Abstract
Isocitrate dehydrogenase 1 and 2 (IDH1/2) are key enzymes in cellular metabolism, epigenetic regulation, redox states, and DNA repair. IDH1/2 mutations are causal in the development and/or progression of various types of cancer due to supraphysiological production of d-2-hydroxyglutarate. In various tumor types, IDH1/2-mutated cancers predict for improved responses to treatment with irradiation or chemotherapy. The present review discusses the molecular basis of the sensitivity of IDH1/2-mutated cancers with respect to the function of mutated IDH1/2 in cellular processes and their interactions with novel IDH1/2-mutant inhibitors. Finally, lessons learned from IDH1/2 mutations for future clinical applications in IDH1/2 wild-type cancers are discussed.
Similar content being viewed by others
Introduction
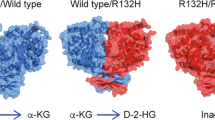
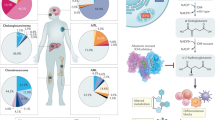
Isocitrate dehydrogenase 1 and 2 (IDH1/2) are key enzymes that function at a crossroads of cellular metabolism, epigenetic regulation, redox states, and DNA repair. Mutations in the genes encoding for these enzymes occur in various types of malignancies, including >80% of low-grade gliomas and secondary glioblastomas [1,2,3], ~60% of chondrosarcomas [4], ~20% of intrahepatic cholangiocarcinomas (ICC) [5] and ~10% of acute myeloid leukemias (AML) [6,7,8]. These mutations occur in a hotspot fashion in the catalytically active sites of these enzymes and the main driver of oncogenesis is the neomorphic production of d-2-hydroxyglutarate (d-2HG; Fig. 1) [9]. The resulting d-2HG accumulation competitively inhibits α-ketoglutarate (αKG)-dependent enzymes, causing cellular alterations in the above-mentioned plethora of cellular metabolism, epigenetic regulation, redox states, and DNA repair, all of which may contribute to carcinogenesis which has been extensively reviewed elsewhere [10,11,12]. As the neomorphic production of d-2HG is essentially a gain of function that is exclusive to mutant IDH1/2 enzymes, it was quickly realized that these frequently-occurring genetic alterations were promising targets for personalized anti-cancer therapy with small-molecule inhibitors [13]. Within 5 years after the initial development of these compounds, the IDH2-mutant inhibitor enasidenib was approved by the FDA as a first-in-class inhibitor for the treatment of relapsed or refractory IDH2-mutated AML [14, 15].
Effects of IDH1/2 mutations and d-2HG accumulation on cellular metabolism, redox states, and DNA damage repair. ALKBH alkylation repair homolog, ATM ataxia-telangiectasia mutated, ATP5 adenosine triphosphate synthase, CoA coenzyme A, COX cytochrome c oxidase, d-2HG d-2-hydroxyglutarate, ETC electron transport chain, FOXO forkhead box proteins, HuR human antigen R, IDH isocitrate dehydrogenase, KDM lysine histone demethylase, NAD(P) nicotinamide dinucleotide (phosphate), NAD(P)H nicotinamide dinucleotide (phosphate), reduced, NAM nicotinamide, NAMPT nicotinamide phosphoribosyltransferase, NMN nicotinamide mononucleotide, NRF2 nuclear factor (erythroid-derived 2)-like, ROS reactive oxygen species
Soon after their discovery, it was appreciated that IDH1/2 mutations were associated with a relatively prolonged patient survival for glioma [3] and glioblastoma [2] but not for AML [16, 8] or chondrosarcoma [4]. For ICC, some studies reported that IDH1/2 mutations were independent predictive factors for prolonged progression-free and overall survival [17], whereas other studies reported no difference between the survival of IDH1/2-mutated vs. IDH1/2 wild-type ICC [18, 19] and one study (reporting only six IDH1/2-mutated cases) even found a worse prognosis of IDH1/2-mutated ICC compared to wild-type counterparts [20]. The assumption that IDH1/2 mutations are causal for the improved clinical outcome in glioma was supported by clinical evidence, as IDH1/2 mutations predicted for improved tumor responses to chemotherapy and/or irradiation in clinical trials [21, 22] and retrospective analyses [23,24,25,26,27]. Furthermore, cancer cells are sensitized to radiation and chemotherapy by the introduction of mutant IDH1/2 or by silencing of wild-type IDH1/2 (Table 1). IDH1/2 mutations or the absence of IDH1/2 wild-type enzymes create downstream vulnerabilities in cancer that can be therapeutically targeted with small-molecule inhibitors, such as poly(ADP-ribose) polymerase (PARP) inhibitors, nicotinamide phosphoribosyltransferase (NAMPT) inhibitors, BCL-2 inhibitors and biguanides. A better understanding of the mechanisms of these vulnerabilities may aid to improve personalized therapy for patients with or without IDH1/2-mutated cancers and is the subject of this review.
IDH1/2 enzymes in metabolism
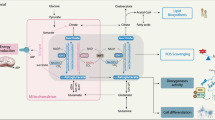
IDH1 and IDH2 catalyze the reversible oxidative decarboxylation of isocitrate to αKG in the cytoplasm and mitochondria, respectively, with concomitant reduction of NADP+ to NADPH (Fig. 2). Although IDH1/2 do not generate NADH, the canonical product of the TCA cycle, IDH1/2 perform the same isocitrate-to-αKG conversion as the traditional TCA cycle enzyme IDH3. IDH1/2 mutations are neomorphic [9], but also confer a loss of function of wild-type IDH1/2 kinetics [28, 29] and redirect carbon metabolites away from the TCA cycle and oxidative phosphorylation towards d-2HG production. This is evidenced by decreased expression of TCA cycle enzymes downstream of IDH [30] and decreased oxidative metabolism in Seahorse metabolic assays using IDH1/2-mutated cancer cells [30, 31]. IDH1-mutant-induced mitochondrial dysfunction is also compensated by an increase in the number of mitochondria in IDH1/2-mutated cells [32–35]. As a result, IDH1/2-mutated cancer cells are vulnerable to inhibition of the residual oxidative metabolism with inhibitors of the electron transport chain (ETC), such as the biguanides metformin and phenformin that inhibit NADH dehydrogenase (complex I) of the ETC [29, 36, 37]. Metformin is currently investigated for safety and efficacy in a clinical trial of patients with IDH1/2-mutated solid tumors [38].
A consequence of rewired metabolism in IDH1/2-mutated cells is the dependence on the glutaminolysis pathway, which provides anaplerosis to the TCA cycle at the level of αKG. IDH1/2-mutated cells need αKG to produce d-2HG but at the same time they restrict αKG production by impairing glycolytic influx and TCA cycle metabolism [30]. αKG production from glutamine provides an alternative source of fuel to satisfy IDH1/2-mutated cells in their d-2HG production, but also render these cells vulnerable to pharmacological inhibition of glutaminolysis with the use of aminooxyacetic acid, BPTES, zaprinast, or chloroquine [35, 39–42]. It has been hypothesized that IDH1-mutated glioma depend on glutamate rather than glutamine for TCA cycle anaplerosis [43, 44].
