Abstract
An enhanced understanding of the pathophysiology of depression would facilitate the discovery of new efficacious medications. To this end, we examined hippocampal transcriptional changes in rat models of disease and in humans to identify common disease signatures by using a new algorithm for signature-based clustering of expression profiles. The tool identified a transcriptomic signature comprising 70 probesets able to discriminate depression models from controls in both Flinders Sensitive Line and Learned Helplessness animals. To identify disease-relevant pathways, we constructed an expanded protein network based on signature gene products and performed functional annotation analysis. We applied the same workflow to transcriptomic profiles of depressed patients. Remarkably, a 171-probesets transcriptional signature which discriminated depressed from healthy subjects was identified. Rat and human signatures shared the SCARA5 gene, while the respective networks derived from protein-based significant interactions with signature genes contained 25 overlapping genes. The comparison between the most enriched pathways in the rat and human signature networks identified a highly significant overlap (p-value: 3.85 × 10–6) of 67 terms including ErbB, neurotrophin, FGF, IGF, and VEGF signaling, immune responses and insulin and leptin signaling. In conclusion, this study allowed the identification of a hippocampal transcriptional signature of resilient or susceptible responses in rat MDD models which overlapped with gene expression alterations observed in depressed patients. These findings are consistent with a loss of hippocampal neural plasticity mediated by altered levels of growth factors and increased inflammatory responses causing metabolic impairments as crucial factors in the pathophysiology of MDD.
Similar content being viewed by others
Introduction
About one third of major depressive disorder (MDD) patients do not respond or respond only partially to available treatments [1]. The challenge to discover new therapies is increased by the incomplete understanding of the pathophysiological bases of the disease. Compelling evidence supports the notion that disease onset is caused by genetic predisposition and environmental triggering challenges, principally stress [2]. The identification of the genes has proven difficult [3,4,5], but a recent study identified 17 independent loci [6]. Stressful experiences during childhood are considered as particularly critical for determining a long-term increase in the risk to develop depressive symptoms.
Research using animal models has provided a crucial contribution to the understanding of MDD neurobiology. The Flinders Sensitive Line (FSL) is a strain that phenotypically presents a number of depression symptoms in comparison with the Flinders Resistant Line (FRL) controls. FSL rats show increased immobility in the forced swim test, reduced activity in novel open-field behaviors, decreased appetite and weight loss, sleep disturbances and neurochemical abnormalities, similar to those found in humans [7]. In the learned helplessness (LH) model, the rats are exposed to uncontrollable, unpredictable, and unavoidable stress and subsequently tested for a deficit in acquiring an escape task. This helplessness response is induced only in some of the animals (termed LH rats and compared with the non-responding nLH rats), thereby modeling the hypothesis of variable predisposition for depression in different subjects [8]. With the aim of reproducing the impact of childhood adverse events, the maternal separation (MS) paradigm was applied to model gene × environment interactions [9]. Since psychopathology in humans has mostly been linked to stressful experiences of social nature, social defeat (SD) models stress are considered ethologically valid. In this model, the experimental rats are introduced into the territory of an aggressive male conspecific that attacks and defeats them, inducing long-lasting behavioral and endocrine effects [10]. Since MDD is heterogeneous, it has been suggested that different animal models may help to identify both the diversity of the causes and the shared mechanisms underlying common symptoms of the human disease [11]. This approach has allowed the identification of common transcriptional dysregulations between rodent models and MDD patients [12,13,14].
The objective of this study was to examine gene expression changes in rat models of depression and in humans and identify common signatures of disease. The transcriptional analysis was carried out in the hippocampus since structural and functional neuroimaging studies have revealed that MDD is accompanied by structural changes in this brain region [15]. In the hippocampus, stress exposure induces an allostatic process aimed at reinstating homeostasis by triggering a plasticity response that causes changes in neural architecture as well as in synaptic functions. In MDD, a stress-related disorder, an allostatic overload impairs adaptive plasticity responses, causing dendritic shrinkage and loss of spines and suppressing neurogenesis in the dentate gyrus [16]. These dysregulations are assumed to sustain hippocampal structural alterations observed in patients and evidence is available that they can be reproduced in the hippocampus of animal models of disease [16]. To increase the sensitivity of the analysis we used a recently introduced algorithm for signature-based clustering of expression profiles to identify a common transcriptional signature in different models of MDD, with subsequent functional annotation by means of network-based analysis. Next, we applied the same workflow to transcriptional data generated in depressed patients and discovered a transcriptional signature able to discriminate disease versus control groups. Finally, we compared the findings obtained in the animal models with the human transcriptional signature to evaluate the relevance of the results for understanding the pathophysiological basis of MDD. We discovered that a common transcriptional signature discriminating LH and FSL rats from controls could be identified and that the main affected cellular pathways converged on growth factors as well as metabolic and inflammatory pathways implicated in sustaining neural plasticity, which were similarly influenced in depressed patients.
Materials and methods
Animals
FSL and FRL male rats (n = 38 and 22) were from colonies maintained in the animal facility at the Karolinska Institutet. In SD experiments, Long Evans rats (400–450 g) were purchased from Harlan, USA; Sprague Dawley female (250–300 g) and male rats (300–350 g) were from Charles River, Italy. In LH experiments, male Sprague Dawley rats weighed 200–240 g (Janvier, France). In chronic mild stress (CMS) experiments, male Wistar rats weighing 220–230 g were used (Charles River, Germany). Animals were housed in standard cages and maintained at a constant temperature of 22 ± 1 °C and 12-h light-dark cycle. Food and water were freely available.
Animal handling and experimental procedures were performed in accordance with the European Community Council Directive 86/609/EEC and were approved by local authorities. All efforts were made to minimize animal distress and to reduce the numbers of animals used.
