Abstract
Epigenetic alterations in DNA methylation might mediate gene expression effects of trauma underlying PTSD symptoms, or effects of PTSD on related health problems. PTSD is associated with all-cause morbidity and premature mortality, suggesting accelerated biological aging. We measured genome-wide DNA methylation (Illumina MethylationEPIC BeadChip) in whole blood in a treatment study for combat-related PTSD - “PROGrESS”, a multisite RCT comparing sertraline plus enhanced medication management (SERT + EMM), prolonged exposure (PE) therapy plus placebo (PE + PLB), and the combination (SERT + PE). DNA methylation was measured in 140 US military veterans who served in Iraq and/or Afghanistan (112 current PTSD cases enrolled in a PTSD treatment study and 28 veterans without PTSD history controls), and also 59 non-trauma exposed controls at baseline posttreatment (24 weeks after baseline). Increased DNA methylation GrimAge acceleration (p = 8.8e−09) was observed in patients with PTSD compared to a pooled control group (trauma exposed and non-trauma exposed), suggesting a higher risk of premature mortality in those with PTSD. There was no difference in GrimAge acceleration between combat trauma and non-trauma exposed controls. No treatment-related changes in GrimAge acceleration were found in within-subject comparisons of PTSD patients pre- to post-treatment.
Similar content being viewed by others
Introduction
Trauma burden and posttraumatic stress disorder (PTSD) may potentially lead to poor health outcomes, higher risk of chronic medical conditions, and mortality, independent of lifestyle factors [1,2,3,4,5]. This higher risk of mortality may be due to accelerated cellular aging, which is a potential consequence of dysregulated autonomic function and increased oxidative stress and inflammation, often observed in those with PTSD [6,7,8].
Cellular aging is associated with highly reliable changes in the epigenome; hence, investigators have been able to leverage DNA methylation data to design epigenetic clocks that accurately capture age [9,10,11,12]. Multiple studies that used Hannum’s epigenetic clock showed that individuals with PTSD have higher DNAm age values compared to their chronological age (i.e., DNAm age acceleration) [6, 13, 14]. In contrast, studies that used Horvath’s epigenetic clock reported that DNAm age acceleration was negatively associated with PTSD [15] and development of PTSD symptoms [16]. The most recent epigenetic clock, DNAm GrimAge, is a better predictor of age-related health outcomes (e.g., time to death, cancer, and coronary heart disease) compared to earlier clocks, since it has been specifically trained on time-to-death, as well as age-related biochemical markers and smoking packs per year [12]. Recent evidence demonstrates a significant association between PTSD and GrimAge acceleration [17,18,19]. Specifically, in a predominantly African American cohort, Katrinli et al. showed that individuals with current and lifetime PTSD have accelerated GrimAge, which also associated with cortical thickness in brain areas related to emotion processing and threat response [17]. Yang et al. reported accelerated GrimAge in male veterans with PTSD, as well as a significant positive association between longitudinal increases in GrimAge acceleration and increases in PTSD symptoms at 3-years follow-up [18]. The longitudinal study from Mehta et al. demonstrated that following exposure to traumatic events, PTSD symptom severity score associated with accelerated GrimAge [19]. The association between PTSD and accelerated GrimAge might partly be explained by immune system dysfunction, since GrimAge acceleration is associated with decreased lymphocyte and increased neutrophil proportions (i.e., higher neutrophil-to-lymphocyte ratio), which is an indicator of immune senescence and inflammation [12, 17].
PTSD can be effectively treated with medications or psychotherapy, and PTSD in veterans is most commonly treated with a combination of medication and psychotherapy [20]. The VA/DOD PTSD Treatment Clinical Practice Guidelines [21] (VA/DOD, 2017) recommend both trauma-focused psychotherapy (e.g., prolonged exposure (PE) therapy) and selective serotonin reuptake inhibitors (SSRIs) as effective treatments for PTSD. Indeed, the treated veterans we examined in the current genetic analyses were enrolled in a randomized controlled trial of PTSD treatment symptom outcome that included PE + SERT, SERT + EMM, and PE + PLB. a “head-to-head” multisite RCT comparing sertraline plus enhanced medication management (SERT + EMM), PE therapy plus placebo (PE + PLB), and the combination (SERT + PE). All showed significant and large reductions in PTSD with no significant differences between groups, and >50% of veterans in each treatment condition had clinically meaningful effects [22]. Successful treatment of PTSD with psychotherapy appears to ameliorate autonomic hyperarousal [23], and may have direct effects on DNA methylation [24], suggesting effects of PTSD on DNA methylation might be reversible.
