Abstract
Newborns are highly susceptible to pathogenic infections with significant worldwide morbidity possibly due to an immature immune system. Recently, we reported that Propionibacterium strain, P. UF1, isolated from the gut microbiota of preterm infants, induced the differentiation of bacteria-specific Th17 cells. Here, we demonstrate that P. UF1 significantly increased the number of protective Th17 cells and maintained IL-10+ regulatory T cells (Tregs) in newborn mice. In addition, P. UF1 protected mice from intestinal Listeria monocytogenes (L. m) infection. P. UF1 also functionally sustained the gut microbiota and induced critical B vitamin metabolites implicated in the regulation of T cell immunity during L. m intestinal infection. Transcriptomic analysis of P. UF1-induced Th17 cells revealed genes involved in the differentiation and regulation of these cells. These results illustrate the potency of P. UF1 in the enhancement of neonatal host defense against intestinal pathogen infection.
Similar content being viewed by others
Introduction
Despite advances, approximately one million neonates under 6 months of age succumb to infections annually.1 Studies demonstrate that the susceptibility of newborns to infections may be due to immature adaptive immune responses to pathogenic challenges.1 Available vaccines provide insufficient protection against pathogenic microorganisms, or require multiple injections to sustain protective immunity.2 The mechanisms for this lack of protection remain elusive. However, there is some evidence that the lack of protection may partially be due to low numbers of protective T cells in newborns compared to adults.3 Thus, mounted T cell immunity may be insufficient to clear the infection leading to tissue injury. Moreover, the development of protective immunity is naturally slow and requires a myriad of additional stimulatory factors, including innocuous dietary antigens and a repertoire of critical microbial metabolites to induce a protective immunity.3,4,5,6 This gradual process of immune development may allow for deterioration due to invading pathogens inducing inflammation that can shut down functional T cell homeostasis against infections. Data demonstrate the critical role of a stable microbial consortium in enhancing immune resistance to various enteric pathogens in infants compared to neonates. This may account in part for the increased susceptibility of neonates to particular foodborne pathogens if the commensal flora is not well established.7,8 Understanding how immunity evolves to protect newborns against life-threatening infections is key to developing novel approaches, including the administration of particular bacteria with bifidogenic properties such as P. UF19,10 to potentially shorten the slow period of neonatal adaptive immunological development,11 resulting in the acceleration of protective T cell differentiation. This may help control and prevent pathological proinflammatory conditions in neonates, including necrotizing enterocolitis (NEC).12 With this in mind, we demonstrate that the administration of our newly discovered P. UF1 bacterium to newborn mice accelerates the neonatal protective CD4+ T cell frequency against intestinal infection.
Results
Acceleration of neonatal T cell immunity by P. UF1
Recently, we reported the discovery of a novel Propionibacterium strain, dubbed P. UF1, capable of inducing the differentiation of dihydrolipoamide acetyltransferase (DlaT)-specific Th17 cells.9 DlaT protein was isolated from bacterial surface layer.9 As an enzyme, DlaT is implicated in bacterial pyruvate decarboxylation that links glycolysis to the citric acid cycle.13 However, DlaT can also be synthesized by other yet-to-be identified mechanism(s) on the S-layer to activate antigen-specific Th17 cells. Induced protective DlaT-specific Th17 cells were highly regulated by P. UF1-induced IL-10+ Tregs to resolve L. m infection in adult mice.9,10 P. UF1 also significantly controlled deleterious signals (e.g., iNOS, IL-1β) involved in necrotizing enterocolitis (NEC)-like injury by regulating Th17 cells and sustaining IL-10+ Tregs, which protected the newborn mice, whose mothers were orally pretreated with P. UF1,9 against NEC manifestation. To investigate the effects of gavaging pregnant dams with P. UF1, which initially resulted in an increased number of Th17 cells and Tregs in newborn mice,9 we first constructed an antibiotic-resistant strain (CmR P. UF1) by insertion of chloramphenicol resistant gene (CmR) into the P. UF1 chromosome (Supplementary Fig. 1a). The possibility of P. UF1-transfer into newborn mice was then discerned by gavaging pregnant mice with CmR P. UF1 (109 CFU/mouse/twice a week). Obtained data demonstrated the presence of P. UF1 in the fecal contents of the newborn mice as early as 5 days after they were delivered by their mothers (Supplementary Fig. 1b). These data may explain the regulated immune protection of newborn mice induced by P. UF1 previously subjected to experimental NEC-like injury.9 Furthermore, to demonstrate any maternal effects on the newborn mouse immunity against pathogen infection, newborn mice without any P. UF1, or PBS oral treatment were kept with their mothers that were gavaged with P. UF1, or PBS for 3 weeks. To limit L. m infection to the gut, these groups of mice were orally infected with ΔactA L. m (108 CFU/mouse). ΔactA L. m lacking the ActA polypeptide cannot interact with actin filaments and is, consequently, nonmotile to spread in the periphery of the mice.9 Interestingly, enhanced Th17 cells and Tregs were induced against ΔactA L. m in mice that were initially delivered by dams gavaged with P. UF1 when compared to the other group (Supplementary Fig. 1c, d). Additionally, the clearance of ΔactA L. m after 4 days demonstrated a significant trend of ΔactA L. m clearance in this group compared to control group (Supplementary Fig. 1e). These data indicate that it may be of significance if mothers and newborn mice would be continuously gavaged with P. UF1 to increase the enhancement of T cell immunity in newborn mice. To expand on this notion and our initial observation,9 we hypothesized that the slow process of T cell differentiation in the neonatal host may be accelerated by P. UF1 to fortify regulated-intestinal T cell response in steady state and during intestinal L. m infection. Thus, we first gavaged groups of pregnant C57BL/6 mice with P. UF1, ΔdlaT P.UF1 (109 CFU/mouse/weekly), or PBS. After birth, the same regimen was used to gavage newborn mice of the respective dams for 8 weeks. Newborn mice were then euthanized at 1, 2, 4, 6, or 8 weeks to study CD4+ T cell response over time (Fig. 1a). Data demonstrated a significant increase in regulated CD4+ T cells, particularly IL-10+ Th17 cells induced by P. UF1 compared to other groups (Fig. 1b, c). Notably, the increased frequency of IL-10+ Th17 cells was followed by IL-10+ IFNγ+ Th1 cells (Fig.1c). Although the frequency of FoxP3+ Tregs was not increased, the numbers of functional IL-10+ TGFβ+ Tregs were significantly enhanced in newborn mice gavaged with P. UF1 for 8 weeks (Fig. 1c). These data may highlight the potency of P. UF1 to increase the numbers of regulated functional colonic CD4+ T cells in the steady state.
