;34:5309–16.
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, et al. Molecular portraits of human breast tumours. Nature. 2000;406:747–52.
Slavoff SA, Mitchell AJ, Schwaid AG, Cabili MN, Ma J, Levin JZ, et al. Peptidomic discovery of short open reading frame-encoded peptides in human cells. Nat Chem Biol. 2013;9:59–64.
Bazzini AA, Johnstone TG, Christiano R, Mackowiak SD, Obermayer B, Fleming ES, et al. Identification of small ORFs in vertebrates using ribosome footprinting and evolutionary conservation. EMBO J. 2014;33:981–93.
Aspden JL, Eyre-Walker YC, Phillips RJ, Amin U, Mumtaz MA, Brocard M, et al. Extensive translation of small Open Reading Frames revealed by Poly-Ribo-Seq. Elife. 2014;3:e03528.
Mackowiak SD, Zauber H, Bielow C, Thiel D, Kutz K, Calviello L, et al. Extensive identification and analysis of conserved small ORFs in animals. Genome Biol. 2015;16:179.
Chu Q, Rathore A, Diedrich JK, Donaldson CJ, Yates JR 3rd, Saghatelian A. Identification of microprotein-protein interactions via APEX tagging. Biochemistry. 2017;56:3299–306.
D’Lima NG, Ma J, Winkler L, Chu Q, Loh KH, Corpuz EO, et al. A human microprotein that interacts with the mRNA decapping complex. Nat Chem Biol. 2017;13:174–80.
Zhang Q, Vashisht AA, O’Rourke J, Corbel SY, Moran R, Romero A, et al. The microprotein Minion controls cell fusion and muscle formation. Nat Commun. 2017;8:15664.
Basrai MA, Hieter P, Boeke JD. Small open reading frames: beautiful needles in the haystack. Genome Res. 1997;7:768–71.
Andrews SJ, Rothnagel JA. Emerging evidence for functional peptides encoded by short open reading frames. Nat Rev Genet. 2014;15:193–204.
Anderson DM, Anderson KM, Chang CL, Makarewich CA, Nelson BR, McAnally JR, et al. A micropeptide encoded by a putative long noncoding RNA regulates muscle performance. Cell. 2015;160:595–606.
Nelson BR, Makarewich CA, Anderson DM, Winders BR, Troupes CD, Wu F, et al. A peptide encoded by a transcript annotated as long noncoding RNA enhances SERCA activity in muscle. Science. 2016;351:271–5.
Kondo T, Plaza S, Zanet J, Benrabah E, Valenti P, Hashimoto Y, et al. Small peptides switch the transcriptional activity of Shavenbaby during Drosophila embryogenesis. Science. 2010;329:336–9.
Magny EG, Pueyo JI, Pearl FM, Cespedes MA, Niven JE, Bishop SA, et al. Conserved regulation of cardiac calcium uptake by peptides encoded in small open reading frames. Science. 2013;341:1116–20.
Kastenmayer JP, Ni L, Chu A, Kitchen LE, Au WC, Yang H, et al. Functional genomics of genes with small open reading frames (sORFs) in S. cerevisiae. Genome Res. 2006;16:365–73.
Wadler CS, Vanderpool CK. A dual function for a bacterial small RNA: SgrS performs base pairing-dependent regulation and encodes a functional polypeptide. Proc Natl Acad Sci USA. 2007;104:20454–9.
Lluch-Senar M, Delgado J, Chen WH, Llorens-Rico V, O’Reilly FJ, Wodke JA, et al. Defining a minimal cell: essentiality of small ORFs and ncRNAs in a genome-reduced bacterium. Mol Syst Biol. 2015;11:780.
Bi P, Ramirez-Martinez A, Li H, Cannavino J, McAnally JR, Shelton JM, et al. Control of muscle formation by the fusogenic micropeptide myomixer. Science. 2017;356:323–7.
