Abstract
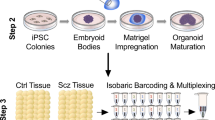
The way in which brain morphology and proteome are remodeled during embryonal development, and how they are linked to the cellular metabolism, could be a key for elucidating the pathological mechanisms of certain neurodevelopmental disorders. Cerebral organoids derived from autism spectrum disorder (ASD) patients were generated to capture critical time-points in the neuronal development, and metabolism and protein expression were investigated. The early stages of development, when neurogenesis commences (day in vitro 39), appeared to be a critical timepoint in pathogenesis. In the first month of development, increased size in ASD-derived organoids were detected in comparison to the controls. The size of the organoids correlates with the number of proliferating cells (Ki-67 positive cells). A significant difference in energy metabolism and proteome phenotype was also observed in ASD organoids at this time point, specifically, prevalence of glycolysis over oxidative phosphorylation, decreased ATP production and mitochondrial respiratory chain activity, differently expressed cell adhesion proteins, cell cycle (spindle formation), cytoskeleton, and several transcription factors. Finally, ASD patients and controls derived organoids were clustered based on a differential expression of ten proteins—heat shock protein 27 (hsp27) phospho Ser 15, Pyk (FAK2), Elk-1, Rac1/cdc42, S6 ribosomal protein phospho Ser 240/Ser 244, Ha-ras, mTOR (FRAP) phospho Ser 2448, PKCα, FoxO3a, Src family phospho Tyr 416—at day 39 which could be defined as potential biomarkers and further investigated for potential drug development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
World Health Organization. The ICD-10 classification of mental and behavioral disorders: clinical descriptions and diagnostic guidelines. World Health Organization. 1992. https://apps.who.int/iris/handle/10665/37958
Forsberg SL, Ilieva M, Maria Michel T. Epigenetics and cerebral organoids: promising directions in autism spectrum disorders. Transl Psychiatry. 2018;8:14 https://doi.org/10.1038/s41398-017-0062-x
Ilieva M, Fex Svenningsen Å, Thorsen M, Michel TM. Psychiatry in a dish: stem cells and brain organoids modeling autism spectrum disorders. Biol Psychiatry. 2018;83:558–68. https://doi.org/10.1016/j.biopsych.2017.11.011.
Courchesne E, Pramparo T, Gazestani VH, Lombardo MV, Pierce K, Lewis NE. The ASD living biology: from cell proliferation to clinical phenotype. Mol Psychiatry. 2019;24:88–107. https://doi.org/10.1038/s41380-018-0056-y.
Mariani J, Coppola G, Zhang P, Abyzov A, Provini L, Tomasini L, et al. FOXG1-dependent dysregulation of GABA/glutamate neuron differentiation in autism spectrum disorders. Cell 2015;162:375–90. https://doi.org/10.1016/j.cell.2015.06.034
Zafeiriou DI, Ververi A, Dafoulis V, Kalyva E, Vargiami E. Autism spectrum disorders: the quest for genetic syndromes. Am J Med Genet B Neuropsychiatr Genet. 2013;162B:327–66. https://doi.org/10.1002/ajmg.b.32152.
Bai D, Yip BHK, Windham GC, Sourander A, Francis R, Yoffe R, et al. Association of genetic and environmental factors with autism in a 5-country cohort. JAMA Psychiatry. 2019;76:1035–43. https://doi.org/10.1001/jamapsychiatry.2019.1411
Satterstrom FK, Kosmicki JA, Wang J, Breen MS, De Rubeis S, An JY, et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell. 2020;180:568–.e23. https://doi.org/10.1016/j.cell.2019.12.036
Lombardo MV, Lai MC, Baron-Cohen S. Big data approaches to decomposing heterogeneity across the autism spectrum. Mol Psychiatry. 2019;24:1435–50. https://doi.org/10.1038/s41380-018-0321-0
Higdon R, Earl RK, Stanberry L, Hudac CM, Montague E, Stewart E, et al. The promise of multi-omics and clinical data integration to identify and target personalized healthcare approaches in autism spectrum disorders. Omics: J Integr Biol. 2015;19:197–208. https://doi.org/10.1089/omi.2015.0020
Gould, Stephen Jay. Ontogeny and phylogeny. Harvard University Press, 1985.
Somel M, Franz H, Yan Z, Lorenc A, Guo S, Giger T, et al. Transcriptional neoteny in the human brain. Proc Natl Acad Sci USA. 2009;106:5743–8. https://doi.org/10.1073/pnas.0900544106.
Bufill E, Agustí J, Blesa R. Human neoteny revisited: the case of synaptic plasticity. Am J Hum Biol. 2011;23:729–39. https://doi.org/10.1002/ajhb.21225.
