Abstract
Suicide is a significant public health concern with complex etiology. Although the genetic component of suicide is well established, the scope of gene networks and biological mechanisms underlying suicide has yet to be defined. Previously, we reported genome-wide evidence that neurexin 1 (NRXN1), a key synapse organizing molecule, is associated with familial suicide risk. Here we present new evidence for two non-synonymous variants (rs78540316; P469S and rs199784139; H885Y) associated with increased familial risk of suicide death. We tested the impact of these variants on binding interactions with known partners and assessed functionality in a hemi-synapse formation assay. Although the formation of hemi-synapses was not altered with the P469S variant relative to wild-type, both variants increased binding to the postsynaptic binding partner, leucine-rich repeat transmembrane neuronal 2 (LRRTM2) in vitro. Our findings indicate that variants in NRXN1 and related synaptic genes warrant further study as risk factors for suicide death.
Similar content being viewed by others
Introduction
Suicide is the 10th leading cause of death in the U.S and accounts for almost 800,000 deaths per year globally [1]. Suicide is preventable but its complex etiology requires further study. While the body of evidence for genes and biological mechanisms underlying suicide is growing, the genetics of suicide death are complex and remain elusive. Many researchers study suicidal behaviors, which are more common than suicide deaths [2]. However, most individuals with suicidal behaviors do not die by suicide, making the extent of shared risk factors between suicidal behaviors and suicide death uncertain [2].
There is evidence of the genetic contribution to suicide death; heritability estimates for suicide is ~50% [3, 4]. Previous studies of large, multi-generation families indicate that the risk of suicidal death extends beyond first-degree relatives [5, 6]. This extended familial risk further implicates genetic factors and not just shared environmental factors between close relatives [5].
The most widely used approach for the identification of suicide genetic risk variants is genome-wide association studies (GWAS) [7,8,9,10,11]. Limitations of GWAS include the genetic heterogeneity inherent in large population cohorts, which may limit statistical power for less common variants [12], and the variants identified are often merely markers for genomic locations (usually intergenic) and seldom impact gene function. Hence, there is a knowledge gap of rare, functional risk variants, which are also important to the study of complex diseases [13].
Our previous novel analyses of 215 suicide deaths in 43 extended high-risk families (7–9 generations) [6] identified genes of interest that allow us to pursue targeted follow-up of rare functional variants in an additional large, population-ascertained cohort of suicide deaths. The extended family design has increased power to detect shared genomic regions associated with risk due to the repetition of the same risk factors across many distantly related suicides. Significantly shared familial regions are highly likely to harbor variants that increase risk of suicide death. Genes in these shared regions are therefore excellent targets for follow-up in complementary designs.
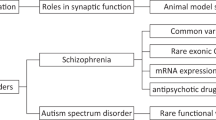
Figure 1 presents our study design. From the 30 significant familially-shared segments previously reported [6], we prioritized a genomic segment from the largest studied family (#601627), which contained a single gene-NRXN1. Other segments contained multiple genes, making it difficult to identify the true risk gene(s) for follow-up work. Prioritization was also based on the known functions of NRXN1. Synapse pathology is thought to underlie neurological and psychiatric disorders [14]. Synapse dysfunction is also an important potential target for prevention and therapeutic intervention. NRXN1 codes for a transmembrane cell adhesion molecule that plays important roles in synaptic assembly and organization, specification, neurotransmission, and synaptic plasticity in both inhibitory and excitatory synapses [15,16,17,18]. NRXN1 can also be prioritized due to its psychiatric relevance, highlighted in a recent review by Hu et al. [19]. Previous studies showed that several missense and structural variations in NRXN1 are associated with autism, schizophrenia, developmental abnormalities, and psychiatric drug response [16, 19,20,21,22,23,24,25,26,27]. There is also evidence that brain expression of NRXN1 is altered in individuals with bipolar disorder or schizophrenia who died by suicide [28, 29]. A synaptic binding partner of NRXN1, LRRTM4, is also implicated in increased suicide risk [30, 31]. Few studies have tested the functional consequences of specific disease-associated NRXN1 variants. Variants found in autism and schizophrenia were tested in human-induced neurons and resulted in alterations in calcium signaling, impairment of synaptic function, and changes in cell fate [32,33,34].
The table summarizes the number of familial and non-familial cases genotyped and analyzed in each study and serves as a reference for the study design. A total of 4382 cases were genotyped for both studies. The analysis strategy started with the prioritization of NRXN1, from a previous study of high-risk families (red arrow), led to the discovery of specific NRXN1 suicide variants (blue arrow) and finally resulted in the elucidation of their functional consequences (blue box). 1Coon H, Darlington T, DiBlasi E, Callor W, Ferris E, Fraser A et al. Genome-wide significant regions in 43 Utah high-risk families implicate multiple genes involved in risk for completed suicide. Mol Psychiatry. 2020;25:3077-90.
In this study, our objectives were to identify NRXN1 variants, in our suicide death cohort, with likely functional consequences on the gene and test their function. We used whole-genome sequencing to search for likely causal functional variants in NRXN1 shared across the six related suicides in the original study [6], supporting the genome-wide significant result. Additionally, we screened genotyping data from 4376 other Utah suicide deaths in our cohort, excluding the six original familial cases, to identify other functional NRXN1 variants. We aimed to identify NRXN1 variants that were at a sufficient frequency in suicide cases to warrant functional analysis (frequency of at least 0.001) and that were rare (frequency < 0.01) in population controls. We also used our suicide death sample to identify functional NRXN1 variants that segregated in other identified high-risk suicide families. To accomplish this analysis, we used genealogical data from a total of 241 ascertained high-risk families. These 241 high-risk families include the 43 described in our previous study and include approximately half of the 4376 genotyped cases (2219 cases) in our suicide death cohort. This approach allowed us to target two functional NRXN1 variants with additional evidence of familial transmission. Finally, we tested the functional consequences of the two identified NRXN1 suicide variants on NRXN1 binding interactions to known partners and in hemi-synapse formation.
Materials/subjects and methods
The Utah suicide research resource
In collaboration with the Utah State Office of the Medical Examiner (OME), and with Institutional Review Board (IRB) permissions from the University of Utah, Utah Department of Health, and Intermountain Healthcare, we have been collecting de-identified DNA samples from suicide deaths since 1997. Suicide status is conservatively determined by the OME based on investigation of the scene of death, interviews with kin and friends of the deceased, review of medical and public records concerning the individual, and autopsy and toxicology reports. Suicide deaths are securely linked to medical information and family records via the Utah Population Database (UPDB), a data resource with state-wide demographic and health information, and with extended genealogical records [35].
Genealogical data allowed the determination of high-risk families through the comparison of the total number of suicide deaths in the family vs. the expected number in the family, accounting for known population rates of suicide death [6]. Our previous study prioritized 43 of the highest-risk families containing 215 suicide deaths [6]. However, in our data resource, a total of 241 high-risk families that meet a familial significance threshold of p < 0.01 have now been ascertained. These families include 2219 Utah suicide deaths with genotyping data (50.71% of the sample). The genealogical information in our data resource not only allowed for prioritization of specific regions, including the region with NRXN1 that is the focus of the follow-up, but the information additionally allowed for prioritization of single functional variants that show familial transmission.