Hashimoto Y, Niikura T, Tajima H, Yasukawa T, Sudo H, Ito Y, et al. A rescue factor abolishing neuronal cell death by a wide spectrum of familial Alzheimer’s disease genes and Abeta. Proc Natl Acad Sci USA. 2001;98:6336–41.
Lee C, Zeng J, Drew BG, Sallam T, Martin-Montalvo A, Wan J, et al. The mitochondrial-derived peptide MOTS-c promotes metabolic homeostasis and reduces obesity and insulin resistance. Cell Metab. 2015;21:443–54.
Slavoff SA, Heo J, Budnik BA, Hanakahi LA, Saghatelian A. A human short open reading frame (sORF)-encoded polypeptide that stimulates DNA end joining. J Biol Chem. 2014;289:10950–7.
Dinger ME, Pang KC, Mercer TR, Mattick JS. Differentiating protein-coding and noncoding RNA: challenges and ambiguities. PLoS Comput Biol. 2008;4:e1000176.
Cancer Genome Atlas N. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70.
Li J, Han L, Roebuck P, Diao L, Liu L, Yuan Y, et al. TANRIC: an interactive open platform to explore the function of lncRNAs in cancer. Cancer Res. 2015;75:3728–37.
Leucci E, Vendramin R, Spinazzi M, Laurette P, Fiers M, Wouters J, et al. Melanoma addiction to the long non-coding RNA SAMMSON. Nature. 2016;531:518–22.
Sharova LV, Sharov AA, Nedorezov T, Piao Y, Shaik N, Ko MS. Database for mRNA half-life of 19 977 genes obtained by DNA microarray analysis of pluripotent and differentiating mouse embryonic stem cells. DNA Res. 2009;16:45–58.
Bhattacharyya S, Tobacman JK. Arylsulfatase B regulates colonic epithelial cell migration by effects on MMP9 expression and RhoA activation. Clin Exp Metastasis. 2009;26:535–45.
Schwanhausser B, Busse D, Li N, Dittmar G, Schuchhardt J, Wolf J, et al. Global quantification of mammalian gene expression control. Nature. 2011;473:337–42.
Ingolia NT, Brar GA, Stern-Ginossar N, Harris MS, Talhouarne GJ, Jackson SE, et al. Ribosome profiling reveals pervasive translation outside of annotated protein-coding genes. Cell Rep. 2014;8:1365–79.
Ingolia NT, Lareau LF, Weissman JS. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell. 2011;147:789–802.
Ma J, Ward CC, Jungreis I, Slavoff SA, Schwaid AG, Neveu J, et al. Discovery of human sORF-encoded polypeptides (SEPs) in cell lines and tissue. J Proteome Res. 2014;13:1757–65.
Mestdagh P, Lefever S, Pattyn F, Ridzon D, Fredlund E, Fieuw A, et al. The microRNA body map: dissecting microRNA function through integrative genomics. Nucleic Acids Res. 2011;39:e136.
Riedl J, Crevenna AH, Kessenbrock K, Yu JH, Neukirchen D, Bista M, et al. Lifeact: a versatile marker to visualize F-actin. Nat Methods. 2008;5:605–7.
Hannus M, Beitzinger M, Engelmann JC, Weickert MT, Spang R, Hannus S, et al. siPools: highly complex but accurately defined siRNA pools eliminate off-target effects. Nucleic Acids Res. 2014;42:8049–61.
Gutschner T, Baas M, Diederichs S. Noncoding RNA gene silencing through genomic integration of RNA destabilizing elements using zinc finger nucleases. Genome Res. 2011;21:1944–54.
Staudt AC, Wenkel S. Regulation of protein function by ‘microProteins’. EMBO Rep. 2011;12:35–42.
Couso JP, Patraquim P. Classification and function of small open reading frames. Nat Rev Mol Cell Biol. 2017;18:575–89.