IDH1/2 mutations or IDH1/2 knockdown disable the oxidative decarboxylation reaction that converts αKG to isocitrate [37]. This reaction occurs predominantly in hypoxia, when glycolytic influx of pyruvate in the TCA cycle is compromised and cells use the reverse IDH1/2 reaction to generate citrate and acetyl-CoA from glutamine and glutamate in order to preserve the capacity to synthesize lipids in hypoxic contexts [37,45–47]. Knockdown of IDH1 sensitized glioma-initiating cells with EGFR amplifications to treatment with erlotinib through decreased fatty acid and cholesterol biosynthesis. This vulnerable phenotype was rescued by treatment with cell membrane-permeable αKG or the fatty acid palmitate plus the cholesterol precursor mevalonate [48].
Finally, the metabolic rewiring by IDH1/2 mutations renders cells vulnerable to inhibitors that exploit a synthetic lethality of this IDH1/2-mutant metabolism. One is that d-2HG inhibits cytochrome c oxidase (complex IV of the ETC) and prevents cytochrome c release into the mitochondrial matrix. Besides restricting oxidative mitochondrial metabolism, cytochrome c release puts IDH1/2-mutated cells on the brink of apoptosis through BAX/BAK-mediated permeabilization of the outer mitochondrial membrane. Under steady-state conditions, this is prevented by BCL-2 binding to the proapoptotic proteins BAX/BAK, but disruption of this binding by the BH3 mimetic ABT-199 (venetoclax, FDA-approved for the treatment of chronic lymphocytic leukemia) results in apoptosis of IDH1/2-mutated cells while IDH1/2 wild-type cells are relatively insensitive [49]. In a clinical study among AML patients, those with IDH1/2 mutations had a higher response rate (36%) to venetoclax than those with IDH1/2 wild type (9%) [50]. A similar effect was observed in glioblastoma models, although BCL-xL and not BCL-2 seemed to be the primary target of the BH mimetic ABT-263 in these glioblastoma models [51]. Another example is that d-2HG downregulates nicotinate phosphoribosyltransferase (NAPRT1), an enzyme in the NAD+ salvage pathway, which causes sensitivity towards depletion of NAD+ by pharmacological inhibition of NAMPT with the preclinical compounds FK866 and GMX1778 leading to AMP kinase-initiated autophagy and cell death in IDH1-mutated cancer cells [52]. Finally, IDH1/2-mutated ICC cells are dependent on the kinase SRC for mTOR-mediated proliferation and survival and these cells are highly sensitive to treatment with the multikinase inhibitor dasatinib, which also has affinity for SRC [53]. These synthetic lethalities are probably the result of complex crosstalk between IDH1/2 mutations and tissue-specific intracellular processes, because among IDH1/2-mutated models, AML and glioblastoma but not ICC was sensitized to BCL-2/BCL-xL inhibition [49, 51, 53], glioma but not ICC was sensitized to NAMPT inhibition [52, 53], and ICC but not chondrosarcoma or lung cancer was sensitized to dasatinib [53].
IDH1/2 enzymes in redox states
Since the reductive carboxylation of isocitrate to αKG by IDH1/2 is redundant besides the function of IDH3 in the TCA cycle, IDH1 and IDH2 presumably arose in evolution for the purpose of NADPH production in the cytoplasm and mitochondria, respectively [12]. NADPH is an important source of cellular reducing power and is required to recharge, activate, or generate reduced glutathione (GSH) [54], thioredoxin [55], catalase tetramers [56], and cytochrome P450 [54, 57], all of which are involved in the reduction of reactive oxygen species (ROS). Moreover, NADPH is necessary for the synthesis of deoxynucleotides and thus for DNA damage repair [58]. IDH1/2 mutations cause loss of IDH1/2 wild-type functions [28, 29], resulting in decreased NADPH and GSH levels and increased ROS levels, both in steady-state conditions and after induction of ROS production [29, 41, 48, 52, 59, 60]. In addition, d-2HG accumulation induces oxidative stress independently of IDH1/2 mutations [32–34]. In AML, the IDH1R132H gene expression signature was enriched in genes that result in a cellular phenotype that is responsive to treatment with small molecules that target ROS and NADP+/NADPH signaling and metabolism [61]. Oxidative stress is possibly induced via inhibition of wild-type IDH1/2 activity due to αKG mimicry of d-2HG, which results in a pseudo-product inhibition of wild-type IDH1/2 [55] or via increased mitochondrial transmembrane proton leakage due to cytochrome c retention in the mitochondrial intermembrane space as described above [49]. Further evidence of increased ROS levels as mediator of increased therapy sensitivity of IDH1/2-mutated cells is shown by the almost complete reversal of this sensitivity in various cell models by the antioxidant and GSH surrogate N-acetyl-cysteine. This reversal of sensitivity has been shown in the presence of carmustine (BCNU) [41], irradiation [29, 62, 63], cisplatin, temozolomide [64], and erlotinib in glioma-initiating cells with EGFR amplification, where increased ROS levels increase erlotinib-induced apoptosis after IDH1 knockdown [48]. Some in vitro studies showed depleted GSH levels and increased ROS levels in cancer cells with IDH1 mutations [29, 64–66] but IDH1/2 mutations did not alter ROS levels in brains and hematopoietic cells of IDH1R132H knock-in mice [67, 68] or immortalized human astrocytes [69] in other studies. However, these studies only interrogated steady-state conditions and used a ROS marker (CM-H2DCFDA) that is insensitive to H2O2 [70], the oxidant that is most probably elevated in cells that are depleted of NADPH and have limited peroxidase and peroxiredoxin activity.
IDH1/2 are the most important NADPH producers in most human organs, including the brain [28, 71], and IDH1 mutations lead to depleted NADPH levels in colorectal cancer cells [29] and glioma cells [64, 66]. In myeloid cells, glucose-6-phosphate dehydrogenase (G6PDH) of the pentose phosphate pathway is the major NADPH provider [12]. IDH1 is the highest upregulated NADPH-producing enzyme when glioblastoma and normal brain tissue are compared and IDH1 mRNA and protein expression is upregulated following radiation, suggesting a role for IDH1 in cellular responses to radiation [48, 63], possibly by induction of IDH1 expression via Forkhead box O (FOXO) transcription factors [72]. In pancreatic cancer cells, IDH1 expression is induced by HuR (ELAVL1) after treatment with gemcitabine and the HuR–IDH1 regulatory axis is essential for adaptive pancreatic cancer cell survival under acute stress [73]. Furthermore, the introduction of a mutant IDH1/2 protein radiosensitizes glioblastoma cells and other cancerous and noncancerous cells [29, 74, 75]. Radiosensitization also occurs after knockdown of IDH1 [63, 77, 78], whereas overexpression of IDH1 protects cancer cells against chemotherapy [73, 79]. These latter two findings are important, because it isolates the role of IDH1 loss-of-function in cellular radiosensitization and rules out d-2HG from being solely responsible for this phenomenon. Radiosensitivity of IDH1−/− cells was related to increased cellular senescence due to depletion of antioxidants and deoxynucleotides in IDH1−/− cells following irradiation, whereas apoptosis, necrosis, autophagy, unrepaired DNA double-strand breaks, and homologous recombination repair remained unchanged [63]. This partly corroborates and partly contrasts the situation in IDH1/2-mutated cells, where radiosensitivity is caused by depletion of antioxidants [29], and by reduced DNA damage responses and double-strand break repair [29, 75]. Since d-2HG accumulation is strongly linked to perturbed DNA damage repair (see below), it seems plausible that ROS-associated therapy sensitivity of IDH1/2-mutated cells mainly occurs via increased cellular senescence due to depleted levels of antioxidants and deoxynucleotides, whereas d-2HG-associated therapy sensitivity of IDH1/2-mutated cells is predominantly associated with inhibition of DNA damage response proteins.