To the best of our knowledge, no studies to date have examined the effect of PTSD treatment and PTSD remission on GrimAge acceleration prospectively. So far, one cross-sectional study reported that GrimAge acceleration did not significantly differ between current PTSD cases and individuals who do not meet current PTSD criteria but had a previous history of PTSD, suggesting that the effect of PTSD on GrimAge acceleration might not be reversible [17] or might reflect differences that were present prior to trauma exposure. In the present study, we first aimed to evaluate the association between PTSD and mortality at baseline, using the most recent epigenetic predictor of lifespan, GrimAge. Then, we sought to investigate whether the impact of PTSD on accelerated GrimAge might be reversible by PTSD treatment and PTSD remission.
Materials and methods
Participants
The participants included in the study were part of a larger study of the PROlonGed ExpoSure and Sertraline Trial (PROGrESS), a four-site [VA Ann Arbor Healthcare System (VAAAHS), Ralph H. Johnson VA Medical Center (CHSVAMC), Massachusetts General Hospital (MGH), and VA San Diego Healthcare System (VASDHCS) randomized-controlled trial (RCT; N = 223), designed to examine: (1) the comparative effectiveness of Prolonged Exposure plus placebo (PE/PLB), Sertraline plus Enhanced Medication Management (SERT/EMM), or combined treatment (PE/SERT) on PTSD, and (2) neurobiological predictors and potential biomarkers of treatment response including hypothalamic-pituitary-adrenal axis, brain, and genetic/genomic biomarkers [22, 25]. PROGrESS was approved by the institutional review boards at VHAAAHS, the University of Michigan, VASDHCS, CHSVAMC, MGH and the Department of Defense Human Research Protection Office. Participants provided written informed consent before enrollment. Participants and clinicians were blinded to pill condition through week 24, and independent evaluators were blinded to treatment assignments for the duration of the study.
The PROGrESS study methods were published in detail [22, 25]. Briefly, inclusion criteria were service members or veterans of the Iraq or Afghanistan wars with combat-related PTSD and significant impairment (Clinicians-Administered PTSD Scale for DSM-IV [CAPS-IV] [26] score ≥ 50) of at least 3 months’ duration. Exclusion criteria included factors related to safety and appropriateness of psychotherapy and sertraline treatment [25, 27].
Measures
Demographics, childhood adversity, and combat experiences history were obtained by self-report at intake, and psychiatric symptoms were assessed by self-report and clinician-administered measures at intake and weeks 6, 12, 24, 36, and 52; blinding was broken at week 24. Adverse childhood experiences were assessed by the Deployment Risk and Resilience Inventory (DRRI) section A - Prior Stressors [28] using a 0–5 point score of dichotomous endorsement (yes vs no) of physical, sexual, and emotional abuse items. Exposure to combat trauma was measured with the Combat Experiences Scale (CES) [29]. PTSD diagnosis and symptom severity in the past month were measured with the CAPS-IV (for DSM-IV) clinician interview [26], and self-reported symptoms of PTSD were assessed using the PTSD Checklist [PCL] Specific Stressor Version [30]. PTSD remission was defined as a CAPS score of 35 or less at week 24 (or last observed CAPS score) [22]. Additional details on measures can be found in the methods and main outcomes publications [22, 25].