P. UF1 accelerates the regulation of neonatal immunity. a Schematic depiction of feeding schedule of newborn mice. C57BL/6 female mice (dams) were gavaged once a week with P. UF1, ΔdlaT P. UF1 (109 CFU/dam), or PBS during pregnancy. After birth, newborn mice of the respective dams were gavaged once a week with P. UF1, ΔdlaT P. UF1 (107 CFU/mouse), or PBS during breastfeeding. After weaning, mice continued to receive P. UF1, ΔdlaT P. UF1 (108 CFU/mouse), or PBS once a week. Mice were euthanized at 1, 2, 4, 6, or 8 weeks old to analyze colonic CD4+ T cell responses by FACS. b Representative flow plots for colonic IL-10+ Th17, IL-10+ TGFβ+ Tregs and IL-10+ IFNγ+ Th1 cells from mice gavaged with P. UF1, ΔdlaT P. UF1, or PBS for 8 weeks. c Frequency (top) and total counts (bottom) of Th17, IL-10+ Th17, IL-10+ TGFβ+ Treg and IL-10+ IFNγ+ Th1 analyzed from 1- (n = 3–4), 2- (n = 4–5), 4- (n = 6–7), 6- (n = 4–5), and 8- (n = 4–5) week-old mice. Data are from 1 (2-, 6- and 8-week-old mice) experiment or representative of 2 (1- and 4-week-old mice) independent experiments. Data are represented as mean ± SEM; * p < 0.05, ** p < 0.01, *** p < 0.001, ANOVA plus Tukey post-test
Regulation of gut homeostasis by P. UF1 during intestinal Listeria infection
To limit tissue damage, activation of protective T cell responses requires regulatory signals to suppress pathogenic inflammation.14 To once again demonstrate the ability of P. UF1 in the regulation of Th17 cells, along with the maintenance of IL-10+ Tregs in a pathogen-induced inflammatory condition, pregnant C57BL/6 were gavaged with P. UF1, ΔdlaT P. UF1 (109 CFU/mouse), or PBS. Once these dams had delivered their newborns, groups of the newborn mice were then gavaged with P. UF1, ΔdlaT P. UF1, or PBS. These groups of mice were orally infected with ΔactA L. m (108 CFU/mouse 3 weeks of age, or 109 CFU/mouse 4–8 weeks of age) at 3, 4, 6, and 8 weeks of age (Fig. 2a). Data revealed that the protective regulatory Th17 cell response was induced by P. UF1 against ΔactA L. m as early as 3 weeks (Fig. 2b, c and Supplementary Fig. 2a). Protective Th17 cells, along with Th1 cells, both potentially controlled by IL-10+ Tregs9 (Fig. 2b, c and Supplementary Fig. 2a), significantly decreased ΔactA L. m infection within 3, or 4 days, while the other groups of mice showed higher burdens of ΔactA L. m (Fig. 2d and Supplementary Fig. 2b). These results demonstrate P. UF1’s ability to enhance protective T cell response against L. m infection.
T helper cells induced by P. UF1 protect neonatal mice against L. m infection. a Schematic depiction of the feeding schedule of newborn mice. Pregnant C57BL/6 dams were gavaged with P. UF1, ΔdlaT P. UF1 (109 CFU/dam) or PBS once a week. After birth, newborn mice of the respective dams were gavaged once a week either with P. UF1, ΔdlaT P. UF1 (107 CFU/mouse), or PBS. After weaning, mice continued to receive P. UF1, ΔdlaT P. UF1 (108 CFU/mouse), or PBS once a week. Mice were then infected with ΔactA L. m (109 CFU/mouse) at 4, 6, or 8 weeks old and euthanized 1 week after infection to analyze colonic CD4+ T cell responses by FACS. b Representative flow plots for IL-10+ Th17, IL-10+ TGFβ+ Treg and IL-10+ IFNγ+ Th1 cells analyzed from 8-week-old mice infected with ΔactA L. m. c Frequency (top) and total counts (bottom) of colonic Th17, IL-10+ Th17, IL-10+ TGFβ+ Treg and IL-10+ IFNγ+ Th1 cells, analyzed from 4- (ΔactA L. m, n = 5; ΔactA L. m + P. UF1, n = 7; ΔactA L. m + ΔdlaT P. UF1, n = 9), 6- (ΔactA L. m, n = 4; ΔactA L. m + P. UF1, n = 8; ΔactA L. m + ΔdlaT P. UF1, n = 5), and 8- (ΔactA L. m, n = 5; ΔactA L. m + P. UF1, n = 8; ΔactA L. m + ΔdlaT P. UF1, n = 10) week-old mice. d Pathogen burdens in the feces of mice infected with ΔactA L. m. e 8-week-old mice (n = 4–5) gavaged with P. UF1, ΔdlaT P. UF1 or PBS were intraperitoneally injected with anti-IL-17A neutralizing antibody (100 μg/mouse/time, days −6 and −2) and then infected with ΔactA L. m (109 CFU/mouse, day 0). Bacterial burdens were monitored on days 1–4 post infection. Dash lines indicate the limit of pathogen detection. f Groups of Rag1-/- mice were gavaged with P. UF1 (4 gavages, 109 CFU/mouse) or PBS, and then orally infected with ΔactA L. m (109 CFU/mouse). ΔactA L. m clearance was determined on days 1–5 post infection. Data are from 1 experiment (e, f), or representative of 3 (a–d) independent experiments. Data are presented as mean ± SEM; * p < 0.05, ** p < 0.01, *** p < 0.01, ANOVA plus Tukey post-test (c, d) or Kruskal–Wallis test (d–f)
Recently, we9,10 and others15,16 demonstrated the implication of Th17 cells in the clearance of pathogen infection. To further demonstrate the role of P. UF1-induced Th17 cells against ΔactA L. m, we neutralized IL-17A by intraperitoneal injection (ip) of anti-IL-17A antibodies (200 μg/mouse) in mice that were initially gavaged with P. UF1, ΔdlaT P. UF1 (109 CFU/mouse), or PBS for 8 weeks. These groups of mice were then infected with ΔactA L. m (109 CFU/mouse). Data demonstrated that neutralizing IL-17A resulted in the persistence of ΔactA L. m infection in all groups of mice illuminating an important role of IL-17A in the clearance of ΔactA L. m (Fig. 2e). Notably, such pathogen persistence was significantly higher on days 2 and 3 in mice gavaged with P. UF1 that received anti-IL-17A neutralizing antibodies, potentially indicating the prominent role of Th17 cells induced by this bacterium controlling ΔactA L. m infection (Fig. 2e). To further emphasize the role of T cell responses against pathogen infection, we then employed Rag1-/- mice exhibiting no mature T and B cells.9 Here, groups of Rag1-/- mice were gavaged either with P. UF1(109 CFU/mouse/every other day), or PBS four times. Subsequently, mice were orally infected with ΔactA L. m (109 CFU/mouse). Data demonstrated that in both groups of Rag1-/- mice, ΔactA L. m clearance was not obvious and ΔactA L. m infection persisted even more in P. UF1-gavaged mice, once again highlighting the protective role of T cells, particularly P. UF1-induced Th17 cells, against ΔactA L. m infection (Fig. 2f).