Finally, IDH1 mutations are implicated in the downregulation and aberrant subcellular localization of nuclear factor-erythroid 2-related factor 2 (NRF2) and NAD(P)H quinine oxidoreductase 1 (NQO1), which are important cellular defense proteins against oxidative stress. Their downregulation and dysfunction is associated with increased sensitivity to chemotherapy with temozolomide, but a regulatory relationship between IDH1 and NRF2 remains unclear.
IDH1/2 enzymes in DNA repair
Besides the indirect effects of IDH1/2 mutations on DNA repair via redox state perturbations, IDH1/2 and DNA repair are interwoven via direct inhibition by d-2HG of αKG-dependent dioxygenases involved in DNA repair. For example, d-2HG inhibits the DNA repair enzyme alkB homolog (ALKBH) [80, 81] and the DNA damage response proteins lysine-specific demethylase 4A/B (KDM4A/B) [75, 82, 83] and suppresses the expression of the DNA damage response protein ATM [75]. These findings are linked with increased DNA damage in IDH1/2-mutated cells as compared to IDH1/2 wild-type cells, either in steady-state conditions or after treatment with cytotoxic or targeted agents [29, 31, 74, 75]. Perturbed steady-state DNA repair may contribute to oncogenesis of IDH1/2-mutated cancers, but a perturbed DNA damage response is even more likely to be related to the increased susceptibility of IDH1/2-mutated cancers to DNA damage-inducing cytotoxic agents.
Inhibition of ALKBH by d-2HG results in sensitization of IDH1/2-mutated cancer cells to alkylating agents such as busulfan and CCNU [81]. These findings provide a molecular basis for the sensitivity of IDH1/2-mutated glioma towards a regimen of radiotherapy in the presence or absence of procarbazine, CCNU, and vincristine (PCV) [21, 22], of which the first two are DNA alkylators. Moreover, temozolomide is another DNA-alkylating agent that is the standard of care first-line therapy for glioblastoma and IDH1/2 mutations predict for glioblastoma responses to temozolomide [26].
IDH1/2-mutated cancers are known to confer defects in homologous recombination, whereas the other major DNA double-strand break repair pathway, non-homologous end-joining, remains intact [75]. The result is increased levels of DNA damage [29, 31, 75], which prompted researchers to perform a focused high-throughput screen of DNA repair inhibitors [75]. This resulted in the observation that IDH1/2-mutated cancers are sensitive to PARP inhibitors in vivo and this sensitivity for PARP inhibitors synergizes with temozolomide or cisplatin treatment in vitro, but there is disagreement on the underlying mechanisms [31, 75, 76]. One study implicated NAD+ deficiency in IDH1/2-mutated cells in PARP dysfunction, because NAD+ is an essential cofactor for PARP-mediated single-strand DNA repair [31]. However, this conclusion was based on associative evidence obtained from findings that the PARP DNA repair machinery was intact in IDH1/2-mutated cells whereas NAD+ levels were more depleted in IDH1/2-mutated cells than in IDH1/2 wild-type cells after DNA-damaging temozolomide treatment in vitro. However, mechanistic experiments to pinpoint NAD+ levels as the critical factor for IDH1/2 mutation-mediated PARP dysfunction were not performed [31]. Another study showed that NAD+ levels have no role in IDH1/2 mutation-induced PARP sensitivity but instead convincingly showed that d-2HG inhibits the αKG-dependent dioxygenases KDM4A/B. Inhibition of KDM4A/B induces a homologous recombination defect that creates a “BRCAness” phenotype in IDH1/2-mutant cells, which results in PARP inhibitor sensitivity in vivo [75]. Of note, IDH1/2 mutations and d-2HG accumulation were already linked to KDM4 inhibition several years earlier but thus far this observation was only linked to global histone hypermethylation and not to decreased DNA damage responses [84]. The mechanism behind the latter phenomenon is that histone methylation, such as demethylation of histone H3 lysine 20 (H3K20) and trimethylation of H3K9, is a barrier to DNA double-strand break repair that can be relieved by KDM4A [82] and KDM4B [83], respectively. Moreover, these demethylases cooperate with or sometimes orchestrate the activity of canonical DNA damage response proteins, such as 53BP1 in the case of KDM4A [82] and PARP1 in the case of KDM4B [83]. TET2 is a major downstream target of d-2HG accumulation and is considered to be a major mediator of IDH1/2-mutant-mediated oncogenesis [11, 12, 85]. However, it is unlikely that TET2 inhibition contributes to sensitization of IDH1/2-mutant cells to PARP inhibitors, because restoration of TET2 function sensitizes rather than protects TET2 haploinsufficient AML cells to PARP inhibitors [86]. This is an intriguing finding that also questions how IDH1/2-mutant inhibitors reverse the PARP inhibitor sensitization of IDH1/2-mutated cells, as IDH1/2-mutant inhibitors reduce d-2HG levels and should restore TET2 function [87]. One study speculated that perturbed DNA damage repair and increased temozolomide sensitivity of IDH1/2-mutated cells are caused by impaired oxidative metabolism because administration of αKG protected IDH1/2-mutated cells against temozolomide treatment. However, it now seems more plausible that αKG administration reduces the competitive inhibition of d-2HG of ALKBH and/or KDM4A/B and restores the activity of these DNA damage repair enzymes [31]. It was also demonstrated that temozolomide treatment made a bigger dent in NAD+ levels in IDH1/2-mutated than in IDH1/2 wild-type cells which was driven by NAD+ consumption by PARP [88]. As a result, combined treatment with temozolomide and NAMPT inhibitors had a synergistic effect in IDH1/2-mutated cancers in vivo and may represent a promising therapeutic avenue for IDH1/2-mutated cancer patients.