DNA methylation and GrimAge
Blood samples were collected at all four sites at baseline and post-treatment (Week 24) by venipuncture at hospital phlebotomy stations, ~7 ml collected into 10 ml lavender-top EDTA vacutainer tubes. Blood samples were centrifuged at 4 C at 2000 × g to separate plasma from cellular components, and care was taken to exclude buffy coat (white blood cells, WBC) from the supernatant. Separate aliquots of packed red blood cells + buffy coat WBC pellet and plasma supernatant were frozen, stored at −80C, and shipped to VAAAHS overnight on dry ice with no thaw. Genomic DNA extracted from blood pellets was assayed using the Methylation EPIC BeadChip (Illumina, Inc) at the University of Michigan Advanced Genomics Core. R package CpGassoc was used for quality control steps, including (i) removal of samples with probe detection call rates <90% and an average intensity value of either <50% of the experiment-wide sample mean or <2000 arbitrary units; (ii) setting probes with detection p values >0.01 as missing; and (iii) filtering out missing probes for >10% of samples [31]. Probes that were known to cross-hybridize between autosomes and sex chromosomes were filtered out [32]. Methylation data was preprocessed and normalized using single-sample Noob background correction implemented in R package minfi [33]. Batch effects of chip and position were removed using ComBat [34].
GrimAge and GrimAge acceleration (i.e., age-adjusted GrimAge) were computed using the DNA methylation age calculator (https://dnamage.genetics.ucla.edu/new) [12]. DNA methylation data were used to estimate cellular heterogeneity (i.e., the proportion of CD8 + T, CD4 + T, natural killer (NK), B cells, monocytes, and neutrophils), using the Robust Partial Correlation method implemented in R package Epidish [35] and the blood reference panel generated by Salas and colleagues [36]. To validate our findings regarding changes in cellular composition, we also used Houseman method [37] with the blood reference panel generated by Reinius et al. [38] as an alternative. DNA methylation data was used to generate methylation-based ancestry principal components (mPCs), following the method described by Barfield et al. [39]. The components mPC1 and mPC2 correlated most with self-reported ancestry (Spearman rho = 0.75, p < 2.2e−16 for mPC1 and Spearman rho = −0.81, p < 2.2e−16 for mPC2; Fig. S1) and were included as covariates in subsequent analyses to adjust for ancestry. DNA methylation based smoking scores for each sample were computed using the weights of 39 CpG sites associated with smoking pack years [40], as previously described [41].
Plasma interleukin-6 (IL-6) concentration
IL-6 levels were measured in the plasma (supernatant) using Immulite™ (Siemens, Inc.), a rapid highly sensitive and precise semi-automated chemiluminescent assay that has an intra-assay variability of <5%.
Statistical analysis
We tested the correlations between GrimAge acceleration and potential confounders, including sex, self-reported ancestry, cellular heterogeneity, and methylation-derived smoking score. Associations of GrimAge acceleration with trauma exposure and PTSD at baseline were evaluated using linear regression models, adjusting for sex, methylation-based ancestry principal components (mPC1 and mPC2), and the estimated proportions of CD8 + T, CD4 + T, NK, B cells and monocytes. Pre- to post-treatment differences in CAPS, cellular variables, IL-6 concentration, and GrimAge acceleration were assessed by non-parametric paired samples Wilcoxon tests, since the variables are not normally distributed (Shapiro–Wilk normality test, p < 0.05). To evaluate whether the pre- to post-treatment change in GrimAge acceleration associated with PTSD remission (as dichotomous PTSD remission variable and as percent change in CAPS score), we conducted longitudinal analyses using linear regression models, where post-treatment GrimAge acceleration was modeled as a function of PTSD remission (as dichotomous PTSD remission variable and as percent change in CAPS score), while adjusting for pre-treatment GrimAge acceleration, sex, ancestry principal components, and changes in estimated CD8T+, CD4T+, NK, B cell, and monocyte cell proportions. Since smoking has already been included in training of GrimAge, we did not include methylation-derived smoking score as a covariate in our initial analyses, as this redundance could introduce error due to overfitting of models. However, for significant findings, we conducted post-hoc sensitivity analyses to explore possible confounding effects of smoking, by including methylation-derived smoking score as a covariate.
Results
Demographic and clinical characteristics
Analytical sample included 199 participants from the PROGrESS study, with DNAm measured at baseline for 59 non-combat exposed controls, 28 combat exposed controls without PTSD, and 112 combat exposed PTSD cases in the treatment study. At post-treatment (24 weeks after baseline) sample included 109 patients with PTSD (out of 112 PTSD at baseline). Demographics and characteristics of the analytical sample are presented in Table 1.