Behavioral procedures
MS was performed as previously published [9]. Briefly, pups assigned to the separation procedure (MS group, 20 rats within the FSL group) were separated from the dam as a litter for 180 min. Control rats were left undisturbed, except for routine cleaning.
Forced-swim test consisted of two exposures to a water tank that does not permit escape [17]. During the first exposure, rats were placed into the tank for 15 min. The second exposure occurred 24 h afterwards and lasted 5 min during which rats behavior was videotaped and subsequently scored by a trained experimenter blind to the animal experimental group. Immobility time, expressed as duration (s), was analysed.
For SD, a resident-intruder protocol was followed [18]. Briefly, Long Evans male rats were used as dominant rats. Experimental subjects were Sprague–Dawley male rats (n = 10) that served as intruders by being transferred into the resident rat cage. Following the attack by the dominant leading to the defeat of the intruder, experimental subjects were physically isolated and left in visual, auditory and olfactory contact with the dominant for the following 30 min. Intruders were randomly exposed to different resident animals for 21 days. Control rats (n = 10) were introduced in a novel clean cage, without resident rat with the same protocol.
Learned Helplessness (LH) consisted of 20 min of inescapable foot-shock in a 40 min session. Twenty-four hours after inescapable shock, animals were tested for learned helplessness behavior. Testing consisted of 15 trials of 0.8 mA current lasting 60 s each with 24 s inter-trial time. Current was accompanied by a light clue to facilitate detection of the lever and animals were able to terminate the current by pressing the lever. Nine rats with more than 10 failures to terminate the current were selected for the experiment as LH; 8 rats with less than 5 failures were selected as the nLH controls. Antidepressant treatment with 25 mg/kg escitalopram (ESC) was carried out for 7 days.
Chronic mild stress (CMS) was performed as previously published [19]; the paradigm consisted in exposure to a chronic mild stress procedure for 7 weeks, while control rats were housed in separate rooms and had no contact with the stressed animals.
Detailed procedures are described in Supplementary Methods.
Gene expression analysis
Rat samples were processed as reported in [17] and in [19]. The present study compared publicly available hippocampal gene expression profiles measured in post-mortem tissue from subjects with MDD and matched unaffected controls (n = 19/group). The data have been made available to researchers world-wide and are available from the Gene Expression Omnibus (accession ID:GSE53987; www.ncbi.nlm.nih.gov/geo). Human microarray data were generated by hybridization to Affymetrix Human Genome U133 Plus 2.0 arrays (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE53987). Additional details are available in Supplementary Methods.
Bioinformatics
Transcriptional signatures were identified by means of an enhanced version of the rank-based classification method previously published [20, 21]. In the enhanced version used here, the original classification method has been extended with a genetic algorithm optimizer that automatically selects the method parameters (signature length and feature selection stringency) to maximize the accuracy of subject classification. In the case of the transcriptional signature reported in Table 1, rat samples coming from the three considered datasets (FSL, LH, and SD) have been merged together in order to obtain one single dataset to which the classification algorithm has been applied. The classification algorithm was then trained to perform a dichotomous classification, specifically to accurately classify control subjects, which consist of all the control samples from all the datasets, from the class of diseased ones, which includes all FSL, LH, and SD rats. A training version of the merged dataset was obtained by excluding samples difficult to classify, specifically FSL with MS, which share the FSL group as their controls, and SD rats, which were not separated into susceptible and resilient responders. The signatures resulting from the training were then used to run the algorithm in validation mode and to perform a classification on the complete dataset. In the case of the human transcriptional signature reported in Table 2, the classification algorithm has been applied to distinguish control samples from subjects with MDD without any pre-processing of the dataset.
To facilitate the interpretation of the biological processes underlying the phenomenon under investigation, we characterized the list of genes constituting the transcriptional signature by means of a network analysis technique implemented in NetWalker. Orthologs of the rat signature genes were obtained using Biomart (www.biomart.org). We used the protein-protein interaction network in the Netwalker suite (which includes HPRD, BIND, HomoMINT, Gene, and IntAct) as background network, the genes included in the signature as the set of genes of interest, and the negative log2 of the p-values computed during the probe filtering step in the signature identification algorithm (Kruskal–Wallis test) as gene scores. The NetWalker output consisted of a list of interactions of the background network and their scores as computed by the algorithm. We then extracted the connected sub-network containing all interactions above the empirically-selected threshold (n = 543 selected interactions for the rat and n = 448 for the human). The final list of genes was given by the genes included in such a sub-network (n = 272 genes for the rat and n = 274 for the human). The gene lists obtained from the transcriptional signature and from the network enrichment analysis were used to extract the most representative GO Biological Process terms (i.e., among statistically significant enriched terms, we manually discarded the most generic biological processes deemed as too general to provide specific information; the complete lists are reported in Tables S1, S3, S6, S8). For identifying and visualizing enriched GO terms, we used GOrilla and REVIGO tools. Pathway analysis was performed using ConsensusPathDB. Hypergeometric distribution with an adjusted p-value (FDR) threshold of 0.001 was applied to test GO and pathway enrichments, using HPRD gene list as reference background.
Detailed procedures are described in Supplementary Methods.
Comparison between pathway enrichment of the rat and human network lists
The pathway lists resulting from the enrichment analysis of the rat network list were ranked based on the negative logarithm of the p-value computed by the ConsensusPathDB tool. The same ranking operation was performed on the pathways list obtained for the human network gene list. In order to extract the top elements of each list for the rat-human comparison, rather than selecting an arbitrary threshold for p-values (i.e., p-value < 0.05), we adopted a more sensitive approach. Using the rrho package in R, we identified the top n1 elements in the rat pathway list and the top n2 in the human pathway list that resulted in the most significant overlap, where the significance was computed using the hypergeometric statistics. The value of n1 and n2 identified in this way were n1 = 106, n2 = 169, for an overlap of 67 terms (hypergeometric test p-value: 3.85 × 10−6).