Crespi B. Developmental heterochrony and the evolution of autistic perception, cognition and behavior. BMC Med. 2013;11:119 https://doi.org/10.1186/1741-7015-11-119
Thorsen M, Bilenberg N, Thorsen L, Michel TM. Oxidative stress in adults with autism spectrum disorder: a case control study. J Autism Dev Disord. 2022;52:275–82. https://doi.org/10.1007/s10803-021-04897-x.
American Psychiatric Association, DSM-5 Task Force. Diagnostic and statistical manual of mental disorders: DSM-5™ (5th ed.). American Psychiatric Publishing, Inc. 2013 https://doi.org/10.1176/appi.books.9780890425596
Okita K, Matsumura Y, Sato Y, Okada A, Morizane A, Okamoto S, et al. A more efficient method to generate integration-free human iPS cells. Nat Methods. 2011;8:409–12. https://doi.org/10.1038/nmeth.1591.
Kamand M, Ilieva M, Forsberg SL, Thomassen M, Svenningsen ÅF, Meyer M, et al. Generation of autism spectrum disorder patient-derived iPSC line SDUKIi004-A. Stem Cell Res. 2020;49:102038 https://doi.org/10.1016/j.scr.2020.102038.
Kamand M, Forsberg SL, Thomassen M, Ilieva M, Meyer M, Svenningsen ÅF, et al. Establishment of an induced pluripotent stem (iPS) cell line (SDUKIi006-A) from a 21-year-old male patient diagnosed with atypical autism disorder. Stem Cell Res. 2021;51:102185 https://doi.org/10.1016/j.scr.2021.102185.
Kamand M, Ilieva M, Louise Forsberg S, Thomassen M, Meyer M, Fex Svenningsen Å, et al. Derivation of induced pluripotent stem cells (SDUKIi003-A) from a 20-year-old male patient diagnosed with Asperger syndrome. Stem Cell Res. 2020;48:101974 https://doi.org/10.1016/j.scr.2020.101974.
Kamand M, Ilieva M, Forsberg SL, Thomassen M, Fex Svenningsen Å, Holst B, et al. Generation of human induced pluripotent stem cells (SDUKIi002-A) from a 22-year-old male diagnosed with autism spectrum disorder. Stem Cell Res. 2020;46:101834 https://doi.org/10.1016/j.scr.2020.101834.
Lancaster MA, Knoblich JA. Generation of cerebral organoids from human pluripotent stem cells. Nat Protoc. 2014;9:2329–40. https://doi.org/10.1038/nprot.2014.158.
Biemann K. Mass spectrometry in organic chemistry applications. New York: MrGraw. 1962:223-7.
Bak LK, Schousboe A, Sonnewald U, Waagepetersen HS. Glucose is necessary to maintain neurotransmitter homeostasis during synaptic activity in cultured glutamatergic neurons. J Cereb Blood Flow Metab. 2006;26:1285–97.
Treindl F, Ruprecht B, Beiter Y, Schultz S, Döttinger A, Staebler A, et al. A bead-based western for high-throughput cellular signal transduction analyses. Nat Commun. 2016;7:12852 https://doi.org/10.1038/ncomms12852
Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques. 2003;34:374–8. https://doi.org/10.2144/03342mt01
Berndt N, Kann O, Holzhütter HG. Physiology-based kinetic modeling of neuronal energy metabolism unravels the molecular basis of NAD(P)H fluorescence transients. J Cereb Blood Flow Metab. 2015;35:1494–506. https://doi.org/10.1038/jcbfm.2015.70.
Yu G, He QY. ReactomePA: an R/bioconductor package for reactome pathway analysis and visualization. Mol Biosyst. 2016;12:477–9. https://doi.org/10.1039/c5mb00663e
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7. https://doi.org/10.1089/omi.2011.0118.
Huang ZX, Chen Y, Guo HR, Chen GF. Systematic review and bioinformatic analysis of microrna expression in autism spectrum disorder identifies pathways associated with cancer, metabolism, cell signaling, and cell adhesion. Front Psychiatry. 2021;12:630876 https://doi.org/10.3389/fpsyt.2021.630876
Hetman M, Slomnicki LP. Ribosomal biogenesis as an emerging target of neurodevelopmental pathologies. J Neurochem. 2019;148:325–47. https://doi.org/10.1111/jnc.14576.
Lindström MS, Jurada D, Bursac S, Orsolic I, Bartek J, Volarevic S. Nucleolus as an emerging hub in maintenance of genome stability and cancer pathogenesis. Oncogene. 2018;37:2351–66. https://doi.org/10.1038/s41388-017-0121-z.