Impact of P. UF1 on gut microbiota and the induced metabolites during Listeria infection
Intestinal infection may manifest in gut dysbiosis and pathogen growth that can induce inflammatory changes leading to deterioration of intestinal homeostasis and eventual tissue damage.17 Accordingly, we investigated the status of gut microbiota before and after ΔactA L. m infection in newborn mice gavaged with P. UF1 for 8 weeks compared with other groups. Here, the principal coordinate analysis (PcoA) of mice gavaged with P. UF1, ΔdlaT P. UF1, or PBS and subsequently infected with ΔactA L. m showed significant separation in their gut microbiota (PERMANOVA p = 0.036), which was not evident before infection (PERMANOVA p = 0.141) (Supplementary Fig. 3a, b). To identify intestinal commensal species contributing to such PcoA-differences, we then performed linear discriminant analysis effect size (LEfSe) among the infected groups with ΔactA L. m. Analyzed data demonstrated that the family of Lactobacillaceae, particularly Lactobacillus genera, and Clostridiaceae producing short-chain fatty acids (SCFA),18 were highly enriched in mice gavaged with P. UF1 while Prevotella was dominantly represented in the microbiota of PBS-gavaged group after infection (Fig. 3c and Supplementary Fig. 3b). Thus, we focused on the significant species identified through LEfSe analyses of mice infected with ΔactA L. m, and compared the relative abundance of those species before and after infection. Data revealed that P. UF1 sustained beneficial microbes (e.g., Lactobacillus and Peptostreptococcaceae) and even propagated Clostridiaceae after infection (Fig. 3a, b, and Supplementary Fig. 3c). Accordingly, lowering pH and increasing propionic acid level in the proximity of beneficial bacteria could potentially alter the growth of pathogenic bacteria. This aspect may require further investigation, as data illuminated the effects of diverse gut bacteria (e.g., Clostridium species) reducing L. m infectivity19 in mice. This indicates that gut microbes may function as the first line of protection that potentially controls pathogen growth20 through various bacterial metabolites, including propionate.21
P. UF1 modulates the microbiota composition during ΔactA L. m infection. Pregnant C57BL/6 dams were gavaged weekly with P. UF1, ΔdlaT P. UF1 (109 CFU/dam) or PBS. After birth, newborn mice of the respective dams continued to receive PBS, P. UF1, or ΔdlaT P. UF1 (107 CFU/mouse during breastfeeding, 108 CFU/mouse after weaning, once a week). Mice (n = 10) were then infected with ΔactA L. m (109 CFU/mouse) at 8 weeks of age. Seven days after infection, fecal samples were collected and microbiota composition was analyzed by 16S rDNA sequencing. a Histogram of linear discriminant analysis (LDA) scores reveals the most differentially abundant taxa among the indicated treatment groups. Only taxa meeting an LDA significant threshold > 2 are shown. b Taxonomic cladogram created using LDA effect size (LEfSe) method showing the phylogenetic distribution of differentially enriched taxa was compared among the indicated treatment groups. P. UF1-enriched taxa are indicated in green, ΔdlaT P. UF1-enriched taxa are indicated in blue and taxa enriched in PBS control are shown in red
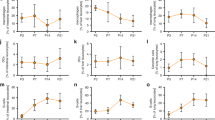
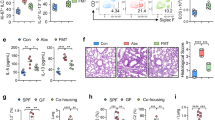
A myriad of microbiota-associated metabolites may affect the host-specific signaling that initiates functional gut homeostasis.22 Notably, analyzing the induced metabolites in fecal samples of newborn mice gavaged with P. UF1 for 8 weeks and then infected with ΔactA L. m when compared with other groups revealed unique features of induced metabolites. Here, the principal component analysis (PCA) showed a distinct clustering of fecal metabolomes from P. UF1, ΔdlaT P. UF1, and PBS-gavaged mice (Fig. 4a). Vitamin B2 metabolism, CoA catabolism/biosynthesis, prostaglandin formation, arachidonic acid metabolism and leukotriene metabolism were exclusively altered in P. UF1-gavaged mice in comparison to other groups (Fig. 4b). Significantly increased intensities of metabolite features, particularly riboflavin (vitamin B2), pantetheine (vitamin B5), and 5-methyltetrahydrofolate (vitamin B9), were observed in fecal samples of P. UF1-gavaged mice potentially regulating T cells, including Th17 cells,23,24,25,26 against pathogen infection (Fig. 4c). By contrast, proinflammatory metabolites, including prostaglandin E1, and 20-Hydroxyleukotriene E4, exhibited lower intensities in P. UF1-gavaged mice compared to other groups (Fig. 4c). These data may highlight the role of metabolite-associated signals that may regulate induced protective host immunity against pathogen invasion and the induced inflammation.6
P. UF1 induces metabolite changes during ΔactA L. m infection. Pregnant C57BL/6 dams were gavaged weekly with P. UF1, ΔdlaT P. UF1 (109 CFU/dam), or PBS. After birth, newborn mice of the respective dams were gavaged with P. UF1, ΔdlaT P. UF1 (107 CFU/mouse during breastfeeding, 108 CFU/mouse after weaning, once a week), or PBS. Mice (n = 10) were then orally infected with ΔactA L. m (109 CFU/mouse) at 8 weeks of age. Seven days after infection, fecal samples were collected and metabolome was analyzed by mass spectrometry. a PCA plot showing distinct clustering of fecal metabolomes from P. UF1-, ΔdlaT P. UF1- and PBS-gavaged mice. b Metabolic pathway analysis of metabolites with intensities significantly altered both in comparisons between P. UF1- versus PBS-gavaged group and P. UF1- versus ΔdlaT P. UF1-gavaged group. Red dashed line shows the permutation p value of 0.05 using Mummichog. c Scatter plots for selected metabolite features, with putative annotation, in the significant pathways identified by Mummichog. The m/z, retention time (in seconds) and adduct ion were labeled for each metabolite. Riboflavin, pantetheine, 5-methyltetrahydrofolate 2-arachidonoyl-glycerophosphocholine were identified in AE column; 20-hdroxyleukotriene E4 and prostaglandin E1 were identified in C18 column. Level of significance is shown as * p < 0.05, ** p < 0.01, *** p < 0.001
Transcriptomic regulation of Th17 cells by P. UF1 during intestinal infection
To investigate the transcriptomic program(s) potentially involved in the regulation of P. UF1-dependent Th17 cells, IL-17AeGFP mice receiving P. UF1 or ΔdlaT P. UF1 for 8 weeks were infected with ΔactA L. m, and CD4+ EGFP+ Th17 cells were sorted for transcriptomic analysis. Here, the PcoA demonstrated the differential transcriptomes of P. UF1- and ΔdlaT P. UF1-induced Th17 cells (Fig. 5a). Furthermore, among the 248 identified differentially expressed transcripts (FDR p < 0.05), a signature of key Th17 cell-related transcripts with significantly higher expression was observed in the P. UF1 group (e.g., lkzf3, Nr1d1, Cxcr6 and ll22) compared to the other group (Fig. 5b, c, and Supplementary Fig. 4a). To illuminate potential pathways enriched in either Th17 cells induced by P. UF1, or ΔdlaT P. UF1, Gene Set Enrichment Analysis (GSEA) was performed. Data demonstrated pathways related to extracellular matrix involved in the regulation of tissue homeostasis and healing were significantly enriched in P. UF1-induced Th17 cells (Fig. 5d). Additionally, genes implicated in TCR signaling and T cell activation (e.g., Lck, Cd69, Prkcq, and Zap70) were also upregulated in P. UF1-induced Th17 cells and supported by GSEA analysis demonstrating a gene set for a TCR signaling pathway (p < 0.005) (Supplementary Fig. 4b). In contrast, gene sets involved in inflammation and chemokine activity (e.g., Alox5, Cybb, Ccl3 and Ccl4) were highly enriched in ΔdlaT P. UF1-induced Th17 cells, suggesting the proinflammatory property of these cells during ΔactA L. m infection (Fig. 5c–e). Finally, we computed overlaps of the significantly upregulated genes in either P. UF1-, or ΔdlaT P. UF1-induced Th17 cells with the gene ontology (GO) term in the molecular signature database (MSigDB). Here, gene sets involved in immune regulation were observed in P. UF1-induced Th17 cells (Supplementary Fig. 4c, d). Furthermore, genes (e.g., Pias2 and Nr4a1) involved in the modulation of P. UF1-induced Th17 cell signaling were significantly increased in these cells compared to their counterparts (Fig. 5b). Together, these data may further demonstrate the critical role of P. UF1 in regulating DlaT-specific Th17 cells during L. m infection to clear this pathogen.