Of the 109 subject pairs with baseline and post-treatment data available, 40 were SERT + EMM, 40 were SERT + PE, and 29 were PE + PLB. Baseline and post-treatment characteristics of those subject pairs are described in Table S1 and categorized based on treatment arms in Tables S2 and S3.
Correlations between GrimAge acceleration and demographic and cellular variables
GrimAge was strongly correlated with chronological age at baseline (N = 199, r = 0.93, p < 2.2e−16, Fig. S2A) and at post-treatment (N = 109, r = 0.91, p < 2.2e−16, Fig. S2B). GrimAge acceleration positively correlated with smoking score and Neu proportions (Fig. S3A), and negatively correlated with CD4T, NK, and B cell proportions both at baseline and at post-treatment (Fig. S3B). CD8T proportions negatively correlated with GrimAge acceleration at baseline (Fig. S3A), but not at post-treatment (Fig. S3B). Self-reported ancestry and IL-6 levels were not correlated with GrimAge acceleration either at baseline or at post-treatment (Fig. S3).
Baseline PTSD associates with GrimAge acceleration
Although combat trauma exposed controls had slightly higher GrimAge acceleration compared to non-trauma exposed controls (Table 1), this difference was not statistically significant (Table 2). GrimAge acceleration did not associate with CES in participants exposed to combat trauma. However, participants with current PTSD at baseline had higher GrimAge acceleration when compared to all controls (combat exposed controls and non-trauma exposed controls pooled together), when compared to only combat exposed controls, and also when compared to only non-trauma exposed controls (Table 2, Fig. 1), suggesting an increased mortality risk for those with PTSD. The associations between baseline PTSD and GrimAge acceleration were still significant in sensitivity analyses with smoking score (p < 0.01, Table S4).
GrimAge acceleration did not significantly differ between combat controls and non-combat controls (p = 0.25). GrimAge acceleration at baseline was higher in PTSD cases compared to all controls (p = 8.77e−09), compared to combat controls (p = 3.31e−03), and compared to non-combat controls (p = 1.0e−07). ***p < 0.001, ns: not significant (p > 0.05).
Pre- to post-treatment changes in cellular variables and GrimAge acceleration
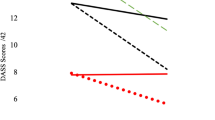
We observed decrease in estimated CD4T, CD8T, monocyte, B cell, and NK proportions, and an increase in neutrophil proportions and IL-6 levels from pre- to post-treatment (Table S1, Fig. 2), suggestive of increased physiologic stress and immune dysregulation. Although CAPS score significantly declined after 24 weeks post-treatment (p < 2.2e−16), we did not observe a significant change in GrimAge acceleration pre- to post-treatment (p = 0.14; Table S1). In addition, the change in GrimAge acceleration did not associate with percent change in CAPS score and PTSD remission (Table S5).
Shown are differences in (A) CD4T proportions, (B) CD8T proportions, (C) Monocyte proportions, (D) B cell proportions, (E) Neutral killer cell proportions, (F) Neutrophil proportions, (G) CAPS score, (H) GrimAge acceleration, and (I) Plasma IL-6 (pIL6) concentration. Significant differences are indicated with asterisk (*).
Changes in cellular variables and GrimAge acceleration based on treatment type
While previously reported outcomes on the full sample found no significant differences in remission (Rauch et al., 2019), PTSD remission rate appeared to be higher in SERT + EMM and SERT + PE, compared to PE + PLB in the current smaller sub-sample (Tables S2 and S3). The demographic analysis conducted in this smaller sub-sample was not adjusted for site, sex, or baseline CAPS. GrimAge acceleration did not significantly change pre- to post-treatment in either treatment arm (Table S3). Furthermore, the change in GrimAge acceleration did not associate with percent change in CAPS score and PTSD remission in either treatment group (Table S5). However, the estimated proportions of CD4T, CD8T, and B cells were decreased in SERT + PE, and neutrophil proportions were elevated in both SERT + EMM and SERT + PE after treatment (Table S3, Fig. 3), which may indicate dysregulated immune response in these treatment arms.
Shown are differences in (A) CD4T proportions, (B) CD8T proportions, (C) Monocyte proportions, (D) B cell proportions, (E) Neutral killer cell proportions, (F) Neutrophil proportions, (G) CAPS score, (H) GrimAge acceleration, and (I) Plasma IL-6 (pIL6) concentration. Significant differences are indicated with asterisk (*).