Cabrera-Quio LE, Herberg S, Pauli A. Decoding sORF translation—from small proteins to gene regulation. RNA Biol. 2016;13:1051–9.
Thurnher M, Nussbaumer O, Gruenbacher G. Novel aspects of mevalonate pathway inhibitors as antitumor agents. Clin Cancer Res. 2012;18:3524–31.
Gill S, Stevenson J, Kristiana I, Brown AJ. Cholesterol-dependent degradation of squalene monooxygenase, a control point in cholesterol synthesis beyond HMG-CoA reductase. Cell Metab. 2011;13:260–73.
Brown DN, Caffa I, Cirmena G, Piras D, Garuti A, Gallo M, et al. Squalene epoxidase is a bona fide oncogene by amplification with clinical relevance in breast cancer. Sci Rep. 2016;6:19435.
Sui Z, Zhou J, Cheng Z, Lu P. Squalene epoxidase (SQLE) promotes the growth and migration of the hepatocellular carcinoma cells. Tumour Biol: J Int Soc Oncodev Biol Med. 2015;36:6173–9.
Zhang HY, Li HM, Yu Z, Yu XY, Guo K. Expression and significance of squalene epoxidase in squamous lung cancerous tissues and pericarcinoma tissues. Thorac Cancer. 2014;5:275–80.
Helms MW, Kemming D, Pospisil H, Vogt U, Buerger H, Korsching E, et al. Squalene epoxidase, located on chromosome 8q24.1, is upregulated in 8q+breast cancer and indicates poor clinical outcome in stage I and II disease. Br J Cancer. 2008;99:774–80.
Simigdala N, Gao Q, Pancholi S, Roberg-Larsen H, Zvelebil M, Ribas R, et al. Cholesterol biosynthesis pathway as a novel mechanism of resistance to estrogen deprivation in estrogen receptor-positive breast cancer. Breast Cancer Res. 2016;18:58.
Ryder NS. Terbinafine: mode of action and properties of the squalene epoxidase inhibition. Br J Dermatol. 1992;126:2–7.
Ryder NS, Dupont MC. Inhibition of squalene epoxidase by allylamine antimycotic compounds. A comparative study of the fungal and mammalian enzymes. Biochem J. 1985;230:765–70.
Ta MT, Kapterian TS, Fei W, Du X, Brown AJ, Dawes IW, et al. Accumulation of squalene is associated with the clustering of lipid droplets. FEBS J. 2012;279:4231–44.
Stevenson J, Luu W, Kristiana I, Brown AJ. Squalene mono-oxygenase, a key enzyme in cholesterol synthesis, is stabilized by unsaturated fatty acids. Biochem J. 2014;461:435–42.
Ji Z, Song R, Regev A, Struhl K. Many lncRNAs, 5′UTRs, and pseudogenes are translated and some are likely to express functional proteins. Elife. 2015;4:e08890.
Housman G, Ulitsky I. Methods for distinguishing between protein-coding and long noncoding RNAs and the elusive biological purpose of translation of long noncoding RNAs. Biochim Biophys Acta. 2016;1859:31–40.
Sleator RD. An overview of the current status of eukaryote gene prediction strategies. Gene. 2010;461:1–4.
Saghatelian A, Couso JP. Discovery and characterization of smORF-encoded bioactive polypeptides. Nat Chem Biol. 2015;11:909–16.
Couso JP. Finding smORFs: getting closer. Genome Biol. 2015;16:189.
Brar GA, Weissman JS. Ribosome profiling reveals the what, when, where and how of protein synthesis. Nat Rev Mol Cell Biol. 2015;16:651–64.
Hall A, Nobes CD. Rho GTPases: molecular switches that control the organization and dynamics of the actin cytoskeleton. Philos Trans R Soc Lond Ser B Biol Sci. 2000;355:965–70.