Psychiatric diagnoses on Utah suicide deaths from electronic medical record (EMR) data are available for ~80% of suicide deaths with genotyping data. The Utah suicide research sample is population-based and reflects suicide deaths that crosscut diagnoses.
Our data also allows for testing background polygenic risk, through the derivation of polygenic risk scores (PRS) [7]. We tested for NRXN1 variant carrier vs. non-carrier differences in polygenic risks for a subset of diagnostic and behavioral traits hypothesized to have NRXN1 associations: suicide attempt, major depressive disorder, depressive symptoms, schizophrenia, autism, bipolar disorder, anxiety disorder, attention deficit hyperactivity disorder, alcohol abuse, smoking, post-traumatic stress disorder, neuroticism, and extraversion. Polygenic risk scores for these traits had already been computed for our GWAS study [7]. To test for carrier vs. non-carrier differences, we used logistic regression, adjusting for genetically determined ancestry principal components derived as part of our GWAS [7].
Genotyping
In total, 4382 cases (215 in the previously analyzed high-risk extended families and 4167 additional cases) were genotyped using the genome-wide Illumina PsychArray, an array with 265,000 tag single nucleotide polymorphisms (SNPs), 245,000 markers from Illumina Exome BeadChip, and 50,000 additional markers with clinical or psychiatric relevance [36]. The SNPs were aligned in the forward strand HG19 configuration by comparing the SNP calls within the 1000 Genomes Project [37]. SNPs were removed if the forward strand status was unclear, or if variants were not polymorphic, and non-autosomal. Additional quality control of genotyping was done using PLINK [38], as previously described [6].
Sequencing
To search for potentially causal risk variants in the 0.92 Mb prioritized candidate region containing NRXN1, we generated sequence data for the six suicide deaths in the previously studied family that shared this region (#601627). Whole-genome sequencing was performed on cases using Illumina NGS technology [39] using DNA from blood at 30x-60x coverage. We used BWA-MEM [40] for mapping sequencing reads to the human genome reference sequence and for variant detection and calling, we used best practice GATK-based procedures [41].
Genetic analyses
We used the GEMINI software tool [42] to search for causal single nucleotide functional variants directly shared in family #601627, and the LUMPY tool [43] to search for shared structural variants. Single nucleotide variants of interest were defined as those in the 0.92 Mb shared region where the minor allele was present in all six cases, the minor allele frequency was <0.01 in 525 Utah controls and publicly-available data (gnomAD; 44), and the variant had medium or high impact severity annotations. The 525 Utah controls used in this analysis included 160 unrelated Utah individuals selected for the absence of disease who were in the CEPH genetic mapping sample [44], 78 unrelated healthy Utahns over age 90 from a study of extreme longevity [45], and 287 1000 Genomes Utah samples [37]. We used SIFT (sorting intolerant from tolerant; [46]) and PolyPhen (polymorphism phenotyping; [47]) to determine the predicted functional consequences of the missense variants we discovered. The algorithms for these computational tools are based on variables such as whether the variants occur in conserved regions, the physical and chemical properties of the amino acids involved, and the structural properties of a protein. All amino acid substitutions across the proteome are assessed to determine which mutations are likely to result in pathogenicity. We searched for additional evidence of NRXN1 functional risk variants using 4376 additional Utah suicides with Illumina PsychArray genotyping. These analyses included 209 cases in the original extended family study who were not related to family #601627 and 4167 new Utah cases. Inclusion of the 209 original cases is justified because this analysis is focused on single NRXN1 functional variants which could occur on a sufficiently small haplotype to escape detection in our original familial analysis, or variants may be shared across too few cases within an extended family to be detected in the original analysis. We analyzed functional NRXN1 variants that spanned the entire gene, comparing to frequencies in Non-Finnish European control samples from 1000 Genomes and gnomAD. Because our original evidence suggested that NRXN1 may be associated with familial suicide risk, we not only tested for case-control frequency differences but also used our extensive genealogical data to estimate the nominal probability of familial segregation of single functional variants. Segregation probabilities were computed by considering variant frequency, number of meioses between sharing cases, and the chance of introduction of the variant through a marry-in spouse vs. transmission at each generation (see Supplementary Methods, and Supplementary Fig. 1 for details). Variants showing significant familial segregation with suicide death were pursued for further functional analysis.
Plasmids
N-terminally myc-tagged LRRTM2-4 expression constructs were previously described [48] and gifted by Dr. Joris de Wit (Katholieke Universiteit Leuven, Belgium). pCAG-HA-rat Nrxn1α, lacking alternatively spliced segment 4 (-AS4), was generated by Dr. Peter Scheiffele [49] and obtained from Addgene (Addgene plasmid # 58266; http://n2t.net/addgene:58266; RRID:Addgene_58266). To generate the wild-type HA-Nrxn1α-ecto-Fc construct, the HA-rat-Nrxn1α extracellular domain was PCR amplified from the pCAG-HA-ratNrxn1α (-AS4) construct and subcloned to be in frame with pTEV-Fc containing a human IgG Fc. To generate single variant Fc (P469S or H885Y) and double mutated (P469S/D1216A or H885Y/D1216A) Fc constructs, we used site-directed-mutagenesis (Q5® Site-Directed Mutagenesis Kit, New England Biolabs) and constructs were verified by sequencing. The D1216A mutation prevents Ca2+-binding at laminin-neurexin-sex hormone-binding globulin domain 6 (LNS6). For full-length Nrxn1α variant constructs, similar mutagenesis procedures were performed, and the full-length Nrxn1α (-AS4) regions were then subcloned back into the original pCAG vector using standard PCR cloning and verified by sequencing. Plasmids for rat neurexophilin 1 (Nxph1; pCMVD2), dystroglycan (DAG; pCMVDAG), neuroligin 1 (Nlgn1; pCMVNL1-B), neuroligin 2 (Nlgn2; pCMVNL2-1) and for like-acetyl-glucosaminyl-transferase (LARGE; pCMV6-XL4 LARGE), have been previously described [50]. Full-length YFP-LRRTM2 used in Fig. 2f was a gift from Fredrik Sterky (SciLifeLab, Stockholm, Sweden).
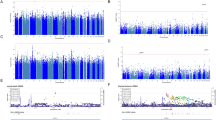
a Schematic of Nrxn1α protein lacking insert at splice site #4 (SS#4), including locations of the suicide-associated variants identified in this study (magenta). Laminin neurexin sex-hormone binding globulin domain (LNS); epidermal growth factor-like domain (EGF); transmembrane region (TM); O-glycosylation sequence (oGlyc); heparan sulfate (HS); PDZ bs.; PSD-95/Dlg/ZO-1 binding site. b In the left panel, ribbon diagram of variant P469S (magenta) in the αLNS2 domain, and binding of Nxph1 (green). In the right panel, stick model of variant H885Y, proximal to the Ca2+-binding site at the LNS4 domain. c Western blot of Nrxn1α and Nxph1 pull-downs. Note that neither variant alters binding. d Western blot of pull-down of α-DAG and β-DAG with the extracellular Fc-tagged domain of Nrxn1α shows undisturbed DAG binding with wild-type and variant Nrxn1α. e Quantification of 2d (n = 3); n.s. indicates not significant by one-way ANOVA. f Western blot pull-down shows Nrxn1α variants bind Nlgn1 and Nlgn2 at similar levels to WT Nrxn1α but the P469S variant binds YFP-LRRTM2 at increased levels to WT Nrxn1α. g Quantification of 2f (n = 3); One-way ANOVA using GraphPad Prism followed by pairwise posttests. *p < 0.05, n.s. indicates not significant.