Whole-proteome analyses of Idh1 wild-type and Idh1-mutated murine hematopoietic stem cells revealed that the latter had lower levels of the (phosphorylated) DNA damage response proteins phospho-ATM, phospho-CHEK2, and γH2AX. Mutant IDH1 downregulates the DNA damage response protein ATM via an epigenetic mechanism that involves chromatin modifications via histone lysine demethylation. Mechanistic experiments involving inhibitors of epigenetic modifiers affecting H3K9 and H3K27 showed a direct link between repressive trimethylation at these histone marks and downregulation of ATM. The authors speculated that KDM4 inhibition by d-2HG is responsible for ATM suppression and excluded TET2 as a mechanistic link between d-2HG accumulation and epigenetic suppression of ATM expression, because Tet2−/− mice had normal ATM levels. At the therapeutic level, reduced ATM activity was associated with increased sensitivity to irradiation and the DNA-damaging chemotherapeutic agent daunorubicin [74]. Reduced ATM expression has also been observed in human primary IDH1/2-mutated AML cells as compared to their wild-type counterparts and was rescued by using an IDH1/2-mutant inhibitor. In these human cell models, IDH1/2 mutations and ATM suppression caused sensitivity to irradiation and daunorubicin [76].
IDH1/2 enzymes in epigenetic regulation
αKG and d-2HG, the products of wild-type and mutant IDH1/2, respectively, are closely involved in epigenetic regulation of gene expression. Via this mechanism, IDH1/2 form a link between metabolism and epigenetics, as has been extensively reviewed elsewhere [89]. d-2HG is an inhibitor of various αKG-dependent dioxygenases necessary for DNA and histone demethylation, such as TET2 and Jumonji-domain containing lysine histone demethylases [10, 11]. The downstream epigenetic effects of IDH1/2 mutations have been predominantly investigated in the context of IDH1/2-mutant-mediated oncogenesis but not as targets for personalized therapy. However, IDH1/2-mutant-induced epigenetic modifications do have potential as therapeutic targets. For example, IDH1 mutations induce a gene expression signature that render AML cells vulnerable to all-trans retinoic acid (ATRA/tretinoin), which promotes the expression of genes that induce differentiation of IDH1-mutated AML cells [61]. Furthermore, the epigenetic effects of IDH1/2 mutations may underlie many of the synthetic lethalities already described in this review, including the downregulation of NAPRT1 to sensitize cells to NAMPT inhibition [52], the downregulation of NRF2 and ATM to sensitize cells to chemo/radiotherapy [60, 74], the downregulation of BCAT1 to increase the dependency on glutaminolysis [90], and the induction of homologous recombination defects to sensitize cells to PARP inhibitors, as described above in more detail [75].
IDH1/2-mutant inhibitors and therapy responses
IDH1/2 mutations are inaugural or at least early events in the formation of glioma [91], chondrosarcoma [4], ICC [92], and AML [93] (although in that case the data are conflicting [8]) and are thus present in the large majority, if not all, cancer cells. This makes IDH1/2 mutations attractive therapeutic targets, because such tumor homogeneity decreases the risk of therapy resistance since targeting IDH1/2 mutations affects all cancer cells. The appreciation of the role of IDH1/2 mutations in oncogenesis and their early occurrence prompted the development of IDH1/2-mutant inhibitors [13]. Enasidenib (AG-221/CC-90007) is now registered for the treatment of refractory/relapsed IDH2-mutated AML and the IDH1-mutant inhibitor ivosidenib (AG-120) is in clinical trials. Enasidenib suppresses d-2HG production, reverses epigenetic dysregulation, and induces cellular differentiation in IDH2-mutated AML, where it achieved an overall response rate of 40% and a complete remission rate of 19% [15, 94, 95]. MDS patients with ancestral IDH1/2 mutations had a worse survival than patients with subclonal IDH1/2 mutations [8], which may suggest that patients with ancestral IDH1/2 mutations are the best candidates for therapy with IDH1/2-mutant inhibitors. However, an association between the mutant IDH2 variant allelic frequency and clinical response to enasidenib was not observed in AML patients [94].
These results with enasidenib monotherapy are promising for the difficult-to-treat population of patients with refractory/relapsed AML, but even before the first clinical trials with IDH1/2-mutant inhibitors started it was doubted whether combination regimens of IDH1/2-mutant inhibitors and conventional chemotherapy or targeted DNA damage-inducing agents would be safe and efficacious [12]. In the context of our increasing understanding of the therapy response-modulating effects of IDH1/2 mutations, which almost exclusively point at an increased sensitization to cytotoxic agents of most types of cancer, it is plausible that concomitant administration of such cytotoxic agents and IDH1/2-mutant inhibitors counteract each other. For example, IDH1/2-mutant inhibitors protect IDH1/2-mutated glioma, AML, chondrosarcoma, and colorectal carcinoma cells against irradiation, daunorubicin, and PARP inhibitors [29, 75, 76]. In all cases, the mechanism of therapy protection by IDH1/2-mutant inhibitors was based on the reversal of the mechanism that rendered IDH1/2-mutated cancer cells sensitive to the therapeutic agent. For example, pretreatment with the IDH1-mutant inhibitor AGI-5198 (the preclinical version of ivosidenib/AG-120) decreased d-2HG levels, restored NADPH production, and decreased ROS levels and these phenomena collectively or ultimately resulted in less IDH1/2-mutated cell death after irradiation [29]. In the case of protection of IDH1-mutated cells against pharmacological PARP inhibition, pretreatment with AGI-5198 decreased d-2HG levels and the number of DNA double-strand breaks and reverted the PARP inhibitor sensitivity of IDH1-mutated cells to levels observed in IDH1 wild-type cells [75]. In both instances, the therapy-protective effects of AGI-5198 was overcome by administration of exogenous d-2HG to increase d-2HG levels independently of the inhibited mutant IDH1 enzyme [29, 75].