Statistical analysis
The hypergeometric statistic and the formula of the Fisher’s method for combining p-values were computed in Excel 2010. The Kruskal Wallis test was implemented in Matlab as part of our signature identification method [20]. One-way and two-way ANOVA and Tukey post-hoc comparisons were performed with InVivoStat software [22]. The effects of CMS were analyzed with Student’s t-test. Significance was assumed for p-value < 0.05.
Results
Analysis of rat MDD models
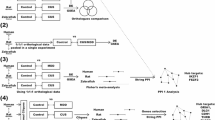
Different rat models of MDD sharing depression-like behaviors were investigated (Fig. 1a) within the GENDEP project. The analysis included FSL/FRL rats, FSL rats that experienced MS, LH/nLH, and SD/controls. In FRL/FSL/FSL + MS rats, depression-like responses were tested in the forced-swim test (Fig. S1). In LH rats, learned helplessness behavior was tested, thus the comparison was performed between rats showing resilient vs. susceptible response to stress exposure. In SD, experimental rats were exposed and defeated by a dominant male, while the controls were exposed to a novel cage.
a Schematic representation of the workflow. The hippocampi of rat models of MDD were subjected to transcriptional analysis. Gene expression data were analyzed with a recently established algorithm for signature-based clustering of expression profiles. Network analysis was then used to expand the list of signature genes with the help of protein-protein interactions from the literature and the gene list was functionally annotated. Subsequently, the same workflow tool was applied to transcriptomic data from human MDD patients and respective controls. b Classification of different models of MDD in rats. The graph represents sample classification by means of the rat transcriptional signature. Nodes represent rat samples and the length of connecting edges is proportional to the degree of similarity between the samples according to their transcriptional signature (short edges connect samples with similar signature, missing edges indicate negligible similarities). Based on the identified transcriptional signature (Table 1), all the models of MDD in rats were separated from control subjects except the SD model. Node colors and IDs indicate the considered rat models of MDD: FSL Flinders sensitive line, FSL + MS Flinders sensitive line with maternal separation, LH learned helplessness, SD social defeat, FRL Flinders resistant line, CTRL_LH control rats of the LH dataset, CTRL_SD control rats of the SD dataset
Identification of a transcriptomic signature in MDD models
Transcriptomic profiling was performed in the hippocampus as neural plasticity is altered in this brain region in MDD patients and in models. The gene expression data were analyzed with a new bioinformatic tool designed to identify with high sensitivity a transcriptomic signature able to classify profiles from controls and disease models (Fig. 1a). The tool is based on a method previously developed by our group that identifies signatures by examining the rank of each gene in the individual expression profiles [20, 21]. Interestingly, the tool identified a common transcriptomic signature comprising 70 probesets (Table 1) able to separate controls from FSL and LH rats (Fig. 1b). In contrast, SD or MS were not separated by the transcriptomic signature (Fig. 1b). These findings suggest this experimental design allowed the identification of the common molecular dysregulations sustaining the susceptible response to stress which is responsible for generating depression-like behaviors in the FSL and LH models, whereas this definite transcriptional response could not be detected in the SD group.
Functional annotation of the transcriptional signature showed that ErbB (q-value = 0.047) and TGF-β (q-value = 0.056) contribute to the differential expression (Table S1).
The algorithm used to identify the transcriptional signature has been designed to produce the shortest possible list while maintaining the highest classification accuracy. Therefore, we strived to expand the biological information present in a condensed form in the gene expression signature with the help of existing knowledge about biological processes as encoded in currently available protein interaction networks (Fig. 1a). We constructed a signature network of highly relevant proteins by carefully embedding the gene products belonging to the transcriptional signature (weighted by their p-values) into a literature-derived protein–protein interaction network; the identified signature network included 272 genes (Table S2). The functional annotation analysis highlighted the involvement of the ubiquitin-proteasome system (Fig. 2a; Tables S3–S4), which regulates cellular processes through degradation of specific functional proteins and controls synaptic plasticity, synaptic size, and energy metabolism. Functional annotation analysis also indicated the participation of the innate and adaptive immune responses, of ErbB receptors, of growth factor signaling, of leptin and insulin pathways, and of glutamatergic transmission (Fig. 2a; Tables S3–S4).
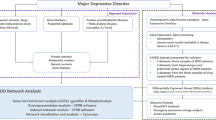
a Schematic graphs of over-represented Gene Ontology biological process terms in human or rat networks respectively. GO terms are represented as nodes, and the strongest GO term pair-wise similarities are designated as edges in the graph. GO terms overlapping in the human and rat networks are grouped. Colors and node size represent the p-values (low values in red and big size, high in green and small size). Complete lists of GO terms can be found in Tables S3 and S8). b Overlap between genes belonging to the rat and human transcriptional signatures and the genes where the top 10,000 most significant SNPs in the discovery dataset of the MDD GWAS are located (4177 SNPs falling within 333 genes [6])
Identification of a transcriptomic signature in MDD patients
While in healthy subjects allostatic plasticity mechanisms allow the development of an adaptive response to stressful experiences promoting resilience, in depressed patients changes in neural circuitry activated in response to stressors become “stuck” [2]. This impairment of structural and functional allostatic plasticity characterizing depressed patients, which is particularly evident in the hippocampus, is shared by animal models of depression [16]. Therefore, we next examined whether evidence could be found that the same transcriptional dysregulations characterizing rat models play a relevant role in the human disease. For this purpose we employed the same bioinformatics tool to analyze transcriptomic profiles of depressed patients from data made available to researchers world-wide (Fig. 1a, Fig. S2). Remarkably, we could identify a transcriptional signature able to separate human MDD patients from controls, based on the expression of 171 probesets (Table 2). The SCARA5 gene was common between the rat and human transcriptional signatures, with a combined p-value of 1.98 × 10−7 (Fisher’s method applied to the p-values for the gene computed in the two datasets data-sets using the Kruskal–Wallis test). The Fth1 gene, which is included in the rat transcriptional signature (Table 1), is reported to interact with SCARA5 (Fig. S3), providing further evidence of the potential relevance of this pathway. Available data suggest that the expression of SCARA5 and its interacting genes Fth1 and Ftl1 are modulated by antidepressant treatment in mouse MDD models (Table S5). In addition, these genes are altered after immobilization stress, a model of posttraumatic stress disorder, and in female mice exposed to chronic mild stress (Table S5).