Discussion
Recent consortium studies show that PTSD associates with changes in DNA methylation patterns involved in inflammatory and oxidative stress pathways [42,43,44]. In addition to the alterations in DNA methylation signatures, PTSD is associated with accelerated GrimAge, which is an epigenetic marker of mortality [12, 17,18,19]. While some previous studies show PTSD-associated changes in DNA methylation markers were reversed with successful treatment [24, 45], the effect of PTSD treatment and remission on accelerated GrimAge is yet to be elucidated.
In our sample and consistent with previous findings, PTSD associated with GrimAge acceleration at baseline; and thus, premature mortality in those with PTSD could be inferred [17,18,19]. GrimAge acceleration positively correlated with estimated neutrophil proportion, and negatively correlated with T and B lymphocyte proportions. Increased neutrophil and decreased lymphocyte proportions indicate inflammation and immune dysfunction, and also associate with mortality [46]. Our findings align with previous research that reported a positive correlation between GrimAge acceleration and CD8 + CD28 − T (CD8T cells without CD28 surface marker) proportions, which is a biomarker of immunosenescence, suggesting the immune system as an important contributor to epigenetic aging [18].
In the present study, we did not observe a significant change in GrimAge acceleration between baseline and treatment follow-up at 24 weeks. Moreover, pre- to post-treatment change in GrimAge acceleration did not associate with PTSD remission when all treatment arms were combined or analyzed separately. Interestingly, estimated neutrophil proportions increased pre- to post-treatment in the overall sample, SERT + EMM and SERT + PE, while proportions of T and B lymphocytes decreased in the overall sample and SERT + PE. We also observed increase in IL-6 levels pre- to post-treatment in the overall sample. Taken together, these findings indicate that successful PTSD treatment may not change the association betweenPTSD and accelerated GrimAge, at least not within 24 weeks. Immune dysregulation represented by increased neutrophils and decreased lymphocytes, alongside with increased IL-6 levels, may explain the lack of treatment effect on GrimAge acceleration. Of note, it may be true that the impact of treatment on GrimAge acceleration and even on the immune response may require a longer period of follow-up and assessment. Indeed, these markers may take a period of months post-treatment or even years to be reflecting in neurobiology [47]. In a recent clinical trial aimed to regenerate the thymus, authors observed protective changes in immunological markers and reversal of epigenetic aging [48]. Specifically, the rate of GrimAge reversal relative to chronological age accelerated from −0.72 year/year from 0–9 month to −6.48 year/year from 9–12 month, suggesting that the rate of reversal accelerates substantially with increasing treatment time [48]. In addition, a 3-year follow-up study reported that longitudinal changes in GrimAge acceleration correlated with longitudinal changes in PTSD symptom severity, albeit the study did not prospectively assess treatment status [18]. Additional examinations in large ethnically diverse cohorts over a longer follow-up period is warranted to fully understand the effect of PTSD treatment on reversal of GrimAge acceleration.
This study is not without limitations. First, due to relatively low sample size, our association studies are underpowered. Even so, we report significant associations of GrimAge acceleration with PTSD at baseline. However, we are underpowered to test the interaction effect of treatment type, which would be useful to further characterize the observed associations. Second, since all follow-up over time occurred in the context of treatment, it is unclear how these may compare to no treatment time control group. Third, our findings are based on combat-related PTSD; therefore, may not extrapolate to other types of PTSD. It is possible that GrimAge acceleration results from other factors that increase risk for PTSD development and not reflective of the impact a specific trauma that results in PTSD.
In conclusion, accelerated GrimAge was apparent in those with PTSD, suggesting a shorter lifespan. In addition, the effect of PTSD on GrimAge acceleration may not revert with successful treatment, at least not within 24 weeks. Further prospective studies with larger sample sizes and longer follow-up times are required to investigate the effect of successful PTSD treatment on accelerated GrimAge.
References
Kubzansky LD, Koenen KC, Jones C, Eaton WW. A prospective study of posttraumatic stress disorder symptoms and coronary heart disease in women. Health Psychol. 2009;28:125–30.