Cell culture
Cell cultures were kept in a humidified incubator and maintained at 37° C and 5% CO2. Cell lines used: Chinese Hamster Ovary (CHO-K1) cells (RRID: CVCL_0214, female) were grown in F12K media (Life Technologies, Carlsbad, CA, United States), 10% FBS (Fetal Bovine Serum) (Life Technologies), and penicillin/streptomycin; HEK293 cells (RRID: CVCL_0063, female) were grown in DMEM (Life Technologies), 10% FBS, and penicillin/streptomycin. Cells were transfected using polyethylenimine (PEI, Polysciences, Warrington, PA, United States) at a ratio of 10 mg PEI/1mg DNA for CHO cells, 4 mg PEI/1mg DNA for 293 cells used in the expression of Fc proteins, and 10 mg PEI/1mg DNA for 293 cells transfected for the synapse formation assay.
Neurons: preparation of hippocampal rat cultures were previously described [51, 52]. Briefly, P0 rat cortical glia were cultured to form monolayers on Poly-D-Lysine/collagen-coated coverslips in 24-well plates. P0 rat hippocampi from Sprague Dawley rats of both sexes (RRID: RGD_734476) were dissected one week later in cold 4-(2-hydroxyethyl)-1-piperazinee-thanesulfonic acid (HEPES)-buffered saline solution (Life Technologies), treated with enzymatic papain for 30 min, dissociated, and plated onto glial monolayers at a density of 5 × 104 cells/well.
Fc-binding assay
Wild-type (WT), P469S, and H885Y HA-Nrxn1α-ecto-Fc constructs were transfected into HEK293 cells using PEI and proteins were extracted from the media based on previously described methods [51]. Briefly, transfected cells were incubated in OptiMEM (Life Technologies) media for 5 days. Then, the conditioned media was harvested, concentrated, and filtered using 100 kD Amicon Ultra filter units (Millipore, Billerica, MA, United States). Fc protein concentrations were estimated by Western blot (Supplementary Fig. 2a) using known concentrations of purified human Fc (Jackson ImmunoResearch, West Grove, PA, United States) as a standard. Fc proteins were run on 8% SDS-PAGE gels and transferred to nitrocellulose membranes using the iBlot system (Life Technologies). Membranes were incubated in blocking solution (50 mM Tris pH 7.5, 300 mM NaCl, 3% wt/vol dry milk powder, and 0.05% Tween-20) for 1 h, and, incubated in HRP-conjugated secondary antibody (Jackson ImmunoResearch) for 1 h at room temperature. The Bio-Rad Clarity ECL kit on a Bio-Rad ChemiDoc XRS+ imaging system was used for protein detection. Nrxn1α WT-ecto-Fc, Nrxn1α variant-ecto-Fc, and control-Fc (at ~1 μg/ml concentrations) were then added to GFP-only and LRRTM + GFP co-expressing CHO cells 24 h after transfection with PEI. Cells were incubated on ice for 30 min to facilitate binding of the Fc proteins. Following Fc treatment, the cells were washed gently with 1x Phosphate Buffered Saline (PBS) (Life Technologies) and fixed with 4% paraformaldehyde (PFA).
Biochemical procedures and pull-down assays
DNA constructs coding for soluble ecto-Fc of WT or mutated Nrxn1α, were transfected into HEK293 cells using calcium phosphate as described previously [50]. Similarly, co-secreted complexes of Nxph1 with WT or variant Nrxn1-Fc were harvested after calcium phosphate co-transfection of constructs. To obtain recombinant full-length αβDAG, DNA constructs coding for αβDAG and LARGE were co-transfected in N2A cells using calcium phosphate, and cells were lysed 72 h after transfection using 1% Triton in Tris-buffer (50 mM Tris pH 7.5, 80 mM NaCl, 5 mM CaCl2, protease inhibitor cocktail III, Merck) for 30 min at 4 °C. After centrifugation (1 min, 21,000 × g, 4 °C) supernatant was used immediately or frozen by liquid nitrogen and stored at −80 °C. Full-length Nlgn1, Nlgn2, or LRRTM2 were expressed in COS7 cells using DEAE-dextran transfection, based on previously described methods [50] and cells were incubated for 2 days before lysis using 1% Triton in Tris-buffer, as described above.
For pull-down assays, Protein A-conjugated Sepharose beads were added to either media containing co-secreted Nxph1/Nrxn1α-Fc complexes or cell lysates, incubated overnight, washed three times, and analyzed directly. N2A lysates with DAG were incubated with Nrxn1α-bound Protein A-beads overnight in the presence of 5mM CaCl2. Similarly, COS lysates containing either Nlgn1, Nlgn2, or YFP-LRRTM2 were also incubated with Nrxn1α-bound Protein A-beads overnight in the presence of 5mM CaCl2. Complexes were centrifugated (30 s, 11,000 × g, 4 °C), washed three times, boiled for 5 min in sample buffer, and analyzed by Western Blot using stain-free 4–15% gradient gels and the Mini-Protean system (Bio-Rad, Feldkirchen, Germany). Gray values of each blot have been normalized to input values which were detected by UV using a stain-free gel (Bio-Rad), anti-Fc antibody (I9135, Sigma), and anti-HA antibody (16B12, Biolegend) for HA-Nrxn1a-Fc variants (n = 3). Intensities have been measured using ImageJ and analyzed in a one-way ANOVA using GraphPad Prism.
Immunocytochemistry
Cells were fixed in 4% PFA for 10 min and washed three times with 1x PBS. Cells were blocked for 30 min in blocking buffer (1x PBS + 3%BSA + 0.1% Triton-X100). Primary antibodies were diluted in blocking buffer without Triton-X100 and cells were incubated for 2 h at room temperature with the primary antibodies. The following primary antibodies were used: goat anti-GFP 1: 3000, (RRID:AB_305643, Abcam, Cambridge, MA, United States), rabbit anti-myc 1:1000 (RRID:AB_439680, Sigma Aldrich, St. Louis, MO, United States), chicken anti-MAP2 1:5000 (RRID:AB_2138153, Abcam), guinea pig anti-vGLUT1 1:2000 (RRID:AB_2301751, Millipore), mouse anti-PSD-95 1:1000 (RRID:AB_2315909, NeuroMab, UC Davis/NIH NeuroMAB Facility, Davis, CA, United States), guinea pig anti-vGAT 1:1000 (RRID: AB_212625, Millipore), mouse anti-gephyrin 1:1000 (RRID:AB_2619837, Synaptic Systems, Goettingen, Germany). After incubation with primary antibodies, cells were washed three times and incubated in the dark with secondary antibodies for 1 h, at room temperature. Cy3-conjugated donkey anti-human IgG 1:1000 (RRID:AB_2340534, Jackson ImmunoResearch) was used to visualize Fc proteins. All secondary antibodies were donkey-derived and diluted 1:1000 (Jackson ImmunoResearch) and included, donkey anti-chicken-DyLight 405 (RRID:AB_2340373), donkey anti-goat-Alexa 488 (RRID:AB_2336933), donkey anti-guinea pig Alexa 647 (RRID: AB_2340476), and donkey anti-rabbit Alexa 647 (RRID: AB_2492288). Slides were prepared for imaging with Fluoromount-G (Southern Biotech, Birmingham, AL, United States).