The response to antidepressant treatment was then investigated. Seven day treatment with ESC of LH and nLH revealed statistically significant effects in both Fth1 and Ftl1 levels for pharmacological treatment (two-way ANOVA, F(1,27) = 29.19, p < 0.0001; F(1,27) = 18.20, p = 0.0002, respectively), for the response to stress (F(1,27) = 8.34, p = 0.0076; F(1,27) = 20.72, p = 0.0001, respectively), and for interaction (F(1,27) = 5.21, p = 0.031; F(1,27) = 7.21, p = 0.012, respectively) in the hippocampus. The expression of both genes was lower in LH rats (p = 0.0048 and p < 0.0001, respectively) and increased after ESC treatment (p < 0.0001 and p = 0.0002, respectively; Fig S4). Scara5 showed significant effects for the response to stress (F(1,27) = 28.03, p < 0.0001, Fig S4). In the prefrontal cortex, Scara5 revealed a significant effect for the response to stress (F(1,29) = 24.50, p < 0.0001), treatment (F(1,29) = 5.69, p = 0.024), and interaction (F(1,29) = 72.13, p < 0.0001); post-hoc analysis showed a significant reduction after antidepressant administration (p = 0.001, Fig S4). Treatment effects were detected in both Fth1 and Ftl1 (F(1,28) = 6.62, p = 0.016; F(1,29) = 31.65, p < 0.0001, respectively), as well as stress effects (F(1,28) = 54.43, p < 0.0001; F(1,29) = 45.49, p < 0.0001, respectively) and an interaction effect was observed in Fth1 (F(1,28) = 8.16, p = 0.008). Both genes were downregulated in LH rats (p = 0.022; p = 0.0006), whereas only Ftl1 was significantly upregulated by ESC treatment (p = 0.0043, Fig. 4S).
In order to further investigate whether dyregulations of the Scara5 pathway are involved in the molecular underpinning of different MDD models, we compared the mRNA levels of these genes in the dorsal and ventral hippocampus of rats exposed to 7 weeks of CMS as compared to the respective controls. In the ventral hippocampus, decreased Scara5 mRNA levels were detected in CMS rats (−18%, t(23.129) = 2.465, p = 0.022, Fig. S5). In the dorsal hippocampus the decrease amounted to a similar extent, although the difference was not statistically significant (−16%, t(16) = 1.896, p = 0.076, Fig. S5), possibly due to a lower number of samples. Fth1 and Ftl1 mRNA levels were not modulated by CMS exposure (Fig. S5).
We next looked at the overlap between the genes belonging to the rat and human transcriptional signatures and expanded signatures with data from [6]. By taking the 10,000 most significant SNPs reported for the discovery dataset, we could identify 333 genes mapped by a total 4177 SNPs. As shown in the Venn diagram (Fig. 2b, Fig. S6), three of the 333 genes were in common with the human signature genes (SPPL3, NHP2L1, and ZBP1) and one gene with the rat signature (COL12A1), suggesting that the associated polymorphisms may influence gene expression. Two out of three human genes were also significantly modulated in the rat data-set, although at p-values that didn’t reach the cut-off needed to be included into the signature (NHP2L1: p = 1.49 × 10−4; ZBP1: p = 0.0475). We also compared the overlap with data from the new GWAS study by [23] and identified SPPL3 and CACNA1E, CACNA2D1, CTNND1, MLF1, PAX6 as common genes with the human signature and expanded signature, respectively (Fig. S6).
Subsequently, the same workflow was followed for the analysis of the human signature (Fig. 1a): it was subjected to functional annotation (Table S6) and a protein interaction network was constructed, which included 274 genes (Table S7). A comparison between rat and human networks revealed 25 overlapping genes (Table S7), with a probability of an overlap of this magnitude corresponding to p = 6.80547 × 10−09 (Hypergeometric test). Functional analysis of the human signature network was then investigated (Fig. 2a, Tables S8-S9).
Identification of overlapping enriched pathways between human and rat signature networks
The enrichments in GO terms and pathways in the signature network constructed from depressed patients’ data were compared with enrichments derived from the signature network generated from rat models. Notably, the comparison between the top enriched pathways in rat (n = 169) and the top pathways in human (n = 106) identified a highly significant overlap of 67 terms (p = 3.85 × 10−6, Table 3). The common pathways confirmed the relevance of mechanisms observed in the GO enrichments, highlighting prominent roles for the ErbB pathway and for neurotrophin and other growth factors, chiefly FGF, VEGF, and IGF (Table 3). Other common pathways encompass the innate and adaptive immune responses; insulin and leptin signaling (Table 3).