P. UF1 induces differentially expressed transcripts in Th17 cells during ΔactA L. m infection. IL-17AeGFP dams were gavaged with P. UF1 or ΔdlaT P. UF1. After birth, the respective pups were continuously gavaged with P. UF1 or ΔdlaT P. UF1 once a week for 8 weeks. Mice were then infected with ΔactA L. m and colonic Th17 cells were sorted for RNA-seq. a PCA plot of all genes identified. b Volcano plot of gene expression profile of Th17 cells derived from P. UF1- or ΔdlaT P. UF1-gavaged mice. c qRT-PCR analysis of expression of selected differentially expressed genes. d Bar graphs of the significantly enriched (FDR p < 0.05) gene sets among GO term, KEGG, Hallmark curated database. e Representative GSEA plots of “Chemokine Activity” from GO term database and “Inflammatory Response” from Hallmark database. * p < 0.05, ** p < 0.01, *** p < 0.01, two-tailed unpaired t-test
Discussion
The developmental process and the regulation of the fetal and neonatal immune system critically require the maternal resources, including metabolites.27 These factors are pivotal to induce the regulation in the developing fetus,27 initiating a homeostatic equilibrium that controls the developing neonatal adaptive immunity, including T cells.11 Thus, regulated neonatal T cell immunity should evolve rapidly to override the acquired immature immunity, which may not be sufficient to protect against infectious diseases.28,29 The protective immune regulation is even more relevant for newborns fed formula instead of HBM.30 The developmental process of regulated immune protection is highly intertwined with exposure to the mother’s microbiota, bacterial-associated metabolites, and nutritional diets. Errors in this complex may disrupt the gut homeostasis, potentially manifesting in microbial dysbiosis31 that further skews immature immunity toward inflammation that can potentially induce tissue damage.32 Although the multifactorial mechanisms implicated in intestinal inflammatory conditions33 are currently elusive, infiltrating uncontrolled intestinal T cells, particularly proinflammatory Th17 cells, may contribute to devastating intestinal disorders.12 Thus, the scientific effort is currently centered on the novel regulatory mechanisms that can accelerate the neonatal immune regulation to swiftly resolve potential intestinal infections to prevent tissue damage. Available data thus far indicate that these regulatory mechanisms are highly dictated by critical nutrients in HBM, along with the mother’s commensals synthesizing critical metabolites34 that positively affect the gut commensals and the neonatal immune homeostasis.35 However, further mechanistic insights are urgently required to better understand the molecular mechanisms involved in the regulation of neonatal T cells,36 particularly Th17 cells involved in mucosal protection and the pathogenesis of intestinal diseases.14
We previously demonstrated P. UF1 induced DlaT-dependent Th17 cell regulation in adult mice in steady state and during intestinal infection.9 To further elucidate the impact of this beneficial bacterium and its gene products (e.g., DlaT) on the development of intestinal T cell immunity in newborn mice, we have established a neonatal mouse model and feeding regimen to elaborate on the role of P. UF1 in regulating and accelerating the evolution of intestinal T cell immunity, particularly protective Th17 cells, which are critically involved in the clearance of intestinal infection.37 Our work highlighted the ability of a beneficial bacterium, P. UF1, with bifidogenic properties to significantly direct resistance against pathogen-induced inflammation in the developing newborn mice through highly regulated CD4+ T cell subsets, including Th17 cells, over time. This is critically relevant because pathogen exposure in the early postnatal phase may impact lifelong immune homeostasis and disease susceptibility.29 Gavaging newborn mice with P. UF1 for 8 weeks also resulted in a better outcome of gut microbiota and the induced metabolites, including vitamin B group, particularly VB9 that is highly involved in the maintenance of Tregs.24 Thus, prime bacterial candidate(s), such as P. UF1, may possess the ability to accelerate the frequency and the regulation of intestinal CD4+ T cells, and sustain functional gut microbiota and their critical metabolites in evolving newborn mouse immunity, all of which potentially promote postnatal immune protection against pathogen infection such as L. m. Furthermore, as demonstrated in this report, P. UF1-induced Th17 cells differed in nature from segmented filamentous bacterium (SFB)-promoted Th17 cells.38 Mainly, P. UF1-dependent Th17 cells functionally maintain their committed property while SFB-induced Th17 cells acquire functional characteristics of Th1 cells adapting a proinflammatory role during Listeria infection.38 Such a Th17 cell plasticity39 may be dictated by the nature of SFB creating an environment that skews Th17 cells toward Th1 polarization.38 Yet, this aspect of P. UF1-induced Th17 cells may be explained by the transcriptional regulation of these cells by P. UF1, as demonstrated in this report. Furthermore, the nature of this beneficial bacterium synthesizing critical metabolites, including propionate, may direct the regulation of T cell molecular machinery. Nonetheless, this notion is currently under intensive scrutiny in our laboratory to clearly demonstrate the role of P. UF1-induced metabolites on immune cells, particularly Th17 cells, during the intestinal infection. Furthermore, our studies are also designed to elucidate the impact of P. UF1-induced metabolites, including propionate, that seem to be implicated in acidifying the bacterial milieu resulting in a significant reduction of pathogen growth.21 Thus, elucidating the role of beneficial bacteria-induced metabolites will further illuminate the therapeutic aspects of these critical compounds in health and disease treatments in the near future.