Hemi-synapse formation assay
Mixed-culture assays for hemi-synapse formation were performed based on standard methods previously described [53]. Briefly, HEK293 cells were transfected, using PEI, with GFP-only, GFP+ WT full-length HA-Nrxn1α, GFP + P469S HA-Nrxn1α, and GFP+ H885Y full-length HA-Nrxn1α constructs. 24 hours later, the transfected HEK293 cells were washed and dissociated in neuronal feeding media and seeded at 3 × 104 cells/well in a 24-well plate of primary neuron cultures grown on a glia monolayer (DIV 10). The co-cultures were incubated for 36 h and fixed on DIV 12 in 1x PBS + 4% PFA for 10 min at room temperature.
Confocal imaging and analysis
Z-stack images were captured using a Zeiss LSM 710 confocal microscope at 40x to observe Fc binding in CHO cells. The maximum intensity projection of the Z-stack was analyzed using NIH ImageJ software under blind conditions. For the Fc-binding assay analysis, CHO cells were traced in the 647 nm channel based on myc-LRRTM immunoreactivity. The traced cell area selection was expanded by 1 µm to establish an ROI for each cell. The ROI area and mean-gray values (MGV) were then measured. The MGV and ROI areas of each respective cell were also measured in the 555 nm channel, which detected Fc immunoreactivity. To account for expression differences in the CHO cells, the ratio of the MGV of both channels was calculated (Fc MGV/LRRTM MGV). Values across all conditions were analyzed using a one-way ANOVA analysis in GraphPad Prism software.
For the mixed co-culture assay, HEK293 cells were imaged at 63x and chosen for further analysis if MAP2 staining (405 nm channel) revealed at least one dendrite crossing or touching a GFP-positive HEK293 cell (488 nm channel). Images were de-speckled and optimal thresholds were determined for PSD-95 (555 nm) and vGLUT1 (647 nm) channels, for excitatory synapses, and gephyrin (555 nm) and vGAT channels (647 nm) for inhibitory synapses. The GFP cell filler channel (488 nm) was thresholded and the selected area of a cell was enlarged by 0.5 µm; this selection became the cell ROI, and cell ROI area measurements were recorded. In addition, the remaining channels were thresholded and particle count and particle area measurement were taken for puncta in the 555 nm and 647 nm channels. A constant particle parameter of 0.2–0.8 µm was imposed to define puncta. vGLUT or vGAT puncta that overlapped with PSD-95 and gephyrin puncta respectively were subtracted from the images to remove false-positive neuron-neuron synapses. The puncta count and puncta area of PSD-95-only and gephyrin-only puncta were recorded and indicated induced hemi-synapses. A one-way ANOVA analysis based on the area of hemi-synapse puncta was used for comparison across all conditions in GraphPad Prism software.
Results
NRXN1 sequence variants in six familial cases
From our previous high-risk family study [6], family #601627 showed significant sharing for a chromosome 2 genomic segment containing the single gene NRXN1. The familial analysis method used in this previous study implicated this target gene but was not designed to identify specific risk variants. Therefore, we searched for causal single-nucleotide and structural variants in NRXN1 directly shared across the six suicide deaths in this original high-risk family using whole-genome sequence data. We did not discover any obvious rare, medium, or high impact causal or structural variants shared by these six familial cases indicating regulatory variant(s) must be responsible for the significance in this extended family.
Functional NRXN1 variants from genotyping
Though direct evidence in the discovery high-risk family was not apparent, the familial genome-wide significant result implicating NRXN1 was sufficiently strong to warrant a search for additional risk variants in our full suicide research cohort. Therefore, we analyzed 4376 additional cases that were genotyped on the Illumina PsychArray platform, retaining protein-coding variants in NRXN1. We found two non-synonymous variants where both Polyphen [47] and SIFT [46] annotations indicated likely functional effects, and the minor allele was observed in at least five suicides (frequency of at least 0.001). These variants are GRCh37 2-50847195-G-A; rs78540316; P469S and GRCh37 2-50724817-G-A; rs199784139; H885Y, with annotations for “possibly” and “probably” damaging in Polyphen and “deleterious” in SIFT. Twenty-eight suicide cases were heterozygous for P469S (frequency: 28 / 8752 = 0.0032); the 1000Genomes European frequency of this variant is 29 / 8758 = 0.0033 and 413 / 126558 = 0.0033 in the gnomAD [54] European non-Finnish population. For H885Y, 10 suicide cases were heterozygous (frequency: 10 / 8752 = 0.0011); the 1000 Genomes European frequency of this variant is 11 / 8762 = 0.0013, and the frequency in gnomAD European non-Finnish controls is 134 / 127740 = 0.0011. One case in our cohort carried both risk variants. While the suicide case frequencies were not significantly elevated over population control frequencies in a case-control design, both NRXN1 variants showed familial segregation in extended families with familial risk of p < 0.01. Segregation of P469S occurred in two different high-risk families: one with seven meioses separating the cases, and the other with 11 meioses separating the cases. Segregation of H885Y occurred in one other high-risk family, separated by 12 meioses (see Supplementary Methods, and Supplementary Fig. 1).
Post hoc diagnostic and polygenic risk characterization of suicide deaths with the NRXN1 variants
Cases with the two NRXN1 variants that are the focus of this study had diagnoses across 27 different phenotypic clusters related to psychiatric and medical co-morbidities (Supplementary Table 1). Of the 38 cases with these variants, 19 had evidence of at least one other medical or psychiatric diagnosis. However, no unusual clustering of phenotypes was observed relative to what was observed in our suicide cohort without variants, who had records for diagnoses (2722 cases). The most frequent diagnoses among all cases were chronic pain (15 cases), depression (12 cases), and anxiety (9 cases). Bipolar diagnoses were found in five cases.
Tests for NRXN1 variant carrier vs. non-carrier differences among the polygenic risk scores for suicide attempt, major depressive disorder, depressive symptoms, schizophrenia, autism, bipolar disorder, anxiety disorder, attention deficit hyperactivity disorder, alcohol abuse, smoking, post-traumatic stress disorder, neuroticism, and extraversion did not result in any nominal significant differences.
Suicide-associated variants show increased binding to LRRTM2 in vitro
We next tested whether either rs78540316 (P469S) or rs199784139 (H885Y) altered Nrxn1 function. There are multiple isoforms of Nrxn1, and the variants we identified alter residues in the LNS2 and LNS4 extracellular domains of the α-isoform (Fig. 2a). The extracellular domain is necessary for the binding of Nrxn1 to its many ligands [18, 50]. Thus, we tested the ability of each variant to bind to several Nrxn1α ligands using pull-down assays. Ligands tested included neurexophilin 1 (Nxph1), the dystroglycan complex (DAG), leucine rich repeat transmembrane neuronal 2 (LRRTM2), neuroligin 1 (Nlgn1), and neuroligin 2 (Nlgn2).