Ridley AJ, Schwartz MA, Burridge K, Firtel RA, Ginsberg MH, Borisy G, et al. Cell migration: integrating signals from front to back. Science. 2003;302:1704–9.
Lambrechts A, Van Troys M, Ampe C. The actin cytoskeleton in normal and pathological cell motility. Int J Biochem Cell Biol. 2004;36:1890–909.
Chhabra ES, Higgs HN. The many faces of actin: matching assembly factors with cellular structures. Nat Cell Biol. 2007;9:1110–21.
Campellone KG, Welch MD. A nucleator arms race: cellular control of actin assembly. Nat Rev Mol Cell Biol. 2010;11:237–51.
Gachet Y, Tournier S, Millar JB, Hyams JS. A MAP kinase-dependent actin checkpoint ensures proper spindle orientation in fission yeast. Nature. 2001;412:352–5.
Lee K, Song K. Actin dysfunction activates ERK1/2 and delays entry into mitosis in mammalian cells. Cell Cycle. 2007;6:1487–95.
Mullen PJ, Yu R, Longo J, Archer MC, Penn LZ. The interplay between cell signalling and the mevalonate pathway in cancer. Nat Rev Cancer. 2016;16:718–31.
Inder KL, Zheng YZ, Davis MJ, Moon H, Loo D, Nguyen H, et al. Expression of PTRF in PC-3 Cells modulates cholesterol dynamics and the actin cytoskeleton impacting secretion pathways. Mol Cell Proteom. 2012;11:M111 012245.
Miettinen TP, Bjorklund M. Mevalonate pathway regulates cell size homeostasis and proteostasis through autophagy. Cell Rep. 2015;13:2610–20.
Gutschner T, Hammerle M, Eissmann M, Hsu J, Kim Y, Hung G, et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013;73:1180–9.
Schmid I, Sakamoto KM (2001). Analysis of DNA content and green fluorescent protein expression. Curr Protoc Cytom. Chapter 7: Unit 7.16.
Acknowledgements
We thank Drs. K. Boulay and A. Roth for helpful discussions and suggestions and for critical reading of the manuscript, S. Wiemann for helpful discussions, L. Chatel-Chaix (University of Heidelberg) and R. Pepperkok (EMBL Heidelberg) for antibodies and suggestions, V. Benes, T. Ivacevic, and S. Schmidt (EMBL Heidelberg, Genome Core Facility) for support with microarray performance and the DKFZ Light Microscopy and Flow Cytometry Core Facilities and the ZMBH Flow Cytometry & FACS Core Facility (University of Heidelberg) for technical support. Research in the Diederichs lab is supported by the German Research Foundation (DFG Di 1421/7-1) and the RNA@DKFZ Cross Program Topic. This work is part of the Ph.D. thesis of M.P.-S.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
S.D. is a co-owner of siTOOLs Biotech GmbH, Martinsried, Germany. The remaining authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Polycarpou-Schwarz, M., Groß, M., Mestdagh, P. et al. The cancer-associated microprotein CASIMO1 controls cell proliferation and interacts with squalene epoxidase modulating lipid droplet formation. Oncogene 37, 4750–4768 (2018). https://doi.org/10.1038/s41388-018-0281-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0281-5
This article is cited by
-
Micropeptides: potential treatment strategies for cancer
Cancer Cell International (2024)
-
The role of DGAT1 and DGAT2 in regulating tumor cell growth and their potential clinical implications
Journal of Translational Medicine (2024)
-
p53-regulated lncRNAs in cancers: from proliferation and metastasis to therapy
Cancer Gene Therapy (2023)
-
Micropeptide MIAC inhibits the tumor progression by interacting with AQP2 and inhibiting EREG/EGFR signaling in renal cell carcinoma
Molecular Cancer (2022)
-
Fatty acids and nutritional components of the seed oil from Wangmo red ball Camellia oleifera grown in the low-heat valley of Guizhou, China
Scientific Reports (2022)