Abstract
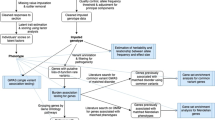
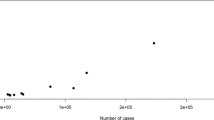
Panic disorder (PD) has a lifetime prevalence of 2–4% and heritability estimates of 40%. The contributory genetic variants remain largely unknown, with few and inconsistent loci having been reported. The present report describes the largest genome-wide association study (GWAS) of PD to date comprising genome-wide genotype data of 2248 clinically well-characterized PD patients and 7992 ethnically matched controls. The samples originated from four European countries (Denmark, Estonia, Germany, and Sweden). Standard GWAS quality control procedures were conducted on each individual dataset, and imputation was performed using the 1000 Genomes Project reference panel. A meta-analysis was then performed using the Ricopili pipeline. No genome-wide significant locus was identified. Leave-one-out analyses generated highly significant polygenic risk scores (PRS) (explained variance of up to 2.6%). Linkage disequilibrium (LD) score regression analysis of the GWAS data showed that the estimated heritability for PD was 28.0–34.2%. After correction for multiple testing, a significant genetic correlation was found between PD and major depressive disorder, depressive symptoms, and neuroticism. A total of 255 single-nucleotide polymorphisms (SNPs) with p < 1 × 10−4 were followed up in an independent sample of 2408 PD patients and 228,470 controls from Denmark, Iceland and the Netherlands. In the combined analysis, SNP rs144783209 showed the strongest association with PD (pcomb = 3.10 × 10−7). Sign tests revealed a significant enrichment of SNPs with a discovery p-value of <0.0001 in the combined follow up cohort (p = 0.048). The present integrative analysis represents a major step towards the elucidation of the genetic susceptibility to PD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kessler RC, Petukhova M, Sampson NA, Zaslavsky AM, Wittchen HU. Twelve-month and lifetime prevalence and lifetime morbid risk of anxiety and mood disorders in the United States. Int J Methods Psychiatr Res. 2012;21:169–84.
Hendriks SM, Spijker J, Licht CM, Beekman AT, Hardeveld F, de Graaf R, et al. Disability in anxiety disorders. J Affect Disord. 2014;166:227–33.
Schumacher J, Deckert J. Serotonin transporter polymorphisms and panic disorder. Genome Med. 2010;2:40.
Hettema JM, Neale MC, Kendler KS. A review and meta-analysis of the genetic epidemiology of anxiety disorders. Am J Psychiatry. 2001;158:1568–78.
Erhardt A, Akula N, Schumacher J, Czamara D, Karbalai N, Muller-Myhsok B, et al. Replication and meta-analysis of TMEM132D gene variants in panic disorder. Transl Psychiatry. 2012;2:e156.
Erhardt A, Czibere L, Roeske D, Lucae S, Unschuld PG, Ripke S, et al. TMEM132D, a new candidate for anxiety phenotypes: evidence from human and mouse studies. Mol Psychiatry. 2011;16:647–63.
Otowa T, Kawamura Y, Nishida N, Sugaya N, Koike A, Yoshida E, et al. Meta-analysis of genome-wide association studies for panic disorder in the Japanese population. Transl Psychiatry. 2012;2:e186.
Otowa T, Hek K, Lee M, Byrne EM, Mirza SS, Nivard MG, et al. Meta-analysis of genome-wide association studies of anxiety disorders. Mol Psychiatry. 2016;21:1391–9.
Schmermund A, Mohlenkamp S, Stang A, Gronemeyer D, Seibel R, Hirche H, et al. Assessment of clinically silent atherosclerotic disease and established and novel risk factors for predicting myocardial infarction and cardiac death in healthy middle-aged subjects: rationale and design of the Heinz Nixdorf RECALL Study. Risk factors, evaluation of coronary calcium and lifestyle. Am Heart J. 2002;144:212–8.
Muglia P, Tozzi F, Galwey NW, Francks C, Upmanyu R, Kong XQ, et al. Genome-wide association study of recurrent major depressive disorder in two European case-control cohorts. Mol Psychiatry. 2010;15:589–601.
Ripke S, O’Dushlaine C, Chambert K, Moran JL, Kahler AK, Akterin S, et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat Genet. 2013;45:1150–9.
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–7.
Witt SH, Streit F, Jungkunz M, Frank J, Awasthi S, Reinbold CS, et al. Genome- wide association study of borderline personality disorder reveals genetic overlap with bipolar disorder, major depression and schizophrenia. Transl Psychiatry. 2017;7:e1155.
Delaneau O, Marchini J, Zagury JF. A linear complexity phasing method for thousands of genomes. Nat Methods. 2011;9:179–81.
Howie B, Marchini J, Stephens M. Genotype imputation with thousands of genomes. G3. 2011;1:457–70.