Nxph1, a glycoprotein secreted by subpopulations of neurons [50], forms a complex with the extracellular LNS2 domain of Nrxn1α (Fig. 2b, left) in the secretory pathway of cells, and co-secretes into media [50]. Therefore, to test Nxph1 binding with Nrxn1α variants, we co-expressed Nxph1 with the extracellular domain of Nrxn1α fused to an Fc tag in HEK293 cells. The media was collected and the co-secreted recombinant Nxph1/Nrxn1α-ecto-Fc complex was bound to Protein A-beads for western blot analysis (Fig. 2c). Our results indicate that both variants bound to Nxph1 similar to WT (Fig. 2c). This suggests that neither NRXN1 variant interferes with secretion and binding to Nxph1.
The extracellular domain of Nrxn1 also binds the alpha subunit of the adhesion receptor protein DAG ([50]; Supplementary Fig. 3a). α-DAG directly interacts with residues in the LNS2 and LNS6 domains of Nrxn1α and physically covers the intervening LNS3, LNS4, and LNS5 domains (Supplementary Fig. 3a). We reasoned that the suicide-associated Nrxn1α variants could interfere with α-DAG binding because the P469S mutation lies in the LNS2 domain (Fig. 2b, left) and the H885Y mutation is proximal to the Ca2+ binding site within the LNS4 domain of Nrxn1α (Fig. 2b, right). To test DAG binding with suicide-associated Nrxn1α variants, we expressed the DAG complex in N2A cells along with its glycosylating co-factor LARGE, which is necessary for LNS domain binding [50]. We then used Nrxn1α-ecto-Fc pre-bound to Protein A beads as bait to pull down DAG from N2A cell lysates. Western blot analysis of the precipitate revealed that each Nrxn1α variant and WT Nrxn1α bound to DAG at similar levels (Fig. 2d–e). Because DAG binds Nrxn1 in multiple locations, we also tested if weakening the interaction between Nrxn1α and DAG would reveal compromised binding with either of the suicide-associated variants. We did this by combining each variant with a D1216A mutation in Nrxn1α. This mutation prevents DAG from interacting with the LNS6 domain. However, combining the D1216A mutation with either suicide-associated variant did not prevent binding of α-DAG compared to controls (Supplementary Fig. 3b).
Next, we tested binding of Nrxn1α to its trans-synaptic receptors, Nlgn1, Nlgn2 and LRRTM2 (Supplementary Fig. 3c). We did this by binding purified Nrxn1α-ecto-Fc proteins to Protein A beads and incubating them with lysates prepared from COS7 cells expressing each receptor. We found no influence of either suicide-associated variant on binding to Nlgn1 or Nlgn2, but we observed that the P469S variant resulted in increased binding to the LRRTM2 receptor (Fig. 2f–g). In summary, binding of purified Nrxn1α variants to all the ligands tested was similar to WT Nrxn1α binding, with the exception of variant P469S binding to LRRTM2, which was increased relative to WT Nrxn1α binding.
We further tested if the suicide-associated neurexin variants altered binding between Nrxn1 and the LRRTM family using a cell-based assay. This in vitro assay used shorter incubation times than those used in the biochemical assays and a different environmental milieu since these factors could influence binding. Previous studies have shown that LRRTM1 is poorly expressed on the surface of non-neuronal cells [48] so we tested Nrxn1α binding to LRRTM2, 3, and 4. We treated CHO cells expressing myc-tagged LRRTMs with conditioned media containing WT and variant Nrxn1α-Fc proteins (Supplementary Fig. 2a) and Fc-binding was detected by immunofluorescence (Fig. 3a, Supplementary Fig 2b).
a Representative confocal images of Fc cell surface binding assays. b-c Quantification of Fc-binding assays, normalized to control-Fc. b Quantification of experiments using 1.0 ug/mL concentration of Fc proteins. c Quantification of the same assay but using 0.1 and 0.5 ug/mL of P469S-Fc only. (****p < 0.0001, One-way ANOVA with Dunnett’s T3 multiple comparisons tests, *p = 0.02, multiple t tests; n = 39 (WT), n = 48 (P469S), n = 37 (H885Y) cells from three different cultures; ±SEM reported).
Interestingly, we found that both suicide-associated variants had significantly increased binding to LRRTM2 in vitro compared to WT with P469S showing a two-fold increase in binding to LRRTM2 (Fig. 3b). The other conditions tested showed no change from WT (Supplementary Fig. 2c). To determine whether the concentration of Nrxn1α ecto-Fc had any effect on its ability to interact with LRRTM2, we repeated this experiment using one-half and one-tenth the original concentration of Nrxn1α ecto-Fc (Fig. 3c). We observed enhanced binding of Nrxn1α ecto-Fc to LRRTM2 at 0.5 µg/mL (half the concentration used in the initial binding assay), but the effect was diminished at 0.1 ug/mL.
P469S variant induces hemi-synapses similar to WT in vitro
Given that variant P469S showed the strongest increase in binding with LRRTM2, we tested whether this variant had altered hemi-synaptogenic properties compared to WT. Previous studies showed that when non-neuronal cells expressing Nrxn1α (-AS4) are plated with neurons, the neurons will form postsynaptic specializations or hemi-synapses in response to Nrxn1α-expressing cells [53, 55, 56]. Thus, we compared hemi-synapse formation induced by WT Nrxn1α and the P469S variant. Newly formed hemi-synapses between HEK293 cells and neurons were defined by the presence of the excitatory postsynaptic marker PSD-95 or the inhibitory postsynaptic marker gephyrin (Fig. 4a). Although we observed an induction of excitatory synapses (Fig. 4b), we did not observe differences between WT Nrxn1α and the P469S variant in the formation of hemi-synapses (Fig. 4b–c).
a Representative confocal images of the hemi-synapse assay. 293 cells were transfected with GFP (control) or co-transfected with full-length WT+GFP and variant+GFP constructs. MAP2 staining (white) indicates dendrites. For excitatory synapses, presynaptic vGLUT1 (blue) and postsynaptic PSD-95 (red) markers were used. For inhibitory synapses, presynaptic vGAT (blue) and postsynaptic gephyrin (red) markers were used. b-c Quantification of the excitatory and inhibitory hemi-synapse assays. b P469S variant induces excitatory synapses similar to WT(***p = 0.0004, **p = 0.0036, One-way ANOVA analysis with Dunnett’s T3 multiple comparisons tests, ns p > 0.05 for all other comparisons; n = 41 [GFP], n = 33 [WT], n = 29 [P469S] cells from three different cultures; ±SEM reported). c Nrxn1α does not induce inhibitory synapses and variants do not change its activity (n = 46 [GFP], n = 50 [WT], n = 27 [P469S] cells from three different cultures; ±SEM reported); no significant differences by one-way ANOVA.