Concluding remarks and future perspectives
IDH1/2 mutations are attractive therapeutic targets for various reasons, but most prominently because they are early events in oncogenesis. As a consequence, this tumor homogeneity ensures that the chance of relapse of IDH1/2-mutated cancers is theoretically small after a complete response/remission obtained by the application of targeted therapeutic agents. Another prediction on the basis of the early occurrence of IDH1/2 mutations in oncogenesis and the plethora of downstream cellular effects of IDH1/2 mutations is the profoundly altered tumor biology during oncogenesis. Consequently, many research efforts have been devoted to the discovery of specific vulnerabilities, especially in the domains of metabolism and DNA damage responses. This has resulted in a better understanding of the sensitization of cancer cells by IDH1/2 mutations to conventional chemo/radiotherapy, but also of the susceptibility to targeted agents that have maximal efficacy and minimal side effects. We expect that pharmacological inhibition of BCL-2, NAMPT, and PARP are the most promising therapeutic avenues in this category [49, 52, 75], especially in combination with each other or with chemo/radiotherapy. Olaparib, rucaparib, and niraparib are PARP inhibitors [95], and venetoclax is a BCL-2 inhibitor [97] that are all FDA-approved for the treatment of other types of cancer, which may ease future clinical trials with these drugs for the treatment of IDH1/2-mutated cancers. Because of the inherent counteracting nature of IDH1/2-mutant inhibitors on the one hand and personalized targeted therapy of IDH1/2-mutated cancers on the other hand, it is doubtful whether there will ever be clinically safe and effective combinations of IDH1/2-mutant inhibitors and cytotoxic agents to which a particular IDH1/2-mutated cancer is sensitized [29, 75]. The future looks bright for preclinical and clinical research on IDH1/2-mutated cancers and its advances may eventually trickle down to the much larger populations of patients with IDH1/2 wild-type cancers. It is becoming increasingly clear that inhibition of wild-type IDH1/2 to mimic therapy responses of IDH1/2-mutant cancer biology may have therapeutic potential. This was recently demonstrated in three preclinical models [48, 63, 73]. Another example of the trickle-down effect of the increased research on IDH1/2 mutations is that reasonably potent pharmacological inhibitors of wild-type IDH1 such as GSK864 (IC50: ~470 nM) have become available whereas we previously only had RNA interference, genetic modification, or the unspecific and impotent oxalomalate [78] to inhibit wild-type IDH1. GSK864 was originally developed as IDH1-mutant inhibitor but also showed activity against wild-type IDH1 [98]. In this way, we learn from lessons from nature so that ultimately IDH1/2 wild-type patients may benefit as well from our understanding of the increased therapy sensitivity of IDH1/2-mutated cancers.
Change history
23 August 2018
The originally published version of this article contained an error in Figure 2. On the IDH1/2 mutated arrow, it incorrectly read ‘NADP+ ->NADPH’ instead of the correct ‘NADPH ->NADP+‘. This has now been corrected in the PDF and HTML versions of the article. The authors wish to apologize for any inconvenience caused.
References
Molenaar RJ, Verbaan D, Lamba S, Zanon C, Jeuken JW, Boots-Sprenger SH, et al. The combination of IDH1 mutations and MGMT methylation status predicts survival in glioblastoma better than either IDH1 or MGMT alone. Neuro Oncol. 2014;16:1263–73.
Parsons DW, Jones S, Zhang X, Lin JC, Leary RJ, Angenendt P, et al. An integrated genomic analysis of human glioblastoma multiforme. Science. 2008;321:1807–12.
Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W, et al. IDH1 and IDH2 mutations in gliomas. N Engl J Med. 2009;360:765–73.
Pansuriya TC, van Eijk R, d’Adamo P, van Ruler MA, Kuijjer ML, Oosting J, et al. Somatic mosaic IDH1 and IDH2 mutations are associated with enchondroma and spindle cell hemangioma in Ollier disease and Maffucci syndrome. Nat Genet. 2011;43:1256–61.
Borger DR, Tanabe KK, Fan KC, Lopez HU, Fantin VR, Straley KS, et al. Frequent mutation of isocitrate dehydrogenase (IDH)1 and IDH2 in cholangiocarcinoma identified through broad-based tumor genotyping. Oncologist. 2012;17:72–9.
Figueroa ME, Abdel-Wahab O, Lu C, Ward PS, Patel J, Shih A, et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell. 2010;18:553–67.
Mardis ER, Ding L, Dooling DJ, Larson DE, McLellan MD, Chen K, et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N Engl J Med. 2009;361:1058–66.
Molenaar RJ, Thota S, Nagata Y, Patel B, Clemente M, Przychodzen B, et al. Clinical and biological implications of ancestral and non-ancestral IDH1 and IDH2 mutations in myeloid neoplasms. Leukemia. 2015;29:2134–42.
Dang L, White DW, Gross S, Bennett BD, Bittinger MA, Driggers EM, et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature. 2009;462:739–44.
LM G, Boulay K, Topisirovic I, Huot ME, Mallette FA. Oncogenic activities of IDH1/2 mutations: from epigenetics to cellular signaling. Trends Cell Biol. 2017;27:738–52.
Losman JA, Kaelin WG Jr. What a difference a hydroxyl makes: mutant IDH, (R)-2-hydroxyglutarate, and cancer. Genes Dev. 2013;27:836–52.
Molenaar RJ, Radivoyevitch T, Maciejewski JP, van Noorden CJ, Bleeker FE. The driver and passenger effects of isocitrate dehydrogenase 1 and 2 mutations in oncogenesis and survival prolongation. Biochim Biophys Acta. 2014;1846:326–41.
Popovici-Muller J, Saunders JO, Salituro FG, Travins JM, Yan S, Zhao F, et al. Discovery of the first potent inhibitors of mutant IDH1 that lower tumor 2-HG in vivo. ACS Med Chem Lett. 2012;3:850–5.
Dang L, Su SM. Isocitrate dehydrogenase mutation and (R)-2-hydroxyglutarate: from basic discovery to therapeutics development. Annu Rev Biochem. 2017;86:305–31.
Stein EM, DiNardo CD, Pollyea DA, Fathi AT, Roboz GJ, Altman JK, et al. Enasidenib in mutant IDH2 relapsed or refractory acute myeloid leukemia. Blood. 2017;130:722–31.
Xu Q, Li Y, Lv N, Jing Y, Xu Y, Li Y, et al Correlation between isocitrate dehydrogenase gene aberrations and prognosis of patients with acute myeloid leukemia: a systematic review and meta-analysis. Clin Cancer Res. 2017;23:4511–22.
Wang P, Dong Q, Zhang C, Kuan PF, Liu Y, Jeck WR, et al. Mutations in isocitrate dehydrogenase 1 and 2 occur frequently in intrahepatic cholangiocarcinomas and share hypermethylation targets with glioblastomas. Oncogene. 2013;32:3091–100.
Goyal L, Govindan A, Sheth RA, Nardi V, Blaszkowsky LS, Faris JE, et al. Prognosis and clinicopathologic features of patients with advanced stage isocitrate dehydrogenase (IDH) mutant and IDH wild-type intrahepatic cholangiocarcinoma. Oncologist. 2015;20:1019–27.
Zhu AX, Borger DR, Kim Y, Cosgrove D, Ejaz A, Alexandrescu S, et al. Genomic profiling of intrahepatic cholangiocarcinoma: refining prognosis and identifying therapeutic targets. Ann Surg Oncol. 2014;21:3827–34.
Jiao Y, Pawlik TM, Anders RA, Selaru FM, Streppel MM, Lucas DJ, et al. Exome sequencing identifies frequent inactivating mutations in BAP1, ARID1A and PBRM1 in intrahepatic cholangiocarcinomas. Nat Genet. 2013;45:1470–3.
Buckner JC, Shaw EG, Pugh SL, Chakravarti A, Gilbert MR, Barger GR, et al. Radiation plus Procarbazine, CCNU, and Vincristine in low-grade glioma. N Engl J Med. 2016;374:1344–55.
Cairncross JG, Wang M, Jenkins RB, Shaw EG, Giannini C, Brachman DG, et al. Benefit from procarbazine, lomustine, and vincristine in oligodendroglial tumors is associated with mutation of IDH. J Clin Oncol. 2014;32:783–90.