Biever A, Valjent E, Puighermanal E. Ribosomal protein S6 phosphorylation in the nervous system: from regulation to function. Front Mol Neurosci. 2015;8:75 https://doi.org/10.3389/fnmol.2015.00075
Kim TS, Jang CY, Kim HD, Lee JY, Ahn BY, Kim J. Interaction of Hsp90 with ribosomal proteins protects from ubiquitination and proteasome-dependent degradation. Mol Biol Cell. 2006;17:824–33. https://doi.org/10.1091/mbc.e05-08-0713.
Yano T, Ferlito M, Aponte A, Kuno A, Miura T, Murphy E, et al. Pivotal role of mTORC2 and involvement of ribosomal protein S6 in cardioprotective signaling. Circ Res. 2014;114:1268–80. https://doi.org/10.1161/CIRCRESAHA.114.303562.
Stetler RA, Gao Y, Signore AP, Cao G, Chen J. HSP27: mechanisms of cellular protection against neuronal injury. Curr Mol Med. 2009;9:863–72. https://doi.org/10.2174/156652409789105561
Concannon CG, Orrenius S, Samali A. Hsp27 inhibits cytochrome c-mediated caspase activation by sequestering both pro-caspase-3 and cytochrome c. Gene Expr. 2001;9:195–201. https://doi.org/10.3727/000000001783992605
de Agustín-Durán D, Mateos-White I, Fabra-Beser J, Gil-Sanz C. Stick around: cell-cell adhesion molecules during neocortical development. Cells. 2021;10:118 https://doi.org/10.3390/cells10010118
Redies C, Hertel N, Hübner CA. Cadherins and neuropsychiatric disorders. Brain Res. 2012;1470:130–44. https://doi.org/10.1016/j.brainres.2012.06.020.
Mastrolia V, Al Massadi O, de Pins B, Girault JA. Pyk2 in dorsal hippocampus plays a selective role in spatial memory and synaptic plasticity. Sci Rep. 2021;11:16357 https://doi.org/10.1038/s41598-021-95813-x
Zhu X, Bao Y, Guo Y, Yang W. Proline-rich protein tyrosine kinase 2 in inflammation and cancer. Cancers. 2018;10:139 https://doi.org/10.3390/cancers10050139
Lancaster MA, Knoblich JA. Spindle orientation in mammalian cerebral cortical development. Curr Opin Neurobiol. 2012;22:737–46. https://doi.org/10.1016/j.conb.2012.04.003.
Vadodaria KC, Brakebusch C, Suter U, Jessberger S. Stage-specific functions of the small Rho GTPases Cdc42 and Rac1 for adult hippocampal neurogenesis. J Neurosci. 2013;33:1179–89. https://doi.org/10.1523/JNEUROSCI.2103-12.2013
Benayoun BA, Caburet S, Veitia RA. Forkhead transcription factors: key players in health and disease. Trends Genet. 2011;27:224–32. https://doi.org/10.1016/j.tig.2011.03.003.
McLaughlin CN, Broihier HT. Keeping neurons young and foxy: FoxOs promote neuronal plasticity. Trends Genet. 2018;34:65–78. https://doi.org/10.1016/j.tig.2017.10.002
Paik JH, Ding Z, Narurkar R, Ramkissoon S, Muller F, Kamoun WS, et al. FoxOs cooperatively regulate diverse pathways governing neural stem cell homeostasis. Cell Stem Cell. 2009;5:540–53. https://doi.org/10.1016/j.stem.2009.09.013
Besnard A, Galan-Rodriguez B, Vanhoutte P, Caboche J. Elk-1 a transcription factor with multiple facets in the brain. Front Neurosci. 2011;5:35 https://doi.org/10.3389/fnins.2011.00035
Pagani M, Barsotti N, Bertero A, Trakoshis S, Ulysse L, Locarno A, et al. mTOR-related synaptic pathology causes autism spectrum disorder-associated functional hyperconnectivity. Nat Commun. 2021;12:6084 https://doi.org/10.1038/s41467-021-26131-z
Wang H, Doering LC. Reversing autism by targeting downstream mTOR signaling. Front Cell Neurosci. 2013;7:28 https://doi.org/10.3389/fncel.2013.00028
Tyler WA, Gangoli N, Gokina P, Kim HA, Covey M, Levison SW, et al. Activation of the mammalian target of rapamycin (mTOR) is essential for oligodendrocyte differentiation. J Neurosci. 2009;29:6367–78. https://doi.org/10.1523/JNEUROSCI.0234-09.2009
Kitagishi Y, Kobayashi M, Kikuta K, Matsuda S. Roles of PI3K/AKT/GSK3/mTOR pathway in cell signaling of mental illnesses. Depress Res Treat. 2012;2012:752563 https://doi.org/10.1155/2012/752563.