O’Donovan A, Cohen BE, Seal KH, Bertenthal D, Margaretten M, Nishimi K, et al. Elevated risk for autoimmune disorders in iraq and afghanistan veterans with posttraumatic stress disorder. Biol Psychiatry. 2015;77:365–74.
Rosenbaum S, Stubbs B, Ward PB, Steel Z, Lederman O, Vancampfort D. The prevalence and risk of metabolic syndrome and its components among people with posttraumatic stress disorder: a systematic review and meta-analysis. Metabolism 2015;64:926–33.
Hung YH, Cheng CM, Lin WC, Bai YM, Su TP, Li CT, et al. Post-traumatic stress disorder and asthma risk: A nationwide longitudinal study. Psychiatry Res. 2019;276:25–30.
Schlenger WE, Corry NH, Williams CS, Kulka RA, Mulvaney-Day N, DeBakey S, et al. A Prospective Study of Mortality and Trauma-Related Risk Factors Among a Nationally Representative Sample of Vietnam Veterans. Am J Epidemiol. 2015;182:980–90.
Wolf EJ, Logue MW, Stoop TB, Schichman SA, Stone A, Sadeh N, et al. Accelerated DNA Methylation Age: Associations With Posttraumatic Stress Disorder and Mortality. Psychosom Med. 2018;80:42–8.
Lohr JB, Palmer BW, Eidt CA, Aailaboyina S, Mausbach BT, Wolkowitz OM, et al. Is Post-Traumatic Stress Disorder Associated with Premature Senescence? A Review of the Literature. Am J Geriatr Psychiatry. 2015;23:709–25.
Fonkoue IT, Marvar PJ, Norrholm S, Li Y, Kankam ML, Jones TN, et al. Symptom severity impacts sympathetic dysregulation and inflammation in post-traumatic stress disorder (PTSD). Brain Behav Immun. 2020;83:260–69.
Horvath S. DNA methylation age of human tissues and cell types. Genome Biol. 2013;14:R115.
Hannum G, Guinney J, Zhao L, Zhang L, Hughes G, Sadda S, et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol Cell. 2013;49:359–67.
Levine ME, Lu AT, Quach A, Chen BH, Assimes TL, Bandinelli S, et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging (Albany NY). 2018;10:573–91.
Lu AT, Quach A, Wilson JG, Reiner AP, Aviv A, Raj K, et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging (Albany NY). 2019;11:303–27.
Wolf EJ, Logue MW, Hayes JP, Sadeh N, Schichman SA, Stone A, et al. Accelerated DNA methylation age: Associations with PTSD and neural integrity. Psychoneuroendocrinology 2016;63:155–62.
Wolf EJ, Maniates H, Nugent N, Maihofer AX, Armstrong D, Ratanatharathorn A, et al. Traumatic stress and accelerated DNA methylation age: A meta-analysis. Psychoneuroendocrinology 2018;92:123–34.
Verhoeven JE, Yang R, Wolkowitz OM, Bersani FS, Lindqvist D, Mellon SH, et al. Epigenetic Age in Male Combat-Exposed War Veterans: Associations with Posttraumatic Stress Disorder Status. Mol Neuropsychiatry. 2018;4:90–9.
Boks MP, van Mierlo HC, Rutten BP, Radstake TR, De Witte L, Geuze E, et al. Longitudinal changes of telomere length and epigenetic age related to traumatic stress and post-traumatic stress disorder. Psychoneuroendocrinology 2015;51:506–12.
Katrinli S, Stevens J, Wani AH, Lori A, Kilaru V, van Rooij SJH, et al. Evaluating the impact of trauma and PTSD on epigenetic prediction of lifespan and neural integrity. Neuropsychopharmacology 2020;45:1609–16.
Yang R, Wu GWY, Verhoeven JE, Gautam A, Reus VI, Kang JI, et al. A DNA methylation clock associated with age-related illnesses and mortality is accelerated in men with combat PTSD. Mol Psychiatry. 2021;26:4999–5009.
Mehta D, Bruenig D, Pierce J, Sathyanarayanan A, Stringfellow R, Miller O, et al. Recalibrating the epigenetic clock after exposure to trauma: The role of risk and protective psychosocial factors. J Psychiatr Res. 2022;149:374–81.