Discussion
Our study aimed to identify a hippocampal transcriptional signature that would discriminate affected animals from controls in MDD models, in the direction of investigating multiple models to capture the common mechanisms underlying shared symptoms [11]. In hippocampus, the dysregulation of the adaptive allostatic response to stressors due to allostatic overload is linked with impaired plasticity, altered electrophysiological activity, and suppressed neurogenesis. Impairments due to allostatic overload are triggered by altered gene expression [16]. Our results show that rat models exhibiting many similar depression-like behaviors shared a common hippocampal transcriptional signature. Whereas the signature could distinguish FSL and LH models from controls, SD rats in our conditions did not display differential expression of the signature genes. This difference is likely due to the experimental design in this investigation. In our design, the FSL strain, a strain characterized by depressive-like behaviors, was the MDD model, whereas the FRL strain that does not show depressive-like behaviors served as control. Similarly, in the LH model, the rats were tested for helpless behavior and animals displaying a helpless response were the MDD model group, whereas those who did not (nLH) were the controls. Therefore, in these two models the MDD model was represented by the susceptible group, which was compared with the controls, represented by the resilient group. However, we did not separate the resilient and susceptible SD animals following encounter with a dominant rat [24]: they were treated as a single group in the comparison with the controls that were not exposed to the dominant rat. SD-resilient and susceptible mice have been shown to differ in transcriptional activity [24, 25], thus it is possible that in a different experimental design the transcriptional signature could discriminate between susceptible and unsusceptible SD rats. Overall, these results suggest that our transcriptional signature genes are specifically involved in the susceptibility to displaying depression-like responses and the lack of resilience skills. This interpretation could explain the significant pathway overlap that we detected between rat models and depressed patients, since resilience in the face of stress is a key aspect of a healthy response with respect to the development of MDD [16].
The rat and human transcriptional signatures shared the SCARA5 gene, which encodes for a scavenger receptor implicated in the innate immune response (Fig. S3) [26]. We speculate that SCARA5 differential expression contributes to the increased inflammatory response mediating the relationship between MDD and metabolic dysregulations [27]. SCARA5 is expressed at low but detectable levels in neuronal cells from human hippocampus (http://www.proteinatlas.org/ENSG00000168079-SCARA5/tissue). The discovery of SCARA5 association with MDD here reported is novel, although previous evidence for association with psychiatric disorders has been reported in a GWAS in schizophrenia [28] and in an infant trauma depression-like model, where the alteration is reversed in the rescue phenotype [29]. Moreover, increased transcriptome variability of SCARA5 was observed in temporal cortex of autistic subjects [30]. We can speculate that this pathway participates in the crucial role of scavenger receptors in microglia inflammatory response, which is recognized in Alzheimer’s disease [31, 32], but is also important in MDD [33]; further studies are needed to confirm our results.
We investigated the biological meaning of rat and human signatures by network reconstruction and gene functional annotation and identified numerous common pathways between MDD patients and FSL/LH models. Enrichment analysis suggested the pathophysiological significance of ErbB/neuregulin signaling. The ErbB family comprises four growth factor receptors that bind neuregulin and related EGF domain-containing proteins implicated in neural development, neurotransmission and synaptic plasticity [34]. Several ErbB/neuregulin members have been associated with MDD [34]. Genetic neuregulin variants were associated with a distinct pattern of the association between symptom severity and psychotic symptoms in depressed patients [35]. Reduced ErbB3 and neuregulin-1 levels have been detected in MDD patients in brain and leukocytes [36, 37]. Neuregulin-1 has also been implicated in the response to antidepressant treatment in genetic analyses in humans and in animal models [38, 39]. Our results supporting the hypothesis that altered levels of neuregulin/ErbB members in allostatic overload conditions play a role in the development of depressive behaviors are corroborated by findings in chronic mild stress rats, where neuregulin/ErbB signaling shows a dysregulation reversed by antidepressant treatment [40].
In addition, the relevance of other growth factor pathways was demonstrated in rat and human networks. In particular, enriched neurotrophic factor pathways have been identified, consistent with a large body of literature showing that BDNF levels are altered in depressed patients [41], that the BDNFVal66Met polymorphism interacting with stress exposure is associated with MDD [42] and that neurotrophic signaling is instrumental for antidepressant efficacy [43]. These findings confirm the hypothesis that stress decreases BDNF expression, thus impairing neurogenesis and neural plasticity activated in the resilient response to stress [16, 44, 45]. Moreover, we discovered dysregulations in FGF, IGF-1, and VEGF, which have been recently associated with MDD [46, 47]. It is especially suggestive that work from the Pritzker Neuropsychiatric Disorder Research Consortium allowed the important discovery that FGF, originally implicated in neural development, plays a crucial role in neurogenesis and in neuroplasticity in adulthood, suggesting a neuromodulatory function in anxiety-like and depression-like behaviors [47, 48], providing support for our findings. In addition, altered levels of FGF family members were revealed in MDD patients, both in blood and in brain [49]. Altered levels of IGF-1, a neurotrophic, neurogenetic and neuroprotective factor, were repeatedly reported in MDD patients, although with conflicting results in the direction of change [46]. In MDD models, IGF administration produced antidepressant-like effects [50], in line with our findings. VEGF exerts direct and indirect neurogenetic effects that have been linked with antidepressant activity [51]. A role in MDD is also supported by the data showing that peripheral VEGF levels are significantly elevated in MDD patients [46, 52]. Overall, our findings corroborate the hypothesis suggested by a body of recent results that modulations of the NGF, FGF, IGF, and VEGF pathways contribute to the molecular underpinning of MDD probably by hampering the adaptive neuroplasticity responses associated with resilient responses to stressors.
We observed a considerable impact on metabolic pathways, including insulin and leptin signaling. Compelling evidence supports the involvement of a metabolic component in the pathophysiology of MDD. A high degree of co-morbidity between MDD and type-2 diabetes was demonstrated [27], with insulin resistance as a connecting link [53, 54]. In agreement with our findings, elevated plasma insulin levels were observed in depressed patients [55, 56], the hypoglycemic agent pioglitazone displayed antidepressant efficacy [57] and decreased insulin signaling induced depression-like behaviors in rats [58]. In line with our results, altered leptin levels have been repeatedly observed in MDD patients [56] and leptin-mediated antidepressant-like effects reported in animal models [2, 54, 59].