Hartmann C, Hentschel B, Tatagiba M, Schramm J, Schnell O, Seidel C, et al. Molecular markers in low-grade gliomas: predictive or prognostic? Clin Cancer Res. 2011;17:4588–99.
Houillier C, Wang X, Kaloshi G, Mokhtari K, Guillevin R, Laffaire J, et al. IDH1 or IDH2 mutations predict longer survival and response to temozolomide in low-grade gliomas. Neurology. 2010;75:1560–6.
Okita Y, Narita Y, Miyakita Y, Ohno M, Matsushita Y, Fukushima S, et al. IDH1/2 mutation is a prognostic marker for survival and predicts response to chemotherapy for grade II gliomas concomitantly treated with radiation therapy. Int J Oncol. 2012;41:1325–36.
SongTao Q, Lei Y, Si G, YanQing D, HuiXia H, XueLin Z, et al. IDH mutations predict longer survival and response to temozolomide in secondary glioblastoma. Cancer Sci. 2012;103:269–73.
Tran AN, Lai A, Li S, Pope WB, Teixeira S, Harris RJ, et al. Increased sensitivity to radiochemotherapy in IDH1 mutant glioblastoma as demonstrated by serial quantitative MR volumetry. Neuro Oncol. 2014;16:414–20.
Bleeker FE, Atai NA, Lamba S, Jonker A, Rijkeboer D, Bosch KS, et al. The prognostic IDH1(R132) mutation is associated with reduced NADP + -dependent IDH activity in glioblastoma. Acta Neuropathol. 2010;119:487–94.
Molenaar RJ, Botman D, Smits MA, Hira VV, van Lith SA, Stap J, et al. Radioprotection of IDH1-mutated cancer cells by the IDH1-mutant inhibitor AGI-5198. Cancer Res. 2015;75:4790–802.
Khurshed M, Molenaar RJ, Lenting K, Leenders WP, van Noorden CJF. In silico gene expression analysis reveals glycolysis and acetate anaplerosis in IDH1 wild-type glioma and lactate and glutamate anaplerosis in IDH1-mutated glioma. Oncotarget. 2017;8:49165–77.
Lu Y, Kwintkiewicz J, Liu Y, Tech K, Frady LN, Su YT, et al. Chemosensitivity of IDH1-mutated gliomas due to an impairment in PARP1-mediated DNA repair. Cancer Res. 2017;77:1709–18.
Gilbert MR, Liu Y, Neltner J, Pu H, Morris A, Sunkara M, et al. Autophagy and oxidative stress in gliomas with IDH1 mutations. Acta Neuropathol. 2014;127:221–33.
Kolker S, Pawlak V, Ahlemeyer B, Okun JG, Horster F, Mayatepek E, et al. NMDA receptor activation and respiratory chain complex V inhibition contribute to neurodegeneration in d-2-hydroxyglutaric aciduria. Eur J Neurosci. 2002;16:21–8.
Latini A, Scussiato K, Rosa RB, Llesuy S, Bello-Klein A, Dutra-Filho CS, et al. D-2-hydroxyglutaric acid induces oxidative stress in cerebral cortex of young rats. Eur J Neurosci. 2003;17:2017–22.
Navis AC, Niclou SP, Fack F, Stieber D, van Lith S, Verrijp K, et al. Increased mitochondrial activity in a novel IDH1-R132H mutant human oligodendroglioma xenograft model: in situ detection of 2-HG and α-KG. Acta Neuropathol Comm. 2013;1:18.
Cuyas E, Fernandez-Arroyo S, Corominas-Faja B, Rodriguez-Gallego E, Bosch-Barrera J, Martin-Castillo B, et al. Oncometabolic mutation IDH1 R132H confers a metformin-hypersensitive phenotype. Oncotarget. 2015;6:12279–96.
Grassian AR, Parker SJ, Davidson SM, Divakaruni AS, Green CR, Zhang X, et al. IDH1 mutations alter citric acid cycle metabolism and increase dependence on oxidative mitochondrial metabolism. Cancer Res. 2014;74:3317–31.
Molenaar RJ, Coelen RJ, Khurshed M, Roos E, Caan MW, van Linde ME, et al. Study protocol of a phase IB/II clinical trial of metformin and chloroquine in patients with IDH1-mutated or IDH2-mutated solid tumours. BMJ Open. 2017;7:e014961.
Elhammali A, Ippolito JE, Collins L, Crowley J, Marasa J, Piwnica-Worms D. A high-throughput fluorimetric assay for 2-hydroxyglutarate identifies Zaprinast as a glutaminase inhibitor. Cancer Discov. 2014;4:828–39.
Emadi A, Jun SA, Tsukamoto T, Fathi AT, Minden MD, Dang CV. Inhibition of glutaminase selectively suppresses the growth of primary acute myeloid leukemia cells with IDH mutations. Exp Hematol. 2014;42:247–51.
Mohrenz IV, Antonietti P, Pusch S, Capper D, Balss J, Voigt S, et al. Isocitrate dehydrogenase 1 mutant R132H sensitizes glioma cells to BCNU-induced oxidative stress and cell death. Apoptosis. 2013;18:1416–25.
Seltzer MJ, Bennett BD, Joshi AD, Gao P, Thomas AG, Ferraris DV, et al. Inhibition of glutaminase preferentially slows growth of glioma cells with mutant IDH1. Cancer Res. 2010;70:8981–7.
van Lith SA, Molenaar R, van Noorden CJ, Leenders WP. Tumor cells in search for glutamate: an alternative explanation for increased invasiveness of IDH1 mutant gliomas. Neuro Oncol. 2014;16:1669–70.
van Lith SA, Navis AC, Verrijp K, Niclou SP, Bjerkvig R, Wesseling P, et al. Glutamate as chemotactic fuel for diffuse glioma cells: are they glutamate suckers? Biochim Biophys Acta. 2014;1846:66–74.
Metallo CM, Gameiro PA, Bell EL, Mattaini KR, Yang J, Hiller K, et al. Reductive glutamine metabolism by IDH1 mediates lipogenesis under hypoxia. Nature. 2011;481:380–4.
Mullen AR, Wheaton WW, Jin ES, Chen PH, Sullivan LB, Cheng T, et al. Reductive carboxylation supports growth in tumour cells with defective mitochondria. Nature. 2011;481:385–8.
Wise DR, Ward PS, Shay JE, Cross JR, Gruber JJ, Sachdeva UM, et al. Hypoxia promotes isocitrate dehydrogenase-dependent carboxylation of alpha-ketoglutarate to citrate to support cell growth and viability. Proc Natl Acad Sci. 2011;108:19611–6.
Calvert AE, Chalastanis A, Wu Y, Hurley LA, Kouri FM, Bi Y, et al. Cancer-associated IDH1 promotes growth and resistance to targeted therapies in the absence of mutation. Cell Rep. 2017;19:1858–73.