Bohnert KM, Sripada RK, Mach J, McCarthy JF. Same-Day Integrated Mental Health Care and PTSD Diagnosis and Treatment Among VHA Primary Care Patients With Positive PTSD Screens. Psychiatr Serv. 2016;67:94–100.
US Department of Veterans Affairs Management of posttraumatic stress disorder and acute stress reaction 2017. https://www.healthquality.va.gov/guidelines/MH/ptsd/. Accessed 15 August 2022.
Rauch SAM, Kim HM, Powell C, Tuerk PW, Simon NM, Acierno R, et al. Efficacy of Prolonged Exposure Therapy, Sertraline Hydrochloride, and Their Combination Among Combat Veterans With Posttraumatic Stress Disorder: A Randomized Clinical Trial. JAMA Psychiatry. 2019;76:117–26.
Maples-Keller JL, Rauch SAM, Jovanovic T, Yasinski CW, Goodnight JM, Sherrill A, et al. Changes in trauma-potentiated startle, skin conductance, and heart rate within prolonged exposure therapy for PTSD in high and low treatment responders. J Anxiety Disord. 2019;68:102147.
Vinkers CH, Geuze E, van Rooij SJH, Kennis M, Schur RR, Nispeling DM, et al. Successful treatment of post-traumatic stress disorder reverses DNA methylation marks. Mol Psychiatry. 2021;26:1264–71.
Rauch SAM, Simon NM, Kim HM, Acierno R, King AP, Norman SB, et al. Integrating biological treatment mechanisms into randomized clinical trials: Design of PROGrESS (PROlonGed ExpoSure and Sertraline Trial). Contemp Clin Trials. 2018;64:128–38.
Blake DD, Weathers FW, Nagy LM, Kaloupek DG, Gusman FD, Charney DS, et al. The development of a Clinician-Administered PTSD Scale. J Trauma Stress. 1995;8:75–90.
Ravi M, Stevens JS, Michopoulos V. Neuroendocrine pathways underlying risk and resilience to PTSD in women. Front Neuroendocrinol. 2019;55:100790.
King DW, King LA, Vogt DS. Manual for the Deployment Risk and Resilience Inventory (DRRI): A Collection of Measures for Studying Deployment-Related Experiences of Military Veterans. Boston, MA: National Center for PTSD;2003.
Keane TM, Fairbank JA, Caddell JM, Zimering RT, Taylor KL, Mora CA. Clinical evaluation of a measure to assess combat exposure. Psychol Assess: J Consult Clin Psychol. 1989;1:53–5.
Adkins JW, Weathers FW, McDevitt-Murphy M, Daniels JB. Psychometric properties of seven self-report measures of posttraumatic stress disorder in college students with mixed civilian trauma exposure. J Anxiety Disord. 2008;22:1393–402.
Barfield RT, Kilaru V, Smith AK, Conneely KN. CpGassoc: an R function for analysis of DNA methylation microarray data. Bioinformatics 2012;28:1280–1.
McCartney DL, Walker RM, Morris SW, McIntosh AM, Porteous DJ, Evans KL. Identification of polymorphic and off-target probe binding sites on the Illumina Infinium MethylationEPIC BeadChip. Genom Data. 2016;9:22–4.
Triche TJ Jr, Weisenberger DJ, Van Den Berg D, Laird PW, Siegmund KD. Low-level processing of Illumina Infinium DNA Methylation BeadArrays. Nucleic Acids Res. 2013;41:e90.
Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 2012;28:882–3.
Teschendorff AE, Breeze CE, Zheng SC, Beck S. A comparison of reference-based algorithms for correcting cell-type heterogeneity in Epigenome-Wide Association Studies. BMC Bioinforma. 2017;18:105.
Salas LA, Koestler DC, Butler RA, Hansen HM, Wiencke JK, Kelsey KT, et al. An optimized library for reference-based deconvolution of whole-blood biospecimens assayed using the Illumina HumanMethylationEPIC BeadArray. Genome Biol. 2018;19:64.
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinforma. 2012;13:86.
Reinius LE, Acevedo N, Joerink M, Pershagen G, Dahlen SE, Greco D, et al. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PLoS ONE. 2012;7:e41361.