It has been proposed that the mechanistic link in the bi-directional relationship between MDD and T2DM is the activation of the inflammatory response associated with both disorders [27]. Our study revealed a transcriptional dysregulation involving immune responses both in patients and in animal models. Moreover, an affected gene (ZBP1) in common with the recent GWAS for MDD [6] is reported to play a role in the innate immune response. Thus, our data are consistent with the hypothesis that there is a link between metabolic dysregulations and inflammatory components in MDD.
Limitations of this study are that hippocampal sub-regions could not be differentiated, whereas distinct plasticity responses are evoked in hippocampal sub-fields. Moreover, the experimental design did not discriminate SD-susceptible and resilient rats, preventing an appropriate comparison of this model. The number of patients was relatively small and no characterization was possible. Therefore, testing the relevance of the identified genes and pathways in additional animal models and human subjects is required in future experiments. Crucially, further experiments are necessary to elucidate the role of SCARA5 in MDD pathophysiology, here revealed for the first time.
In conclusion, the present results demonstrate that both MDD patients and the FSL and LH models show dysregulations in growth factor signaling, in metabolic pathways, and in inflammatory responses. MDD arises when the resilience of healthy brains is lost and instead of promoting neural circuitry adaptation able to cope with stressors, a maladaptive plasticity takes place [2]. Compelling evidence supports the crucial contribution provided by inappropriate activation of these pathways through gene expression regulation and epigenetic mechanisms finally impairing the mechanisms for a resilient response to stress exposure and leading to allostatic overload [44, 53, 54]. Lastly, and speculatively, the molecular alterations identified in the present study could contribute to the identification of molecular alterations characterizing MDD biotypes recently identified by functional magnetic resonance imaging [60].
References
Mrazek Da, Hornberger JC, Altar CA, Degtiar I. A review of the clinical, economic, and societal burden of treatment-resistant depression: 1996–2013. Psychiatr Serv. 2014;65:977–87.
McEwen BS. Neurobiological and systemic effects of chronic stress. Chronic Stress. 2017;1:247054701769232.
CONVERGE consortium. Sparse whole-genome sequencing identifies two loci for major depressive disorder. Nature. 2015;523:588–91.
Flint J, Kendler KS. The genetics of major depression. Neuron. 2014;81:484–503.
Major Depressive Disorder Working Group of the Psychiatric GWAS Consortium, Ripke S, Wray NR, Lewis CM, Hamilton SP, Weissman MM, et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol Psychiatry. 2013;18:497–511.
Hyde CL, Nagle MW, Tian C, Chen X, Paciga SA, Wendland JR, et al. Identification of 15 genetic loci associated with risk of major depression in individuals of European descent. Nat Genet. 2016;48:1–9.
Overstreet DH, Friedman E, Mathé AA, Yadid G. The Flinders Sensitive Line rat: a selectively bred putative animal model of depression. Neurosci Biobehav Rev. 2005;29:739–59.
Vollmayr B, Gass P. Learned helplessness: unique features and translational value of a cognitive depression model. Cell Tissue Res. 2013;354:171–8.
Piubelli C, Carboni L, Becchi S, Mathé AA, Domenici E. Regulation of cytoskeleton machinery, neurogenesis and energy metabolism pathways in a rat gene-environment model of depression revealed by proteomic analysis. Neuroscience. 2011;176:349–80.
Buwalda B, Kole MHP, Veenema AH, Huininga M, Boer SF, de, Korte SM, et al. Long-term effects of social stress on brain and behavior: a focus on hippocampal functioning. Neurosci Biobehav Rev. 2005;29:83–97.
Akil H, Gordon J, Hen R, Javitch J, Mayberg H, McEwen B, et al. Treatment resistant depression: a multi-scale, systems biology approach. Neurosci Biobehav Rev. 2018;84:272–88.
Hervé M, Bergon A, Guisquet A-MLe, Leman S, Consoloni J-L, Fernandez-Nunez N, et al. Translational identification of transcriptional signatures of major depression and antidepressant response. Front Mol Neurosci. 2017;10:248.
Labonté B, Engmann O, Purushothaman I, Menard C, Wang J, Tan C, et al. Sex-specific transcriptional signatures in human depression. Nat Med. 2017;23:1102–11.
Wingo AP, Velasco ER, Florido A, Lori A, Choi DC, Jovanovic T, et al. Expression of the PPM1F gene is regulated by stress and associated with anxiety and depression. Biol Psychiatry. 2018;83:284–95.
Sheline YI, Mittler BL, Mintun MA. The hippocampus and depression. Eur Psychiatry. 2002;17:300–5.
McEwen BS, Bowles NP, Gray JD, Hill MN, Hunter RG, Karatsoreos IN, et al. Mechanisms of stress in the brain. Nat Neurosci. 2015;18:1353–63.
Blaveri E, Kelly F, Mallei A, Harris K, Taylor A, Reid J, et al. Expression profiling of a genetic animal model of depression reveals novel molecular pathways underlying depressive-like behaviours. PLoS One. 2010;5:e12596.
Carboni L, Piubelli C, Pozzato C, Astner H, Arban R, Righetti PGG, et al. Proteomic analysis of rat hippocampus after repeated psychosocial stress. Neuroscience. 2006;137:1237–46.
Luoni A, Macchi F, Papp M, Molteni R, Riva MA. Lurasidone exerts antidepressant properties in the chronic mild stress model through the regulation of synaptic and neuroplastic mechanisms in the rat prefrontal cortex. Int J Neuropsychopharmacol. 2015;18:1–12.