As discussed before,9 classically protective immunity against intracellular pathogens such as L. m was considered to require IFNγ+ Th1 cell responses.40 However, IFNγ- CD4+ T cell response demonstrated protection against this pathogen41 indicating that further critical factors may also be involved in immune protection against pathogen challenges. Accordingly, data demonstrated that patients lacking functional Th17 cells due to errors in IL-17 and IL-6 signaling exhibited weaker immune protection against pathogen infections.42,43 Available data thus far also highlight the fact that primed Th1 cells could then be effective when these cells synergistically cooperate with Th17 cells to resolve the intracellular pathogen infection.44,45 However, such synergistically induced Th17 and Th1 cells must be robustly regulated, as demonstrated here, to potentially limit tissue damage.46 Critically, the frequency of IL-10+ Tregs that reduced during ΔactA L. m infection47 was also maintained, which potentially regulated the induced protective Th17 and IFNγ+ Th1 cells.9,10 Notably, a similar trend was also observed in newborn mice that were continuously gavaged with P. UF1, which clearly enhanced the frequency of protective CD4+ T cells, including Th17 cells, against ΔactA L. m infection. Additionally, the relevance of the protective property of T cells, particularly Th17, against pathogen infection was also shown by neutralizing IL-17A in mice in the present report. Our obtained data from Rag1-/- mice with no T cell transfer supported this notion, once again highlighting the protective role of T cells against pathogen infection. Yet, would other critical factors, including functional microbiota and their induced metabolites, also contribute to the rigid protection of mice against pathogen infection? To elucidate these significant points of interest, further experiments are warranted in the near future using relevant mouse models, including Rag1-/- germfree mice. Nonetheless, the concurrent data clearly demonstrate not only the important contribution of gut microbiota19 and their metabolites21 against pathogen infection (e.g., Listeria), but also that these two factors may be critically involved in the functional improvement of protective T cells in a pathogen-induced condition, as demonstrated in this report.
Collectively, our data demonstrate that P. UF1 significantly increases the frequencies of T cells (e.g., Th17 cells) and influences the commensal phylum and the induced metabolic profile. This beneficial bacterium also fortifies the transcripts in Th17 cell regulation during intestinal pathogen infection potentially contributing to immune regulation that may prevent tissue damage. Further mechanistic analyses are warranted to understand the molecular cues exerted by P. UF1, including its metabolic machinery and gene products, which includes surface layer proteins (S-layers) involved in immune regulation resisting intestinal infection inducing-pathogenic-inflammation that may result in intestinal tissue damage, and to implement novel bacteria with bifidogenic properties to possibly prevent intestinal inflammatory diseases and neonatal pathogenic infections.
Methods
Animals
C57BL/6 and C57BL/6-Il17atm1Bcgen/J (IL-17AeGFP) and Rag1-/- mice were obtained from Jackson Laboratory and maintained under specific pathogen-free, Helicobacter-free conditions, in accordance with the Animal Welfare Act and the Public Health Policy on Humane Care. Procedures were approved by UF’s Institutional Animal Care and Use Committee (IACUC), protocol numbers: 201708484 and 201609388.
Bacterial cultures
Propionibacterium University of Florida 1 (P. UF1) and its derivative strains, were grown anaerobically at 30 °C in MRS medium (Difco Laboratories, Detroit, MI) supplemented with 1% (w/v) sodium lactate (Thermo Fisher Scientific, Rockford, IL), as previously described.9 ΔactA Listeria monocytogenes (L. m) was grown in Brain Heart Infusion (BHI) medium (Difco Laboratories, Detroit, MI) at 37 °C. Antibiotics were added at the following final concentrations: 5 μg/ml chloramphenicol for isogenic P. UF1 strains; 200 μg/ml streptomycin and 50 μg/ml kanamycin for ΔactA L. m.
Bacterial administration
Female mice (dam) were gavaged with PBS, P. UF1, ΔdlaT P. UF1 (109 CFU/dam, once a week) during pregnancy. After birth, newborn mice of the respective dams were gavaged with PBS, P. UF1 or ΔdlaT P. UF1 (107 CFU/dam, once a week). Groups of mice were sacrificed to analyze immunity at the age of 1 and 2 weeks. After weaning, mice were continuously gavaged with PBS, P. UF1 or ΔdlaT P. UF1 (108 CFU/dam, once a week). Mice were sacrificed at the age of 1–8 weeks to analyze CD4+ T cell immune response.
ΔactA L. m infection
C57BL/6 dams were gavaged with PBS, P. UF1, ΔdlaT P. UF1 (109 CFU/dam, once a week) during pregnancy. After birth, newborn mice of corresponding dams were gavaged with PBS, P. UF1 or ΔdlaT P. UF1 (107 CFU/mouse during breastfeeding and 108 CFU/mouse after weaning, once a week) until the age of 3, 4, 6, and 8 weeks. For infection, food was withdrawn for 4 h before mice were orally infected with 109 CFU of ΔactA L. m in 100 μl PBS containing 50 mg/ml CaCO3. To minimize the impact of ΔactA L. m on 3-week-old mice, mice were infected with 108 CFU in 100μl PBS containing 50 mg/ml CaCO3. The ΔactA L. m burdens in fecal samples were then monitored from days 1–4 post infection by plating the fecal samples on streptomycin-resistant BHI agar plates. Additionally, Rag1-/- mice (8 weeks old) were gavaged with PBS, or P. UF1 (4 gavages, every other day, 109 CFU/mouse) and then orally infected with ΔactA L. m (109 CFU/mouse). Fecal samples were collected from days 1 to 5 post infection and the CFU burdens were determined by plating serial dilutions of fecal samples on streptomycin-resistant plates.