Magdalon J, Sánchez-Sánchez SM, Griesi-Oliveira K, Sertié AL. Dysfunctional mTORC1 signaling: a convergent mechanism between syndromic and nonsyndromic forms of autism spectrum disorder? Int J Mol Sci. 2017;18:659 https://doi.org/10.3390/ijms18030659
Morita M, Gravel SP, Hulea L, Larsson O, Pollak M, St-Pierre J, et al. mTOR coordinates protein synthesis, mitochondrial activity and proliferation. Cell Cycle. 2015;14:473–80. https://doi.org/10.4161/15384101.2014.991572
Frye RE. RASopathies: a window into the molecular mechanisms underlying neurodevelopmental disorders. Dev Med Child Neurol. 2015;57:315–6. https://doi.org/10.1111/dmcn.12626.
Ash RT, Buffington SA, Park J, Suter B, Costa-Mattioli M, Zoghbi HY, et al. Inhibition of elevated Ras-MAPK signaling normalizes enhanced motor learning and excessive clustered dendritic spine stabilization in the MECP2-duplication syndrome mouse model of autism. eNeuro. 2021;8:ENEURO.0056–21.2021. https://doi.org/10.1523/ENEURO.0056-21.2021
Fong DK, Rao A, Crump FT, Craig AM. Rapid synaptic remodeling by protein kinase C: reciprocal translocation of NMDA receptors and calcium/calmodulin-dependent kinase II. J Neurosci. 2002;22:2153–64. https://doi.org/10.1523/JNEUROSCI.22-06-02153.2002
Hansel C. Deregulation of synaptic plasticity in autism. Neurosci Lett. 2019;688:58–61. https://doi.org/10.1016/j.neulet.2018.02.003
Parsons S, Parsons J. Src family kinases, key regulators of signal transduction. Oncogene. 2004;23:7906–9. https://doi.org/10.1038/sj.onc.1208160
Hacohen-Kleiman G, Sragovich S, Karmon G, Gao AYL, Grigg I, Pasmanik-Chor M, et al. Activity-dependent neuroprotective protein deficiency models synaptic and developmental phenotypes of autism-like syndrome. J Clin Investig. 2018;128:4956–69. https://doi.org/10.1172/JCI98199.
Grigg I, Ivashko-Pachima Y, Hait TA, Korenková V, Touloumi O, Lagoudaki R, et al. Tauopathy in the young autistic brain: novel biomarker and therapeutic target. Transl Psychiatry. 2020;10:228 https://doi.org/10.1038/s41398-020-00904-4
Ivashko-Pachima Y, Hadar A, Grigg I, Korenková V, Kapitansky O, Karmon G, et al. Discovery of autism/intellectual disability somatic mutations in Alzheimer’s brains: mutated ADNP cytoskeletal impairments and repair as a case study. Mol Psychiatry. 2021;26:1619–33. https://doi.org/10.1038/s41380-019-0563-5.
Steiner P. Brain fuel utilization in the developing brain. Ann Nutr Metab. 2019;75(Suppl 1):S8–S18.
Varum S, Rodrigues AS, Moura MB, Momcilovic O, Easley CA, Ramalho-Santos J, et al. Energy metabolism in human pluripotent stem cells and their differentiated counterparts. PLoS One. 2011;6:e20914.
Goyal MS, Hawrylycz M, Miller JA, Snyder AZ, Raichle ME. Aerobic glycolysis in the human brain is associated with development and neotenous gene expression. Cell Metab. 2014;19:49–57.
Yoxall CW, Weindling AM. Measurement of cerebral oxygen consumption in the human neonate using near infrared spectroscopy: cerebral oxygen consumption increases with advancing gestational age. Pediatr Res. 1998;44:283–90.
Gleason CA, Hamm C, Jones MD Jr. Cerebral blood flow, oxygenation, and carbohydrate metabolism in immature fetal sheep in utero. Am J Physiol. 1989;256:R1264–8.
Altman DI, Perlman JM, Volpe JJ, Powers WJ. Cerebral oxygen metabolism in newborns. Pediatrics. 1993;92:99–104.
Walls AB, Waagepetersen HS, Bak LK, Schousboe A, Sonnewald U. The glutamine-glutamate/GABA cycle: function, regional differences in glutamate and GABA production and effects of interference with GABA metabolism. Neurochem Res. 2015;40:402–9.