Lauria M. Rank-based transcriptional signatures. Syst Biomed. 2013;1:228–39.
Lauria M, Moyseos P, Priami C. SCUDO: a tool for signature-based clustering of expression profiles. Nucleic Acids Res. 2015;43:W188–92.
Clark RA, Shoaib M, Hewitt KN, Stanford SC, Bate ST. A comparison of InVivoStat with other statistical software packages for analysis of data generated from animal experiments. J Psychopharmacol. 2012;26:1136–42.
Wray NR, Ripke S, Mattheisen M, Trzaskowski M, Byrne EM, Abdellaoui A, et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat Genet. 2018;50:668–81.
Krishnan V, Han M-HH, Graham DL, Berton O, Renthal W, Russo SJ, et al. Molecular adaptations underlying susceptibility and resistance to social defeat in brain reward regions. Cell. 2007;131:391–404.
Bagot RCC, Cates HMM, Purushothaman I, Lorsch ZSS, Walker DMM, Wang J, et al. Circuit-wide transcriptional profiling reveals brain region-specific gene networks regulating depression susceptibility. Neuron. 2016;90:969–83.
Ojala JRM, Pikkarainen T, Elmberger G, Tryggvason K. Progressive reactive lymphoid connective tissue disease and development of autoantibodies in scavenger receptor A5-deficient mice. Am J Pathol. 2013;182:1681–95.
Stuart MJ, Baune BT. Depression and type 2 diabetes: inflammatory mechanisms of a psychoneuroendocrine co-morbidity. Neurosci Biobehav Rev. 2012;36:658–76.
Xu C, Aragam N, Li X, Villla EC, Wang L, Briones D, et al. (2013). BCL9 and C9orf5 are associated with negative symptoms in schizophrenia: meta-analysis of two genome-wide association studies. PLoS One 8. e51674
Rincón-Cortés M, Barr GA, Mouly AM, Shionoya K, Nuñez BS, Sullivan RM. Enduring good memories of infant trauma: rescue of adult neurobehavioral deficits via amygdala serotonin and corticosterone interaction. Proc Natl Acad Sci. 2015;112:881–6.
Garbett K, Ebert PJ, Mitchell A, Lintas C, Manzi B, Mirnics K, et al. Immune transcriptome alterations in the temporal cortex of subjects with autism. Neurobiol Dis. 2008;30:303–11.
Cornejo F, Vruwink M, Metz C, Muñoz P, Salgado N, Poblete J, et al. (2017). Scavenger receptor-A deficiency impairs immune response of microglia and astrocytes potentiating Alzheimer’s disease pathophysiology. Brain Behav. Immun. https://doi.org/10.1016/j.bbi.2017.12.007.
Frenkel D, Wilkinson K, Zhao L, Hickman SE, Means TK, Puckett L, et al. Scara1 deficiency impairs clearance of soluble amyloid-β by mononuclear phagocytes and accelerates Alzheimer’s-like disease progression. Nat Commun. 2013;4:1–9.
Brites D, Fernandes A. Neuroinflammation and depression: microglia activation, extracellular microvesicles and microRNA dysregulation. Front Cell Neurosci. 2015;9:1–20.
Mei L, Nave KA. Neuregulin-ERBB signaling in the nervous system and neuropsychiatric diseases. Neuron. 2014;83:27–49.
Bousman CA, Potiriadis M, Everall IP, Gunn JM. Effects of neuregulin-1 genetic variation and depression symptom severity on longitudinal patterns of psychotic symptoms in primary care attendees. Am J Med Genet Part B Neuropsychiatr Genet. 2014;165:62–67.
Aston C, Jiang L, Sokolov BP. Transcriptional profiling reveals evidence for signaling and oligodendroglial abnormalities in the temporal cortex from patients with major depressive disorder. Mol Psychiatry. 2005;10:309–22.
Milanesi E, Minelli A, Cattane N, Cattaneo A, Mora C, Barbon A, et al. ErbB3 mRNA leukocyte levels as a biomarker for major depressive disorder. BMC Psychiatry. 2012;12:145.
Wang N, Zhang GF, Liu XY, Sun HL, Wang XM, Qiu LL, et al. Downregulation of neuregulin 1-ErbB4 signaling in parvalbumin interneurons in the rat brain may contribute to the antidepressant properties of ketamine. J Mol Neurosci. 2014;54:211–8.
Biernacka JM, Sangkuhl K, Jenkins G, Whaley RM, Barman P, Batzler A, et al. The International SSRI Pharmacogenomics Consortium (ISPC): a genome-wide association study of antidepressant treatment response. Transl Psychiatry. 2015;5:e553.
Dang R, Cai H, Zhang L, Liang D, Lv C, Guo Y, et al. Dysregulation of Neuregulin-1/ErbB signaling in the prefrontal cortex and hippocampus of rats exposed to chronic unpredictable mild stress. Physiol Behav. 2015;154:145–50.
Molendijk ML, Spinhoven P, Polak M, Bus BAA, Penninx BWJH, Elzinga BM. Serum BDNF concentrations as peripheral manifestations of depression: evidence from a systematic review and meta-analyses on 179 associations (N = 9484). Mol Psychiatry. 2014;19:791–800.
Hosang GM, Shiles C, Tansey KE, McGuffin P, Uher R. Interaction between stress and the BDNF Val66Met polymorphism in depression: a systematic review and meta-analysis. BMC Med. 2014;12:7.
Björkholm C, Monteggia LM. BDNF—a key transducer of antidepressant effects. Neuropharmacology. 2016;102:72–9.