Chan SM, Thomas D, Corces-Zimmerman MR, Xavy S, Rastogi S, Hong WJ, et al. Isocitrate dehydrogenase 1 and 2 mutations induce BCL-2 dependence in acute myeloid leukemia. Nat Med. 2015;21:178–84.
Konopleva M, Pollyea DA, Potluri J, Chyla B, Hogdal L, Busman T, et al. Efficacy and biological correlates of response in a Phase II study of Venetoclax monotherapy in patients with acute myelogenous leukemia. Cancer Discov. 2016;6:1106–17.
Karpel-Massler G, Ishida CT, Bianchetti E, Zhang Y, Shu C, Tsujiuchi T, et al. Induction of synthetic lethality in IDH1-mutated gliomas through inhibition of Bcl-xL. Nat Commun. 2017;8:1067.
Tateishi K, Wakimoto H, Iafrate AJ, Tanaka S, Loebel F, Lelic N, et al. Extreme vulnerability of IDH1 mutant cancers to NAD+ depletion. Cancer Cell. 2015;28:773–84.
Saha SK, Gordan JD, Kleinstiver BP, Vu P, Najem MS, Yeo JC, et al. Isocitrate dehydrogenase mutations confer Dasatinib hypersensitivity and SRC dependence in intrahepatic cholangiocarcinoma. Cancer Discov. 2016;6:727–39.
Koehler A, Van Noorden CJ. Reduced nicotinamide adenine dinucleotide phosphate and the higher incidence of pollution-induced liver cancer in female flounder. Environ Toxicol Chem. 2003;22:2703–10.
Holmgren A, Lu J. Thioredoxin and thioredoxin reductase: current research with special reference to human disease. Biochem Biophys Res Commun. 2010;396:120–4.
Salvemini F, Franze A, Iervolino A, Filosa S, Salzano S, Ursini MV. Enhanced glutathione levels and oxidoresistance mediated by increased glucose-6-phosphate dehydrogenase expression. J Biol Chem. 1999;274:2750–7.
Van Noorden CJ, Butcher RG. A quantitative histochemical study of NADPH-ferrihemoprotein reductase activity. Histochem J. 1986;18:364–70.
Spitz DR, Azzam EI, Li JJ, Gius D. Metabolic oxidation/reduction reactions and cellular responses to ionizing radiation: a unifying concept in stress response biology. Cancer Metastas- Rev. 2004;23:311–22.
Fu Y, Zheng S, Zheng Y, Huang R, An N, Liang A, et al. Glioma derived isocitrate dehydrogenase-2 mutations induced up-regulation of HIF-1alpha and beta-catenin signaling: possible impact on glioma cell metastasis and chemo-resistance. Int J Biochem Cell Biol. 2012;44:770–5.
Li K, Ouyang L, He M, Luo M, Cai W, Tu Y, et al. IDH1 R132H mutation regulates glioma chemosensitivity through Nrf2 pathway. Oncotarget. 2017;8:28865–79.
Boutzen H, Saland E, Larrue C, de Toni F, Gales L, Castelli FA, et al. Isocitrate dehydrogenase 1 mutations prime the all-trans retinoic acid myeloid differentiation pathway in acute myeloid leukemia. J Exp Med. 2016;213:483–97.
Li S, Chou AP, Chen W, Chen R, Deng Y, Phillips HS, et al. Overexpression of isocitrate dehydrogenase mutant proteins renders glioma cells more sensitive to radiation. Neuro Oncol. 2013;15:57–68.
Wahl DR, Dresser J, Wilder-Romans K, Parsels JD, Zhao SG, Davis M, et al. Glioblastoma therapy can be augmented by targeting IDH1-mediated NADPH biosynthesis. Cancer Res. 2017;77:960–70.
Shi J, Sun B, Shi W, Zuo H, Cui D, Ni L, et al. Decreasing GSH and increasing ROS in chemosensitivity gliomas with IDH1 mutation. Tumour Biol. 2015;36:655–62.
Shi J, Zuo H, Ni L, Xia L, Zhao L, Gong M, et al. An IDH1 mutation inhibits growth of glioma cells via GSH depletion and ROS generation. Neurol Sci. 2014;35:839–45.
Zhu H, Zhang Y, Chen J, Qiu J, Huang K, Wu M, et al. IDH1 R132H mutation enhances cell migration by activating AKT-mTOR signaling pathway, but sensitizes cells to 5-FU treatment as NADPH and GSH are reduced. PLoS ONE. 2017;12:e0169038.
Sasaki M, Knobbe CB, Itsumi M, Elia AJ, Harris IS, Chio II, et al. D-2-hydroxyglutarate produced by mutant IDH1 perturbs collagen maturation and basement membrane function. Genes Dev. 2012;26:2038–49.
Sasaki M, Knobbe CB, Munger JC, Lind EF, Brenner D, Brustle A, et al. IDH1(R132H) mutation increases murine haematopoietic progenitors and alters epigenetics. Nature. 2012;488:656–9.
Koivunen P, Lee S, Duncan CG, Lopez G, Lu G, Ramkissoon S, et al. Transformation by the (R)-enantiomer of 2-hydroxyglutarate linked to EGLN activation. Nature. 2012;483:484–8.
Chaturvedi A, Araujo Cruz MM, Jyotsana N, Sharma A, Yun H, Gorlich K, et al. Mutant IDH1 promotes leukemogenesis in vivo and can be specifically targeted in human AML. Blood. 2013;122:2877–87.
Atai NA, Renkema-Mills NA, Bosman J, Schmidt N, Rijkeboer D, Tigchelaar W, et al. Differential activity of NADPH-producing dehydrogenases renders rodents unsuitable models to study IDH1R132 mutation effects in human glioblastoma. J Histochem Cytochem. 2011;59:489–503.
Charitou P, Rodriguez-Colman M, Gerrits J, van Triest M, Groot Koerkamp M, Hornsveld M, et al. FOXOs support the metabolic requirements of normal and tumor cells by promoting IDH1 expression. EMBO Rep. 2015;16:456–66.
Zarei M, Lal S, Parker SJ, Nevler A, Vaziri-Gohar A, Dukleska K, et al. Posttranscriptional upregulation of IDH1 by HuR establishes a powerful survival phenotype in pancreatic cancer cells. Cancer Res. 2017;77:4460–71.
Inoue S, Li WY, Tseng A, Beerman I, Elia AJ, Bendall SC, et al. Mutant IDH1 downregulates ATM and alters DNA repair and sensitivity to DNA damage independent of TET2. Cancer Cell. 2016;30:337–48.
Sulkowski PL, Corso CD, Robinson ND, Scanlon SE, Purshouse KR, Bai H, et al. 2-Hydroxyglutarate produced by neomorphic IDH mutations suppresses homologous recombination and induces PARP inhibitor sensitivity. Sci Transl Med. 2017;9:eaal2463.