Discussion
High-risk families are insightful data resources for genetic risk discovery. From previous results of extended high-risk families [6], we prioritized NRXN1 as a target for studies of genetic risk for familial suicide. Subsequent follow-up in 4376 additional suicide deaths identified two functional NRXN1 missense variants with evidence of familial association with suicide. These variants showed increased binding of the Nrxn1 receptor LRRTM2 in vitro. Though more work must be done under physiological conditions, our results provide functional evidence that these variants could alter synaptic signaling.
NRXN1 is a suicide risk factor
NRXN1 and its interacting genes are compelling candidates for suicide risk, as they are associated with a broad spectrum of neurological and psychiatric disorders (autism, schizophrenia, anxiety, depression, bipolar disorder, and attention deficit hyperactive disorder) [17, 19, 25, 57,58,59]. This evidence suggests the importance of NRXN1 to affect multiple manifestations of psychopathology, mirroring suicide risk, which also crosscuts psychiatric diagnoses [5]. The suicide cases carrying the variants we tested in this study exhibited a range of diagnoses, suggesting the two NRXN1 variants may be related to suicide risk that is not confined to a specific psychiatric or clinical subpopulation.
Functional implications of suicide risk variants
NRXN1 has known functional roles specifically at the synapse in regulating both excitation, and inhibition in the brain [50, 55, 59,60,61,62]. We showed that two NRXN1 variants associated with familial suicide risk resulted in increased binding of LRRTM2, suggesting that synapse regulation may be important in risk. Although our hemi-synapse formation assays did not result in changes in the overall number of synapses, more subtle changes in types of synapses or synapse activity may be key and could dramatically impact neural behavior. Indeed, differences in neuronal distributions of neurexin isoforms and their interacting proteins in the brain are important in neural circuity [15, 50, 62, 63]. Interestingly, we observed an increase, rather than a decrease in binding to LRRTM2. A similar increased binding of LRRTM2 to Nrxn3α has recently been found for an A687T SNP variant related to intellectual disability and epilepsy, resulting in a gain-of-function presynaptic phenotype at excitatory synapses [64]. Ultimately, generation of in vivo models, such as knock-in mouse mutants [64] will be necessary to uncover the functional consequences of the variants in this study.
Previous studies showed that levels of Nrxn1α are important for LRRTM2-mediated excitatory presynaptic differentiation [48]. The NRXN-LRRTM complex was a target in drug-screening for inhibitors because of its link to neurological disorders [65]. Our synapse formation assays did not specifically assess LRRTM2-Nrxn1 synapses, so future work could be directed toward investigating these specific synapses. Finally, NRXN1 may have other unknown functions that are impacted by these variants which were not tested. The dynamics of NRXN1 and binding partners are complex and may additionally be affected by biological processes that exist in vivo [50, 55, 60, 66, 67].
Nature and impact of suicide risk variants
We observed a lack of elevation in allele frequencies of our variants in our full suicide cohort compared to controls, but also observed their presence in other high-risk families. Together, these observations suggest a small effect size of the variants. We acknowledge that effects may be magnified or diminished by the presence/absence of other moderating factors, including other rare variants and/or underlying polygenic risks of psychiatric diagnoses or traits. Post hoc analyses of diagnostic phenotypes did not show unusual clustering of any co-occurring condition among the suicides that were carriers of the two NRXN1 variants compared to non-carrier suicides. In addition, we did not find significant carrier vs. non-carrier differences in polygenic risk scores for selected psychiatric diagnoses. However, we note that we are powered only to observe large differences in phenotypic and polygenic risk effects.
Although we prioritized functional protein-coding variants, allowing the study of mechanism of action, it is also possible that variants in regulatory regions and haplotypes are important for risk, and should be considered in future studies. In a complex phenotype such as suicide, family designs may help identify variants that elevate, rather than drive the risk for suicide. There may be subtle alterations in the regulation of the gene in question, accompanied by additional evidence for the occurrence of rare functional coding variants in that gene in a small minority of cases. Interestingly, a similar scenario was recently described in a familial discovery in the cancer literature [68]. In addition, there is evidence NRXN1 may be proximally and distally regulated [69] and NRXN1 variants can regulate other functionally relevant genes. For example, using the rSNPBase database [70], we discovered intronic NRXN1 variants associated with expression quantitative trait loci genes such as Rho GTPase activating protein 26 (ARHGAP26), which is differentially expressed in the dorsolateral prefrontal cortex between suicide and non-suicide mood disorder subjects [71]. In addition, we note that other familially shared suggestive regulatory genomic segments in our original family study [6] contained genes encoding for proteins that interact with NRXN1 at synapses (LRRTM3, NXPH1) [50, 67], suggesting a potentially more complex role of NRXN1-related synapse pathology in suicide risk. Future studies should also consider a broader pathway-based approach. In conclusion, our findings suggest changes in the NRXN1 gene involving synaptic dynamics may contribute to increased suicide risk. Suicide risk is complex. Genetic changes may elevate risk but are not alone sufficient to cause suicide in the absence of complex interactions with many other genetic and environmental risk factors.
Supplementary information is available at MP’s website.
Data sharing
Data from the parent Utah suicide study has been uploaded into the NIMH Data Archive (NDA) as part of R01MH099134. This data includes demographic data from 4278 Utah suicides, electronic health records data from a subset of 3682 of these suicides, genotypes from 1317 suicides (plink file format), and exome sequence data (.bam file format) from 184 Utah suicides. The project also has a pairwise kinship table that corresponds to these suicide cases. DNA samples from 3347 Utah suicide deaths are at the NIMH Repository (RUCDR). Additional data from the study may be requested by contacting hilary.coon@utah.edu.
References
Suicide Statistics — AFSP [Internet]. [cited 2019 Oct 8]. Available from: https://afsp.org/about-suicide/suicide-statistics/.
Norris D, Clark MS. Evaluation and treatment of the suicidal patient. Am Fam Physician. 2012;85:602–5.
McGuffin P, Marusic A, Farmer A. What can psychiatric genetics offer suicidology. Crisis.2001;22:61–5.
Pedersen NL, Fiske A. Genetic influences on suicide and nonfatal suicidal behavior: twin study findings. Eur Psychiatry. 2010;25:264–7.
Egeland JA, Sussex JN. Suicide and family loading for affective disorders. JAMA.1985;254:915–8.
Coon H, Darlington TM, DiBlasi E, Callor WB, Ferris E, Fraser A, et al. Genome-wide significant regions in 43 Utah high-risk families implicate multiple genes involved in risk for completed suicide. Mol Psychiatry. 2020;25:3077–90.
Docherty AR, Shabalin AA, DiBlasi E, Monson E, Mullins N, Adkins DE, et al. Genome-wide association study of suicide death and polygenic prediction of clinical antecedents. Am J Psychiatry. 2020;177:917–27.
Sokolowski M, Wasserman J, Wasserman D. Genome-wide association studies of suicidal behaviors: a review. Eur Neuropsychopharmacol. 2014;24:1567–77.
Otsuka I, Akiyama M, Shirakawa O, Okazaki S, Momozawa Y, Kamatani Y, et al. Genome-wide association studies identify polygenic effects for completed suicide in the Japanese population. Neuropsychopharmacology. 2019;44:2119–24.