Gray JD, Milner TA, McEwen BS. Dynamic plasticity: the role of glucocorticoids, brain-derived neurotrophic factor and other trophic factors. Neuroscience. 2013;239:214–27.
Autry AE, Monteggia LM. Brain-derived neurotrophic factor and neuropsychiatric disorders. Pharmacol Rev. 2012;64:238–58.
Sharma AN, Costa E, Silva BFB, Da, Soares JC, Carvalho AF, Quevedo J. Role of trophic factors GDNF, IGF-1 and VEGF in major depressive disorder: a comprehensive review of human studies. J Affect Disord. 2016;197:9–20.
Turner CA, Eren-Koçak E, Inui EG, Watson SJ, Akil H. Dysregulated fibroblast growth factor (FGF) signaling in neurological and psychiatric disorders. Semin Cell Dev Biol. 2016;53:136–43.
Aurbach EL, Inui EG, Turner CA, Hagenauer MH, Prater KE, Li JZ, et al. Fibroblast growth factor 9 is a novel modulator of negative affect. Proc Natl Acad Sci. 2015;112:11953–8.
Wu C, Tseng P, Chen Y, Tu K, Lin P. Significantly higher peripheral fibroblast growth factor-2 levels in patients with major depressive disorder: a preliminary meta-analysis under MOOSE guidelines. Medicine. 2016;95:e4563.
Burgdorf J, Zhang X, Colechio EM, Ghoreishi-Haack N, Gross A, Kroes RA, et al. Insulin-like growth factor I produces an antidepressant-like effect and elicits N-methyl-D-aspartate receptor independent long-term potentiation of synaptic transmission in medial prefrontal cortex and hippocampus. Int J Neuropsychopharmacol. 2016;19:pyv101.
Fournier NM, Duman RS. Role of vascular endothelial growth factor in adult hippocampal neurogenesis: implications for the pathophysiology and treatment of depression. Behav Brain Res. 2012;227:440–9.
Carvalho AF, Köhler CA, McIntyre RS, Knöchel C, Brunoni AR, Thase ME, et al. Peripheral vascular endothelial growth factor as a novel depression biomarker: a meta-analysis. Psychoneuroendocrinology. 2015;62:18–26.
Nasca C, Xenos D, Barone Y, Caruso A, Scaccianoce S, Matrisciano F, et al. L-acetylcarnitine causes rapid antidepressant effects through the epigenetic induction of mGlu2 receptors. Proc Natl Acad Sci. 2013;110:4804–9.
Bigio B, Mathé AAA, Sousa VCC, Zelli D, Svenningsson P, McEwen BSS, et al. Epigenetics and energetics in ventral hippocampus mediate rapid antidepressant action: implications for treatment resistance. Proc Natl Acad Sci. 2016;113:7906–11.
Domenici E, Willé DR, Tozzi F, Prokopenko I, Miller S, McKeown A, et al. Plasma protein biomarkers for depression and schizophrenia by multi analyte profiling of case-control collections. PLoS One. 2010;5:e9166.
Lamers F, Bot M, Jansen R, Chan M, Cooper J, Bahn S, et al. (2016). Serum proteomic profiles of depressive subtypes. Nat Publ Gr. 6. e51674
Kemp DE, Ismail-Beigi F, Ganocy SJ, Conroy C, Gao K, Obral S, et al. Use of insulin sensitizers for the treatment of major depressive disorder: a pilot study of pioglitazone for major depression accompanied by abdominal obesity. J Affect Disord. 2012;136:1164–73.
Grillo CA, Piroli GG, Kaigler KF, Wilson SP, Wilson MA, Reagan LP. Downregulation of hypothalamic insulin receptor expression elicits depressive-like behaviors in rats. Behav Brain Res. 2011;222:230–5.
Stieg MR, Sievers C, Farr O, Stalla GK, Mantzoros CS. Leptin: A hormone linking activation of neuroendocrine axes with neuropathology. Psychoneuroendocrinology. 2015;51:47–57.
Drysdale AT, Grosenick L, Downar J, Dunlop K, Mansouri F, Meng Y, et al. Resting-state connectivity biomarkers define neurophysiological subtypes of depression. Nat Publ Gr. 2016. https://doi.org/10.1038/nm.4246.
Acknowledgements
We thank Dr. Stewart A. Bates for help with gene expression experiments and for critical reading of the manuscript and Dr. Roberto Arban for help with the SD experiments and helpful discussions.
Funding
This work was part of a project funded by the European Commission that combined large-scale clinical pharmacogenomic studies on depressed patients with preclinical investigations on animal models of disease, focusing on treatment with pro-serotonergic and pro-noradrenergic antidepressants, called ‘Genome-based therapeutic drugs for depression (GENDEP)’, contract number LSHB-CT-2003-503428. The work was also supported by the University of Bologna (RFO 2014) to L. Carboni and the Swedish Medical Research Council to AAM (10414). The funding bodies had no role in the design of the study, collection and analysis of data and decision to publish. L. Carboni, MR, ED, and L. Caberlotto were GlaxoSmithKline employees when this work was started. During the past year, Dr Malki received income from and has been an employee and stockholder of Eli Lilly and UCB Celltech. The authors declare no conflicts of interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Carboni, L., Marchetti, L., Lauria, M. et al. Cross-species evidence from human and rat brain transcriptome for growth factor signaling pathway dysregulation in major depression. Neuropsychopharmacol 43, 2134–2145 (2018). https://doi.org/10.1038/s41386-018-0117-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41386-018-0117-6
This article is cited by
-
The effect and mechanism of Jiao-tai-wan in the treatment of diabetes mellitus with depression based on network pharmacology and experimental analysis
Molecular Medicine (2021)
-
Treatment-emergent and trajectory-based peripheral gene expression markers of antidepressant response
Translational Psychiatry (2021)