Molenaar RJ, Radivoyevitch T, Nagata Y, Khurshed M, Przychodzen B, Makishima H, et al. IDH1/2mutations sensitize acute myeloid leukemia to PARP inhibition and this is reversed by IDH1/2-mutantinhibitors, Clin Cancer Res 2018 (in press).
Kim SY, Yoo YH, Park JW. Silencing of mitochondrial NADP(+)-dependent isocitrate dehydrogenase gene enhances glioma radiosensitivity. Biochem Biophys Res Commun. 2013;433:260–5.
Lee JH, Park JW. Oxalomalate regulates ionizing radiation-induced apoptosis in mice. Free Radic Biol Med. 2007;42:44–51.
Wang JB, Dong DF, Wang MD, Gao K. IDH1 overexpression induced chemotherapy resistance and IDH1 mutation enhanced chemotherapy sensitivity in Glioma cells in vitro and in vivo. Asian Pac J Cancer Prev. 2014;15:427–32.
Chen F, Bian K, Tang Q, Fedeles BI, Singh V, Humulock ZT, et al. Oncometabolites d- and l-2-hydroxyglutarate inhibit the AlkB family DNA repair enzymes under physiological conditions. Chem Res Toxicol. 2017;30:1102–10.
Wang P, Wu J, Ma S, Zhang L, Yao J, Hoadley KA, et al. Oncometabolite D-2-hydroxyglutarate inhibits ALKBH DNA repair enzymes and sensitizes IDH mutant cells to alkylating agents. Cell Rep. 2015;13:2353–61.
Mallette FA, Mattiroli F, Cui G, Young LC, Hendzel MJ, Mer G, et al. RNF8- and RNF168-dependent degradation of KDM4A/JMJD2A triggers 53BP1 recruitment to DNA damage sites. EMBO J. 2012;31:1865–78.
Young LC, McDonald DW, Hendzel MJ. Kdm4b histone demethylase is a DNA damage response protein and confers a survival advantage following gamma-irradiation. J Biol Chem. 2013;288:21376–88.
Lu C, Ward PS, Kapoor GS, Rohle D, Turcan S, Abdel-Wahab O, et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature. 2012;483:474–8.
Xu W, Yang H, Liu Y, Yang Y, Wang P, Kim SH, et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of alpha-ketoglutarate-dependent dioxygenases. Cancer Cell. 2011;19:17–30.
Cimmino L, Dolgalev I, Wang Y, Yoshimi A, Martin GH, Wang J, et al. Restoration of TET2 function blocks aberrant self-renewal and leukemia progression. Cell. 2017;170:1079–95.e20.
Kernytsky A, Wang F, Hansen E, Schalm S, Straley K, Gliser C, et al. IDH2 mutation-induced histone and DNA hypermethylation is progressively reversed by small-molecule inhibition. Blood. 2015;125:296–303.
Tateishi K, Higuchi F, Miller JJ, Koerner MVA, Lelic N, Shankar GM, et al. The alkylating chemotherapeutic Temozolomide induces metabolic stress in IDH1-mutant cancers and potentiates NAD+ depletion-mediated cytotoxicity. Cancer Res. 2017;77:4102–15.
Kinnaird A, Zhao S, Wellen KE, Michelakis ED. Metabolic control of epigenetics in cancer. Nat Rev Cancer. 2016;16:694–707.
Tonjes M, Barbus S, Park YJ, Wang W, Schlotter M, Lindroth AM, et al. BCAT1 promotes cell proliferation through amino acid catabolism in gliomas carrying wild-type IDH1. Nat Med. 2013;19:901–8.
Schumacher T, Bunse L, Pusch S, Sahm F, Wiestler B, Quandt J, et al. A vaccine targeting mutant IDH1 induces antitumour immunity. Nature. 2014;512:324–7.
Farshidfar F, Zheng S, Gingras MC, Newton Y, Shih J, Robertson AG, et al. Integrative genomic analysis of cholangiocarcinoma identifies distinct IDH-mutant molecular profiles. Cell Rep. 2017;18:2780–94.
Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND, et al. Genomic classification and prognosis in acute myeloid leukemia. N Engl J Med. 2016;374:2209–21.
Amatangelo MD, Quek L, Shih A, Stein EM, Roshal M, David MD, et al. Enasidenib induces acute myeloid leukemia cell differentiation to promote clinical response. Blood. 2017;130:732–41.
Yen K, Travins J, Wang F, David MD, Artin E, Straley K, et al. AG-221, a first-in-class therapy targeting acute myeloid leukemia harboring oncogenic IDH2 mutations. Cancer Discov. 2017;7:478–93.
Lord CJ, Ashworth A. PARP inhibitors: synthetic lethality in the clinic. Science. 2017;355:1152–8.
Gentile M, Petrungaro A, Uccello G, Vigna E, Recchia AG, Caruso N, et al. Venetoclax for the treatment of chronic lymphocytic leukemia. Expert Opin Investig Drugs. 2017;26:1307–16.
Okoye-Okafor UC, Bartholdy B, Cartier J, Gao EN, Pietrak B, Rendina AR, et al. New IDH1 mutant inhibitors for treatment of acute myeloid leukemia. Nat Chem Biol. 2015;11:878–86.
Acknowledgements
This work was supported by AMC PhD Scholarship (to R.J.M), the Dutch Cancer Society (KWF; UVA 2014-6839, to R.J.M. and C.J.F.v.N and AMC2016.1-10460, to R.J.M., J.W.W. and C.J.F.v.N.), the National Institutes of Health (Bethesda, MD; NIH) grants R01HL118281, R01HL123904, R01HL132071, R35HL135795, a grant from the AA & MDS International Foundation (Rockville, MD), and the Robert Duggan Charitable Fund (Cleveland, OH, all to J.P.M.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
J.P.M. has received honoraria, has performed consultancy, and has served as a speaker on behalf of Celgene. The remaining authors declare that they have no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. If you remix, transform, or build upon this article or a part thereof, you must distribute your contributions under the same license as the original. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/.
About this article
Cite this article
Molenaar, R.J., Maciejewski, J.P., Wilmink, J.W. et al. Wild-type and mutated IDH1/2 enzymes and therapy responses. Oncogene 37, 1949–1960 (2018). https://doi.org/10.1038/s41388-017-0077-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-017-0077-z
This article is cited by
-
Scutellarin activates IDH1 to exert antitumor effects in hepatocellular carcinoma progression
Cell Death & Disease (2024)
-
Isocitrate dehydrogenase 1 mutation drives leukemogenesis by PDGFRA activation due to insulator disruption in acute myeloid leukemia (AML)
Leukemia (2023)
-
TNFα-induced IDH1 hyperacetylation reprograms redox homeostasis and promotes the chemotherapeutic sensitivity
Oncogene (2023)
-
IDH2, a novel target of OGT, facilitates glucose uptake and cellular bioenergy production via NF-κB signaling to promote colorectal cancer progression
Cellular Oncology (2023)
-
Optimizing Patient Pathways in Advanced Biliary Tract Cancers: Recent Advances and a French Perspective
Targeted Oncology (2023)