Maternal effects of P. UF1
To investigate P. UF1 colonization in the mouse intestine, an antibiotic-resistant P. UF1 strain was constructed by integration of chloramphenicol resistant gene (cmR) into the P. UF1 chromosome at the alpha-amylase (amyE) locus. Briefly, a 689-bp downstream fragment of amyE gene was amplified from P. UF1 genome using primers amyE-F (5′-CCCAAGCTTGTATGGCTCATGCTGCACTCG-3′) and amyE-R (5′-CCCGAATTCCGTGACCGGCCTCAGTCCTT-3′), and the cmR gene amplified from pUCC-dlaT9 using cmR-F (5′-CCCGGATCCCGTCACACCCGAACATGTCG-3′) and cmR-R (5′-CCCGGTACCGATGGGTCATCAATTGGCCTC-3′). The purified fragments amyE and CmR were cloned into pUC19 plasmid to construct suicide plasmid pUCCmR. Following electroporation into P. UF1, chloramphenicol-resistant colonies were selected and the CmR P. UF1 strain was identified by PCR using primers P1 (5′-CGGTGGGATGGAAGAGCC-3′) and P2 (5′-GCGGTGGACCCGGAATT-3′). For demonstrating the bacterial transfer from dams to their pups, C57BL/6 dams were gavaged with CmR P. UF1 (109 CFU/dam, twice a week) during pregnancy and breastfeeding. After birth, newborn mice were sacrificed at 5–14 days of age, and colonic luminal contents were collected, weighted, and resuspended in PBS. The presence of CmR P. UF1 was determined by plating the fecal samples on chloramphenicol-resistant MRS-lactate agar plates. For investigating the maternal effect, C57BL/6 dams were gavaged with P. UF1 (109 CFU/dam, twice a week) or PBS during pregnancy and breastfeeding. After birth, newborn mice did not receive any gavages with P. UF1 or PBS. On day 21, the newborn mice were infected with ΔactA L. m (108 CFU/mouse), and fecal samples were collected on days 1–4 post infection for monitoring pathogen burdens. Seven days post infection, mice were sacrificed to analyze colonic CD4+ T cell response.
IL-17A neutralization
C57BL/6 mice were continuously gavaged with PBS, P. UF1, or ΔdlaT P. UF1 as described above until 8 weeks of age, and then intraperitoneally injected with purified anti-mouse IL-17A monoclonal antibody (Biolegend, San Diego, CA) on day −6 and −2 relative to infection (100 μg/mouse/injection). On day 0, mice were orally infected with 109 CFU of ΔactA L.m and fecal Listeria burdens were determined from days 1 to 4 post infection.
Flow cytometry
Colonic cells were isolated as previously described.9 For Supplementary Figs. 1 and 2, an improved protocol for T cell isolation was used. Briefly, after shaking in PBS containing 20 mM hepes and 10 mM EDTA for 30 min at 37 °C, colonic tissues were cut into small pieces and incubated with digestion solution [RPMI 1640 containing 10% FBS (Thermo Fisher Scientific, Waltham, MA), 0.4% β-mercaptoethanol, 400 U/ml collagenase VIII (Sigma Aldrich, St. Louis, MO) and 100 μg/ml DNase I (Sigma Aldrich, St. Louis, MO)] for 1.5 h at 37 °C. Digested tissues were filtered through 100 μm cell strainer (Genesee Scientific, San Diego, CA), and cells were resuspended in 5 ml of 40% Percoll (Sigma Aldrich, St. Louis, MO) and overlaid on 5 ml of 80% Percoll. Percoll gradient separation was performed by centrifugation at 700 × g for 25 min at 25 °C. Cells in the interphase were collected and used for flow cytometry analysis.9 Briefly, after stimulation with phorbol 12-myristate 13-acetate (PMA) (50 ng/ml) and ionomycin (2.5 μg/ml) in the presence of brefeldin A (Sigma-Aldrich, St. Louis, MO), colonic cells were stained with LIVE/DEAD® Fixable Blue Dead Cell Stain Kit (Thermo Fisher Scientific, Waltham, MA). Cells were then stained for cell-surface markers and resuspended in FOXP3 Fix/Perm buffer set (Biolegend, San Diego, CA) for intracellular staining. Antibodies and isotypes were purchased from Biolegend, or eBioscience (San Diego, CA). The following antibodies and their corresponding isotypes were used: CD45 (30-F11), CD3 (145–2C11), CD4 (RM4–5), CD8 (53–607), IFNγ (XMG1.2)/Rat IgG1, κ, IL-17A (TC11–18H10.1)/Rat IgG1, κ, IL-10 (JES5–16E3)/Rat IgG2b, κ, FoxP3 (FJK-16A)/Rat IgG2a, κ, TGFβ (TW7–16B4)/Rat IgG1. Flow cytometric data were collected and analyzed with FlowJo software (TreeStar, Ashland, OR). After dead and doublet cell exclusion and the subsequent CD45+ and FCS/SSC selection, T cells were defined as CD3+CD4+CD8-. IL-10+ IL-17A+ CD4+, IL-10+ IFNγ+ CD4+, IL-10+ FoxP3+ CD4+, and IL-10+ TGFβ+ FoxP3+ CD4+ cells were then plotted (Supplementary Fig. 5).
Sorting Th17 cells
IL-17AeGFP dams and their newborns were gavaged with P. UF1 or ΔdlaT P. UF1 for 8 weeks, as described above. Mice were then orally infected with ΔactA L. m, and colonic cells were isolated 7 days post infection. Th17 cells were sorted with a SONY SH800S Cell Sorter (Sony, Tokyo, Japan) after being stained with anti-CD45, anti-CD3, anti-CD4 antibodies following manufactures’ suggested protocol.
RNA-sequencing
Total RNA was isolated from sorted Th17 cells with an RNeasy Plus Micro kit (Qiagen, Germantown, MD), and cDNA was generated with a SMART-Seq v4 Ultra-Low Input RNA Kit (Clontech, Mountain View, CA). Following fragmentation, cDNA library was constructed using a Nextera XT DNA Library Preparation Kit (Illumina, Inc., San Diego, CA) and sequenced on an Illumina HiSeq instrument (Illumina, Inc., San Diego, CA) at the University of Florida ICBR NextGen DNA Sequencing Core Facility. FastQ files were aligned to the mouse genome (NCBI GRCm38/mm10) using STAR aligner (v2.6.0),48 and count table was generated using SubReads featureCounts (v1.6.0).49 Genes with lower expressed transcripts (mean expression value less than one RPKM throughout the dataset) were excluded. Subsequently, DESeq2 was used to determine differential expression.50 Stringent criteria were set to determine significantly dysregulated genes: FDR p-value less than 0.05 and fold-change (FC) greater than 1.5. Selected genes were mapped on the Volcano plot. Regularized-log-transformation of count data was performed before principal component analysis (PCA) and heatmap plotting. Gene set enrichment analysis (GSEA) was performed in the javaGSEA (v3.0) using the GseaPreranked method, with genes ranked according to their DESeq2 Wald test statistics. GSEA’s “weighted” scoring scheme was used to calculate enrichment scores for all gene sets in database, including GO, Hallmark, KEGG and REACTOME. Nominal enrichment p values were based on 1000 permutations. GSEA enrichment plots were made using code adapted from Rtoolbox. Over-representative test of genes significantly upregulated in either condition (FDR p < 0.05) was performed in GSEA website (http://software.broadinstitute.org/gsea/msigdb/annotate.jsp).