Strawbridge RJ, Ward J, Ferguson A, Graham N, Shaw RJ, Cullen B, et al. Identification of novel genome-wide associations for suicidality in UK Biobank, genetic correlation with psychiatric disorders and polygenic association with completed suicide. EBioMedicine. 2019;41:517–25.
Mirkovic B, Laurent C, Podlipski MA, Frebourg T, Cohen D, Gerardin P. Genetic association studies of suicidal behavior: a review of the past 10 years, progress, limitations, and future directions. Front Psychiatry. 2016;7:158.
Knight S, Abo RP, Abel HJ, Neklason DW, Tuohy TM, Burt RW, et al. Shared genomic segment analysis: the power to find rare disease variants. Ann Hum Genet. 2012;76:500–9.
Lee S, Abecasis GR, Boehnke M, Lin X. Rare-variant association analysis: study designs and statistical tests. Am J Hum Genet. 2014;95:5–23.
van Spronsen M, Hoogenraad CC. Synapse pathology in psychiatric and neurologic disease. Curr Neurol Neurosci Rep. 2010;10:207–14.
Reissner C, Runkel F. Missler M. Neurexins. Genome Biol. 2013;14:213.
Kasem E, Kurihara T, Tabuchi K. Neurexins and neuropsychiatric disorders. Neurosci Res. 2018;127:53–60.
Südhof TC. Neuroligins and neurexins link synaptic function to cognitive disease. Nature.2008;455:903–11.
Südhof TC. Synaptic neurexin complexes: a molecular code for the logic of neural circuits. Cell.2017;171:745–69.
Hu Z, Xiao X, Zhang Z, Li M. Genetic insights and neurobiological implications from NRXN1 in neuropsychiatric disorders. Mol Psychiatry. 2019;24:1400–14.
Béna F, Bruno DL, Eriksson M, van Ravenswaaij-Arts C, Stark Z, Dijkhuizen T, et al. Molecular and clinical characterization of 25 individuals with exonic deletions of NRXN1 and comprehensive review of the literature. Am J Med Genet B Neuropsychiatr Genet. 2013;162B:388–403.
Kirov G, Rujescu D, Ingason A, Collier DA, O’Donovan MC, Owen MJ. Neurexin 1 (NRXN1) deletions in schizophrenia. Schizophr Bull. 2009;35:851–4.
Reichelt AC, Rodgers RJ, Clapcote SJ. The role of neurexins in schizophrenia and autistic spectrum disorder. Neuropharmacology.2012;62:1519–26.
Jenkins A, Apud JA, Zhang F, Decot H, Weinberger DR, Law AJ. Identification of candidate single-nucleotide polymorphisms in NRXN1 related to antipsychotic treatment response in patients with schizophrenia. Neuropsychopharmacology. 2014;39:2170–8.
Lett TA, Tiwari AK, Meltzer HY, Lieberman JA, Potkin SG, Voineskos AN, et al. The putative functional rs1045881 marker of neurexin-1 in schizophrenia and clozapine response. Schizophr Res. 2011;132:121–4.
Grayton HM, Missler M, Collier DA, Fernandes C. Altered social behaviours in neurexin 1α knockout mice resemble core symptoms in neurodevelopmental disorders. PLoS ONE. 2013;8:e67114.
Castronovo P, Baccarin M, Ricciardello A, Picinelli C, Tomaiuolo P, Cucinotta F, et al. Phenotypic spectrum of NRXN1 mono- and bi-allelic deficiency: a systematic review. Clin Genet. 2020;97:125–37.
Dachtler J, Ivorra JL, Rowland TE, Lever C, Rodgers RJ, Clapcote SJ. Heterozygous deletion of α-neurexin I or α-neurexin II results in behaviors relevant to autism and schizophrenia. Behav Neurosci. 2015;129:765–76.
Gaynor SC, Breen ME, Monson ET, de Klerk K, Parsons M, DeLuca AP, et al. A targeted sequencing study of glutamatergic candidate genes in suicide attempters with bipolar disorder. Am J Med Genet B Neuropsychiatr Genet. 2016;171:1080–7.
Higgs BW, Elashoff M, Richman S, Barci B. An online database for brain disease research. BMC Genomics. 2006;7:70.
Reichman RD, Gaynor SC, Monson ET, Gaine ME, Parsons MG, Zandi PP, et al. Targeted sequencing of the LRRTM gene family in suicide attempters with bipolar disorder. Am J Med Genet B Neuropsychiatr Genet. 2020;183:128–39.
de Wit J, O’Sullivan ML, Savas JN, Condomitti G, Caccese MC, Vennekens KM, et al. Unbiased discovery of glypican as a receptor for LRRTM4 in regulating excitatory synapse development. Neuron. 2013;79:696–711.
Lam M, Moslem M, Bryois J, Pronk RJ, Uhlin E, Ellström ID, et al. Single cell analysis of autism patient with bi-allelic NRXN1-alpha deletion reveals skewed fate choice in neural progenitors and impaired neuronal functionality. Exp Cell Res. 2019;383:111469.
Pak C, Danko T, Zhang Y, Aoto J, Anderson G, Maxeiner S. et al. Human neuropsychiatric disease modeling using conditional deletion reveals synaptic transmission defects caused by heterozygous mutations in NRXN1. Cell Stem Cell. 2015;17:316–28.
Avazzadeh S, McDonagh K, Reilly J, Wang Y, Boomkamp SD, McInerney V, et al. Increased Ca2+ signaling in NRXN1α +/- neurons derived from ASD induced pluripotent stem cells. Molecular Autism. 2019 Dec 30 [cited 2020 Sep 6];10. Available from: https://pubmed.ncbi.nlm.nih.gov/31893021/.
Utah Population Database - Huntsman Cancer Institute | University of Utah [Internet]. [cited 2019 Oct 8]. Available from: https://uofuhealth.utah.edu/huntsman/utah-population-database/.
Infinium PsychArray-24 Kit | Psychiatric predisposition microarray [Internet]. [cited 2019 Oct 23]. Available from: https://www.illumina.com/products/by-type/microarray-kits/infinium-psycharray.html.
Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO. et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Whole-Genome Sequencing [Internet]. [cited 2019 Dec 7]. Available from: https://www.illumina.com/techniques/sequencing/dna-sequencing/whole-genome-sequencing.html.
Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics. 2010;26:589–95.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
Paila U, Chapman BA, Kirchner R, Quinlan AR. GEMINI: integrative exploration of genetic variation and genome annotations. PLoS Comput Biol. 2013;9:e1003153.
Layer RM, Chiang C, Quinlan AR, Hall IM. LUMPY: a probabilistic framework for structural variant discovery. Genome Biol. 2014;15:R84.
Dausset J, Cann H, Cohen D, Lathrop M, Lalouel JM, White R. Centre d’etude du polymorphisme humain (CEPH): collaborative genetic mapping of the human genome. Genomics. 1990;6:575–7.
Tschanz JT, Corcoran C, Skoog I, Khachaturian AS, Herrick J, Hayden KM. et al. Dementia: the leading predictor of death in a defined elderly population: the Cache County Study. Neurology. 2004;62:1156–62.
Ng PC, Henikoff S. SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003;31:3812–4.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P. et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
de Wit J, Sylwestrak E, O’Sullivan ML, Otto S, Tiglio K, Savas JN. et al. LRRTM2 interacts with Neurexin1 and regulates excitatory synapse formation. Neuron.2009;64:799–806.