Manco M, Guerrera S, Ravà L, Ciofi Degli Atti M, Di Vara S, et al. Cross-sectional investigation of insulin resistance in youths with autism spectrum disorder. Any role in reduction brain glucose Metab? Transl Psychiatry. 2021;11:229 https://doi.org/10.1038/s41398-021-01345-3
Needham BD, Adame MD, Serena G, Rose DR, Preston GM, Conrad MC, et al. Plasma and fecal metabolite profiles in autism spectrum disorder. Biol Psychiatry. 2021;89:451–62.
Smith AM, Natowicz MR, Braas D, Ludwig MA, Ney DM, Donley ELR, et al. A metabolomics approach to screening for autism risk in the Children’s autism metabolome project. Autism Res. 2020;13:1270–85.
Zou M, Li D, Wang L, Li L, Xie S, Liu Y, et al. Identification of amino acid dysregulation as a potential biomarker for autism spectrum disorder in China. Neurotox Res. 2020;38:992–1000.
Gevi F, Zolla L, Gabriele S, Persico AM. Urinary metabolomics of young Italian autistic children supports abnormal tryptophan and purine metabolism. Mol Autism. 2016;7:47.
Ming X, Stein TP, Brimacombe M, Johnson WG, Lambert GH, Wagner GC. Increased excretion of a lipid peroxidation biomarker in autism. Prostaglandins Leukot Ess Fat Acids. 2005;73:379–84.
Goh S, Dong Z, Zhang Y, DiMauro S, Peterson BS. Mitochondrial dysfunction as a neurobiological subtype of autism spectrum disorder: evidence from brain imaging. JAMA Psychiatry. 2014;71:665–71.
Frye RE. Mitochondrial dysfunction in autism spectrum disorder: unique abnormalities and targeted treatments. Semin Pediatr Neurol. 2020;35:100829.
Al-Otaish H, Al-Ayadhi L, Bjorklund G, Chirumbolo S, Urbina MA, El-Ansary A. Relationship between absolute and relative ratios of glutamate, glutamine and GABA and severity of autism spectrum disorder. Metab Brain Dis. 2018;33:843–54.
Shimmura C, Suda S, Tsuchiya KJ, Hashimoto K, Ohno K, Matsuzaki H, et al. Alteration of plasma glutamate and glutamine levels in children with high-functioning autism. PLoS One. 2011;6:e25340.
Shinohe A, Hashimoto K, Nakamura K, Tsujii M, Iwata Y, Tsuchiya KJ, et al. Increased serum levels of glutamate in adult patients with autism. Prog Neuropsychopharmacol Biol Psychiatry. 2006;30:1472–7.
Karmon G, Sragovich S, Hacohen-Kleiman G, Ben-Horin-Hazak I, Kasparek P, Schuster B, et al., Novel ADNP syndrome mice reveal dramatic sex-specific peripheral gene expression with brain synaptic and tau pathologies. Biol Psychiatry. 2021: S0006-3223(21)01630-9. https://doi.org/10.1016/j.biopsych.2021.09.018.
Malishkevich A, Amram N, Hacohen-Kleiman G, Magen I, Giladi E, Gozes I. Activity-dependent neuroprotective protein (ADNP) exhibits striking sexual dichotomy impacting on autistic and Alzheimer’s pathologies. Transl Psychiatry. 2015;5:e501 https://doi.org/10.1038/tp.2014.138
Amram N, Hacohen-Kleiman G, Sragovich S, Malishkevich A, Katz J, Touloumi O, et al. Sexual divergence in microtubule function: the novel intranasal microtubule targeting SKIP normalizes axonal transport and enhances memory. Mol Psychiatry. 2016;21:1467–76. https://doi.org/10.1038/mp.2015.208
Acknowledgements
This work was supported by Psychiatry Research Foundation, Region Southern Denmark and Jascha Foundation.
Author information
Authors and Affiliations
Contributions
MI, BIA, HSW, TMM conceived and designed the experiments. MI, BIA, KTV, SH, TWW, and RL performed the experiments. MI, BIA, KTV, SH, TWW, and RL analyzed the data, generated figures, and wrote the manuscript. MI and TMM directed and supervised the project. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ilieva, M., Aldana, B.I., Vinten, K.T. et al. Proteomic phenotype of cerebral organoids derived from autism spectrum disorder patients reveal disrupted energy metabolism, cellular components, and biological processes. Mol Psychiatry 27, 3749–3759 (2022). https://doi.org/10.1038/s41380-022-01627-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-022-01627-2
This article is cited by
-
Human pluripotent stem cell (hPSC) and organoid models of autism: opportunities and limitations
Translational Psychiatry (2023)