Quantitative reverse-transcription PCR (qRT-PCR)
Transcripts for genes of interest were quantified by qRT-PCR using cDNA generated for RNA-seq. PCR was performed on a CFX96 real-time PCR system (Bio-Rad, Hercules, CA) using SsoAdvanced Universal SYBR Green Supermix (Bio-Rad, Hercules, CA) with following primers: Gapdh, 5′-GGTGAAGGTCGGTGTGAACG-3′ and 5′-CTCGCTCCTGGAAGATGGTG-3′; Cxcr6, 5′-ACCAGCTTTAAGTATGCCATCGTA-3′ and 5′-TCCGGAACTTTAAGCCAACAA-3′; Prkcq, 5′-TGGGCGGACAGAAATATGGT-3′ and 5′-TCCTTCATTCTCAAACTCACTCATGT-3′; Neu3, 5′- CTCAGTCAGAGATGAGGATGCT-3′ and 5′-GTGAGACATAGTAGGCATAGGC-3′; Alox5, 5′-CTCTTCCAAGCTCGAAGTGC-3′ and 5′-TGATGCTACCGAGTGACGAG-3′; Cxcl1, 5′-TGGCTGGGATTCACCTCAAG-3′ and 5′-CAAGCCTCGCGACCATTCT-3′; Cxcl2, 5′-CGCTGTCAATGCCTGAAGAC-3′ and 5′-ACACTCAAGCTCTGGATGTTCTTG-3′; Ccl5, 5′-TCGTGCCCACGTCAAGGAGTATTT-3′ and 5′-TCTTCTCTGGGTTGGCACACACTT-3′; Cybb, 5′-CATCGGTGACAATGAGAACG-3′ and 5′-AAGGCCGATGAAGAAGATCA-3′. Measurements were made in duplicate wells, and results were normalized to those obtained with Gapdh and are presented as fold change over P. UF1.
Ultra-high-resolution metabolomics analysis
Mouse fecal samples were collected and analyzed on an LTQ-FTICR mass spectrometer coupled with anion exchange (AE) and reversed-phase (C18) liquid chromatography, as previously described.9 We used apLCMS to perform peak detection, noise filtering, mass to charge (m/z) ratio and retention time alignment and feature quantification.51 An in-house informatics pipeline was used to quality control and clean up the feature table, as previously described.9 PCA was performed on log2-transformed intensity values of top 500 metabolite features with highest CV across all samples. Pairwise student’s t-tests were performed among treatment groups. Subsequently, metabolic pathway analysis was performed by Mummichog (v2.0), a software specifically designed for untargeted metabolomics,52 using default parameters. 205 metabolite features (C18 run) and 203 metabolite features (AE run), which are either significantly higher or lower in P. UF1 group compared to PBS or ΔdlaT P. UF1 groups of mice (p < 0.05, Student’s t-test) were used as input to Mummichog. The total list of features was used as a reference. Pathway enrichment p-values were calculated in Mummichog based on permutation. The pathways were represented by at least two significant metabolites and enriched at p < 0.05 in either chromatographic column.
Microbiota analysis
Fecal DNA was isolated and DNA library constructed.9 The 16S rDNA library was sequenced on an Illumina Miseq (Illumina, Inc., San Diego, CA), and sequence analysis was performed as previously described.9 Briefly, FastQ files were jointed, de-multiplexed and analyzed by QIIME (v1.9.1). PcoA and the Permanova analyses were also performed using QIIME (v1.9.1) and replotted using Python. Significant taxons were filtered by LDA Effect Size (LEfSe) analyses with default criteria (p < 0.05 by Kruskal–Wallis test; LDA score > 2).
Statistical analysis
Statistical analyses were performed using GraphPad Prism v7.0. Data are expressed as mean ± SEM of number (n) of mice or samples per group. Prior to statistical analysis, normality was tested using the Shapiro–Wilk normality test. Where the groups follow a Gaussian distribution, two-tailed unpaired t-test (for two variables) or one-way ANOVA followed by Tukey’s post-test (for three variables) was performed. Where the groups did not follow a Gaussian distribution, Mann–Whitney U test (for two variables) or Kruskal–Wallis test followed by Dunn’s post-test (for three variables) was performed. p values lower than 0.05 were considered as significant: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
References
PrabhuDas, M. et al. Challenges in infant immunity: implications for responses to infection and vaccines. Nat. Immunol. 12, 189–194 (2011).
Pollard, A. J., Perrett, K. P. & Beverley, P. C. Maintaining protection against invasive bacteria with protein-polysaccharide conjugate vaccines. Nat. Rev. Immunol. 9, 213–220 (2009).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Richards, J. L., Yap, Y. A., McLeod, K. H., Mackay, C. R. & Marino, E. Dietary metabolites and the gut microbiota: an alternative approach to control inflammatory and autoimmune diseases. Clin. Transl. Immunol. 5, e82 (2016).
Levy, M., Blacher, E. & Elinav, E. Microbiome, metabolites and host immunity. Curr. Opin. Microbiol. 35, 8–15 (2017).
Veldhoen, M. & Brucklacher-Waldert, V. Dietary influences on intestinal immunity. Nat. Rev. Immunol. 12, 696–708 (2012).
Zhang, K. et al. Age-dependent enterocyte invasion and microcolony formation by Salmonella. PLoS Pathog. 10, e1004385 (2014).
Dupont, A. et al. Age-dependent susceptibility to enteropathogenic Escherichia coli (EPEC) infection in mice. PLoS Pathog. 12, e1005616 (2016).
Colliou, N. et al. Commensal Propionibacterium strain UF1 mitigates intestinal inflammation via Th17 cell regulation. J. Clin. Invest. 127, 3970–3986 (2017).
Colliou, N. et al. Regulation of Th17 cells by P. UF1 against systemic Listeria monocytogenes infection. Gut Microbes 9, 279–287 (2018).
Torow, N. & Hornef, M. W. The neonatal window of opportunity: setting the stage for life-long host-microbial interaction and immune homeostasis. J. Immunol. 198, 557–563 (2017).
Neu, J. & Walker, W. A. Necrotizing enterocolitis. N. Engl. J. Med. 364, 255–264 (2011).
Zhou, Z. H., McCarthy, D. B., O’Connor, C. M., Reed, L. J. & Stoops, J. K. The remarkable structural and functional organization of the eukaryotic pyruvate dehydrogenase complexes. Proc. Natl Acad. Sci. USA 98, 14802–14807 (2001).
Littman, D. R. & Rudensky, A. Y. Th17 and regulatory T cells in mediating and restraining inflammation. Cell 140, 845–858 (2010).
Zhang, Z., Clarke, T. B. & Weiser, J. N. Cellular effectors mediating Th17-dependent clearance of pneumococcal colonization in mice. J. Clin. Invest. 119, 1899–1909 (2009).
Lin, Y. et al. Interleukin-17 is required for T helper 1 cell immunity and host resistance to the intracellular pathogen Francisella tularensis. Immunity 31, 799–810 (2009).