Barfield RT, Almli LM, Kilaru V, Smith AK, Mercer KB, Duncan R, et al. Accounting for population stratification in DNA methylation studies. Genet Epidemiol. 2014;38:231–41.
Li S, Wong EM, Bui M, Nguyen TL, Joo JE, Stone J, et al. Causal effect of smoking on DNA methylation in peripheral blood: a twin and family study. Clin Epigenet. 2018;10:18.
Logue MW, Miller MW, Wolf EJ, Huber BR, Morrison FG, Zhou Z, et al. An epigenome-wide association study of posttraumatic stress disorder in US veterans implicates several new DNA methylation loci. Clin Epigenet. 2020;12:46.
Smith AK, Ratanatharathorn A, Maihofer AX, Naviaux RK, Aiello AE, Amstadter AB, et al. Epigenome-wide meta-analysis of PTSD across 10 military and civilian cohorts identifies methylation changes in AHRR. Nat Commun. 2020;11:5965.
Snijders C, Maihofer AX, Ratanatharathorn A, Baker DG, Boks MP, Geuze E, et al. Longitudinal epigenome-wide association studies of three male military cohorts reveal multiple CpG sites associated with post-traumatic stress disorder. Clin Epigenet. 2020;12:11.
Katrinli S, Maihofer AX, Wani AH, Pfeiffer JR, Ketema E, Ratanatharathorn A, et al. Epigenome-wide meta-analysis of PTSD symptom severity in three military cohorts implicates DNA methylation changes in genes involved in immune system and oxidative stress. Mol Psychiatry. 2022;27:1720–28.
Yang R, Xu C, Bierer LM, Flory JD, Gautam A, Bader HN, et al. Longitudinal genome-wide methylation study of PTSD treatment using prolonged exposure and hydrocortisone. Transl Psychiatry. 2021;11:398.
Song M, Graubard BI, Rabkin CS, Engels EA. Neutrophil-to-lymphocyte ratio and mortality in the United States general population. Sci Rep. 2021;11:464.
Duan R, Fu Q, Sun Y, Li Q. Epigenetic clock: A promising biomarker and practical tool in aging. Ageing Res Rev. 2022;81:101743.
Fahy GM, Brooke RT, Watson JP, Good Z, Vasanawala SS, Maecker H, et al. Reversal of epigenetic aging and immunosenescent trends in humans. Aging Cell. 2019;18:e13028.
Funding
The DOD had a role in design in that they wanted the study to include only OEF/OIF/OND service members with combat-related PTSD. DOD had no role in the conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the paper; and decision to submit the paper for publication. This work was supported by the U.S. Department of Defense through the U.S. Army Medical Research and Materiel Command (MRMC; Randomized Controlled Trial of Sertraline, Prolonged Exposure Therapy, and Their Combination in OEF/OIF Combat Veterans with PTSD; (Award #W81XWH-11-1-0073; PI: Rauch); the National Center for Advancing Translational Sciences of the National Institutes of Health (Award #UL1TR000433). This material is the result of work supported with resources and the use of facilities at Massachusetts General Hospital, the VA Ann Arbor Healthcare System, Ralph H. Johnson VA Medical Center, and VA San Diego Healthcare System. The views expressed in this paper presentation are solely those of the author(s) and do not reflect an endorsement by or the official policy of the Department of Veterans Affairs, Department of Defense, or the U.S. Government, or the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Concept and design: APK, MS, IL, SAMR; Acquisition of data: APK, ERD, NR, and SAMR; Statistical analysis: SK; Interpretation of the data: SK, APK, ERD, AKS, SAMR; Drafting of the paper: SK, APK, ERD, SAMR; Critical revision of the paper for important intellectual content: SK, APK, ERD, AKS, MS, SAMR; Final approval of the version to be published: All authors.
Corresponding author
Ethics declarations
Competing interests
APK, SK, ERD, NR, AKS, IL, SAMR report no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Katrinli, S., King, A.P., Duval, E.R. et al. DNA methylation GrimAge acceleration in US military veterans with PTSD. Neuropsychopharmacol. 48, 773–780 (2023). https://doi.org/10.1038/s41386-023-01537-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41386-023-01537-z