Taniguchi H, Gollan L, Scholl FG, Mahadomrongkul V, Dobler E, Limthong N. et al. Silencing of neuroligin function by postsynaptic neurexins. J Neurosci. 2007;27:2815–24.
Reissner C, Stahn J, Breuer D, Klose M, Pohlentz G, Mormann M. et al. Dystroglycan binding to α-neurexin competes with neurexophilin-1 and neuroligin in the brain. J Biol Chem. 2014;289:27585–603.
Martin EA, Muralidhar S, Wang Z, Cervantes DC, Basu R, Taylor MR. et al. The intellectual disability gene Kirrel3 regulates target-specific mossy fiber synapse development in the hippocampus. Elife. 2015;4:e09395.
Basu R, Duan X, Taylor MR, Martin EA, Muralidhar S, Wang Y. et al. Heterophilic Type II cadherins are required for high-magnitude synaptic potentiation in the hippocampus. Neuron. 2017;96:160.
Biederer T, Scheiffele P. Mixed-culture assays for analyzing neuronal synapse formation. Nat Protoc. 2007;2:670–6.
gnomAD. 2019. Available from: http://gnomad.broadinstitute.org/.
Chih B, Gollan L, Scheiffele P. Alternative splicing controls selective trans-synaptic interactions of the neuroligin-neurexin complex. Neuron. 2006;51:171–8.
Kang Y, Zhang X, Dobie F, Wu H, Craig AM. Induction of GABAergic postsynaptic differentiation by alpha-neurexins. J Biol Chem. 2008;283:2323–34.
Selten M, van Bokhoven H, Nadif, Kasri N. Inhibitory control of the excitatory/inhibitory balance in psychiatric disorders. F1000Res. 2018;7:23.
Schroeder A, de Wit J. Leucine-rich repeat-containing synaptic adhesion molecules as organizers of synaptic specificity and diversity. Exp Mol Med. 2018;9:50.
Ahmad M, Missler M. Neurexins. In: Encyclopedia of Neuroscience. Edited by Squire LR. Oxford: Academic Press; 2009:261–67.
Sheckler LR, Henry L, Sugita S, Südhof TC, Rudenko G. Crystal structure of the second LNS/LG domain from neurexin 1alpha: Ca2+ binding and the effects of alternative splicing. J Biol Chem. 2006;281:22896–905.
Siddiqui TJ, Pancaroglu R, Kang Y, Rooyakkers A, Craig AM. LRRTMs and neuroligins bind neurexins with a differential code to cooperate in glutamate synapse development. J Neurosci. 2010;30:7495–506.
Alabi OO, Davatolhagh MF, Robinson M, Fortunato MP, Vargas Cifuentes L, Kable JW. et al. Disruption of Nrxn1α within excitatory forebrain circuits drives value-based dysfunction. Elife. 2020;9:e54838.
Missler M, Südhof TC. Neurexins: three genes and 1001 products. Trends Genet. 1998;14:20–6.
Restrepo S, Langer NJ, Nelson KA, Aoto J. Modeling a Neurexin-3α human mutation in mouse neurons identifies a novel role in the regulation of transsynaptic signaling and neurotransmitter release at excitatory synapses. J Neurosci. 2019;13:39.
Karki S, Maksimainen MM, Lehtiö L, Kajander T. Inhibitor screening assay for neurexin-LRRTM adhesion protein interaction involved in synaptic maintenance and neurological disorders. Anal Biochem. 2019;15:587.
Wilson SC, White KI, Zhou Q, Pfuetzner RA, Choi UB, Südhof TC. et al. Structures of neurexophilin-neurexin complexes reveal a regulatory mechanism of alternative splicing. EMBO J. 2019;38:e101603.
Um JW, Choi TY, Kang H, Cho YS, Choii G, Uvarov P. et al. LRRTM3 regulates excitatory synapse development through alternative splicing and neurexin binding. Cell Rep. 2016;14:808–22.
Waller RG, Darlington TM, Wei X, Madsen MJ, Thomas A, Curtin K. et al. Novel pedigree analysis implicates DNA repair and chromatin remodeling in multiple myeloma risk. PLoS Genet. 2018;2:e1007111.
Mozhui K, Wang X, Chen J, Mulligan MK, Li Z, Ingles J. et al. Genetic regulation of Nrxn1 [corrected] expression: an integrative cross-species analysis of schizophrenia candidate genes. Transl Psychiatry. 2011;1:e25.
Guo L, Du Y, Chang S, Zhang K, Wang J. rSNPBase: a database for curated regulatory SNPs. Nucleic Acids Res. 2014;42:D1033–9.
Sequeira A, Morgan L, Walsh DM, Cartagena PM, Choudary P, Li J, et al. Gene expression changes in the prefrontal cortex, anterior cingulate cortex and nucleus accumbens of mood disorders subjects that committed suicide. PLoS ONE. 2012;7:e35367.
Acknowledgements
Genotyping was performed by the University of Utah Genomic Core with support from the National Center for Advancing Translational Sciences of the NIH (UL1TR002538) and by llumina, Inc. with support from Janssen Research & Development, LLC. This work was also supported by the National Institute of Mental Health, R01MH105426 (MEW), R01MH099134 (HC), R01MH122412 (HC), R01MH123489 (HC), the Clark Tanner Foundation (HC), a seed grant from the Utah Genome Project (HC, MEW), a donation from the Sharon Kae Lehr Endowed Research Fund in Memory of James Raymond Crump (DG), by the Deutsche Forschungsgemeinschaft (SFB1348 TP A03 to MM) and the IZKF Münster (Mi3-004-19 to MM). Partial support for all data sets within the UPDB was provided by the University of Utah Huntsman Cancer Institute (HCI), and the HCI Cancer Center Support grant, P30CA42014 from the National Cancer Institute. We especially thank University of Utah and OME staff whose hours of work made this study possible. We also thank Matthew Taylor (University of Utah, Salt Lake City, UT, United States) for assistance with the mixed-culture assay and analysis, Harneet Kaur Dhillon (University of Utah, Salt Lake City, UT, United States) for assistance with the analysis of confocal images, Dr. Joris de Wit (Katholieke Universiteit Leuven, Belgium) for plasmids, and those who have assisted with the revisions for this article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
NW takes responsibility for the integrity of the data and the accuracy of the data analysis. QSL is an employee at Janssen Research & Development, LLC and owns stock/stock options in the company; no other authors have conflicts of interest relevant to the content of this paper, including no financial interest, relationships, or affiliations.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
William, N., Reissner, C., Sargent, R. et al. Neurexin 1 variants as risk factors for suicide death. Mol Psychiatry 26, 7436–7445 (2021). https://doi.org/10.1038/s41380-021-01190-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-021-01190-2
This article is cited by
-
Genome-wide association study meta-analysis of suicide death and suicidal behavior
Molecular Psychiatry (2023)
-
Meta-analysis of epigenome-wide association studies of major depressive disorder
Scientific Reports (2022)
-
Molecular Dynamic Simulation of Neurexin1α Mutations Associated with Mental Disorder
Journal of Molecular Neuroscience (2022)