Pham, T. A. & Lawley, T. D. Emerging insights on intestinal dysbiosis during bacterial infections. Curr. Opin. Microbiol. 17, 67–74 (2014).
Spencer, S. P. & Belkaid, Y. Dietary and commensal derived nutrients: shaping mucosal and systemic immunity. Curr. Opin. Immunol. 24, 379–384 (2012).
Becattini, S. et al. Commensal microbes provide first line defense against Listeria monocytogenes infection. J. Exp. Med. 214, 1973–1989 (2017).
Kamada, N., Chen, G. Y., Inohara, N. & Nunez, G. Control of pathogens and pathobionts by the gut microbiota. Nat. Immunol. 14, 685–690 (2013).
Jacobson, A. et al. A Gut commensal-produced metabolite mediates colonization resistance to salmonella infection. Cell Host Microbe 24, 296–307 e297 (2018).
Lee, W. J. & Hase, K. Gut microbiota-generated metabolites in animal health and disease. Nat. Chem. Biol. 10, 416–424 (2014).
Kinoshita, M. et al. Dietary folic acid promotes survival of Foxp3+regulatory T cells in the colon. J. Immunol. 189, 2869–2878 (2012).
Kunisawa, J., Hashimoto, E., Ishikawa, I. & Kiyono, H. A pivotal role of vitamin B9 in the maintenance of regulatory T cells in vitro and in vivo. PLoS One 7, e32094 (2012).
Kjer-Nielsen, L. et al. MR1 presents microbial vitamin B metabolites to MAIT cells. Nature 491, 717–723 (2012).
Brestoff, J. R. & Artis, D. Commensal bacteria at the interface of host metabolism and the immune system. Nat. Immunol. 14, 676–684 (2013).
Macpherson, A. J., de Aguero, M. G. & Ganal-Vonarburg, S. C. How nutrition and the maternal microbiota shape the neonatal immune system. Nat. Rev. Immunol. 17, 508–517 (2017).
Goenka, A. & Kollmann, T. R. Development of immunity in early life. J. Infect. 71(Suppl 1), S112–S120 (2015).
Shane, A. L., Sanchez, P. J. & Stoll, B. J. Neonatal sepsis. Lancet, https://doi.org/10.1016/S0140-6736(17)31002-4 (2017).
Schwartz, S. et al. A metagenomic study of diet-dependent interaction between gut microbiota and host in infants reveals differences in immune response. Genome Biol. 13, r32 (2012).
Groer, M. W. et al. Development of the preterm infant gut microbiome: a research priority. Microbiome 2, 38 (2014).
Lim, J. C., Golden, J. M. & Ford, H. R. Pathogenesis of neonatal necrotizing enterocolitis. Pediatr. Surg. Int. 31, 509–518 (2015).
Besner, G. E. A pain in the NEC: research challenges and opportunities. J. Pediatr. Surg. 50, 23–29 (2015).
Degnan, P. H., Taga, M. E. & Goodman, A. L. Vitamin B12 as a modulator of gut microbial ecology. Cell. Metab. 20, 769–778 (2014).
Urbaniak, C. et al. Microbiota of human breast tissue. Appl. Environ. Microbiol. 80, 3007–3014 (2014).
Cervantes-Barragan, L. et al. Lactobacillus reuteri induces gut intraepithelial CD4+CD8alphaalpha+T cells. Science 357, 806–810 (2017).
Peck, A. & Mellins, E. D. Precarious balance: Th17 cells in host defense. Infect. Immun. 78, 32–38 (2010).
Yang, Y. et al. Focused specificity of intestinal TH17 cells towards commensal bacterial antigens. Nature 510, 152–156 (2014).
Muranski, P. & Restifo, N. P. Essentials of Th17 cell commitment and plasticity. Blood 121, 2402–2414 (2013).
Hsieh, C. S. et al. Development of TH1 CD4+T cells through IL-12 produced by Listeria-induced macrophages. Science 260, 547–549 (1993).
Harty, J. T. & Bevan, M. J. Specific immunity to Listeria monocytogenes in the absence of IFN gamma. Immunity 3, 109–117 (1995).
Puel, A. et al. Chronic mucocutaneous candidiasis in humans with inborn errors of interleukin-17 immunity. Science 332, 65–68 (2011).
Milner, J. D. et al. Impaired T(H)17 cell differentiation in subjects with autosomal dominant hyper-IgE syndrome. Nature 452, 773–776 (2008).
Thiemann, S. et al. Enhancement of IFNgamma production by distinct commensals ameliorates salmonella-induced disease. Cell Host Microbe 21, 682–694 e685 (2017).
Ost, K. S. & Round, J. L. A few good commensals: gut microbes use IFN-gamma to fight Salmonella. Immunity 46, 977–979 (2017).
Veldhoen, M. Interleukin 17 is a chief orchestrator of immunity. Nat. Immunol. 18, 612–621 (2017).
Ertelt, J. M. et al. Selective culling of high avidity antigen-specific CD4+T cells after virulent Salmonella infection. Immunology 134, 487–497 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, T., Park, Y., Johnson, J. M. & Jones, D. P. apLCMS–adaptive processing of high-resolution LC/MS data. Bioinformatics 25, 1930–1936 (2009).
Li, S. et al. Predicting network activity from high throughput metabolomics. PLoS Comput. Biol. 9, e1003123 (2013).
Acknowledgements
This work was supported by NIH R01 DK109560 (to M.M.).
Author information
Authors and Affiliations
Contributions
M.M. directed and designed the cellular and molecular experiments, which were executed by N.C., Y.G., M.G., J.L., and M.Z. N.C., Y.G. and M.Z. performed the mouse work and flow cytometry analysis. Y.G. and J.L. performed the molecular studies, including transcriptomic studies. M.G., S.L., and D.J. performed, analyzed, and directed metabolomic studies. M.G. conducted all bioinformatic work for microbiota, metabolic, and transcriptomic data analyses. Y.G., N.C., M.G., J.L., S.L., D.P.J., and M.M. analyzed and interpreted the data. M.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ge, Y., Gong, M., Colliou, N. et al. Neonatal intestinal immune regulation by the commensal bacterium, P. UF1. Mucosal Immunol 12, 434–444 (2019). https://doi.org/10.1038/s41385-018-0125-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41385-018-0125-1
This article is cited by
-
Murine in utero exposure to simulated complex urban air pollution disturbs offspring gut maturation and microbiota during intestinal suckling-to-weaning transition in a sex-dependent manner
Particle and Fibre Toxicology (2022)
-
Improving the drying of Propionibacterium freudenreichii starter cultures
Applied Microbiology and Biotechnology (2021)
-
Regulating colonic dendritic cells by commensal glycosylated large surface layer protein A to sustain gut homeostasis against pathogenic inflammation
Mucosal Immunology (2020)