1000 Genomes Project Consortium, Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, et al. A map of human genome variation from population-scale sequencing. Nature. 2010;467:1061–73.
Lam M, Awasthi S, Watson HJ, Goldstein J, Panagiotaropoulou G, Trubetskoy V, et al. RICOPILI: rapid imputation for COnsortias PIpeLIne. Bioinformatics. 2019 (in press).
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
International Schizophrenia Consortium, Purcell SM, Wray NR, Stone JL, Visscher PM, O’Donovan MC, et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature. 2009;460:748–52.
Bulik-Sullivan B, Finucane HK, Anttila V, Gusev A, Day FR, Loh PR, et al. An atlas of genetic correlations across human diseases and traits. Nat Genet. 2015;47:1236–41.
Zheng J, Erzurumluoglu AM, Elsworth BL, Kemp JP, Howe L, Haycock PC, et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics. 2017;33:272–9.
Duncan LE, Ratanatharathorn A, Aiello AE, Almli LM, Amstadter AB, Ashley- Koch AE, et al. Largest GWAS of PTSD (N=20 070) yields genetic overlap with schizophrenia and sex differences in heritability. Mol Psychiatry. 2018;23:666–73.
International Obsessive Compulsive Disorder Foundation Genetics Collaborative (IOCDF-GC) and OCD Collaborative Genetics Association Studies (OCGAS). Revealing the complex genetic architecture of obsessive-compulsive disorder using meta-analysis. Mol Psychiatry. 2018;23:1181–8.
Speed D, Balding DJ. SumHer better estimates the SNP heritability of complex traits from summary statistics. Nat Genet. 2019;51:277–84.
de Leeuw CA, Mooij JM, Heskes T, Posthuma D. MAGMA: generalized gene- set analysis of GWAS data. PLoS Comput Biol. 2015;11:e1004219.
Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Dold M, Bartova L, Souery D, Mendlewicz J, Serretti A, Porcelli S, et al. Clinical characteristics and treatment outcomes of patients with major depressive disorder and comorbid anxiety disorders - results from a European multicenter study. J Psychiatr Res. 2017;91:1–13.
Demirkan A, Penninx BW, Hek K, Wray NR, Amin N, Aulchenko YS, et al. Genetic risk profiles for depression and anxiety in adult and elderly cohorts. Mol Psychiatry. 2011;16:773–83.
Vohma U, Aluoja A, Vasar V, Shlik J, Maron E. Evaluation of personality traits in panic disorder using Swedish universities Scales of Personality. J Anxiety Disord. 2010;24:141–6.
Bienvenu OJ, Hettema JM, Neale MC, Prescott CA, Kendler KS. Low extraversion and high neuroticism as indices of genetic and environmental risk for social phobia, agoraphobia, and animal phobia. Am J Psychiatry. 2007;164:1714–21.
Kendler KS, Gatz M, Gardner CO, Pedersen NL. Personality and major depression: a Swedish longitudinal, population-based twin study. Arch Gen Psychiatry. 2006;63:1113–20.
Navrady LB, Adams MJ, Chan SWY, Major Depressive Disorder Working Group of the Psychiatric Genomics Consortium, Ritchie SJ, McIntosh AM. Genetic risk of major depressive disorder: the moderating and mediating effects of neuroticism and psychological resilience on clinical and self-reported depression. Psychol Med. 2018;48:1890–9.
Goldstein BL, Klein DN. A review of selected candidate endophenotypes for depression. Clin Psychol Rev. 2014;34:417–27.
Dresler T, Guhn A, Tupak SV, Ehlis AC, Herrmann MJ, Fallgatter AJ, et al. Revise the revised? New dimensions of the neuroanatomical hypothesis of panic disorder. J Neural Transm. 2013;120:3–29.
Kim MJ, Loucks RA, Palmer AL, Brown AC, Solomon KM, Marchante AN, et al. The structural and functional connectivity of the amygdala: from normal emotion to pathological anxiety. Behav Brain Res. 2011;223:403–10.
Pfleiderer B, Zinkirciran S, Arolt V, Heindel W, Deckert J, Domschke K. fMRI amygdala activation during a spontaneous panic attack in a patient with panic disorder. World J Biol Psychiatry. 2007;8:269–72.
Sullivan PF, Agrawal A, Bulik CM, Andreassen OA, Borglum AD, Breen G, et al. Psychiatric genomics: an update and an agenda. Am J Psychiatry. 2018;175:15–27.
GTEx Consortium. Human genomics. The genotype-tissue expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science. 2015;348:648–60.
Acknowledgements
The authors are grateful to all patients and control subjects for their participation. We thank Kristina Annerbrink (Gothenburg), Marie Olsson (Gothenburg) and Monica Hellberg (Stockholm) for their support. The study was supported by the German Research Foundation (DFG; grant SCHU1596/4-1 to JS). Controls for the Germany I sample were drawn from the Heinz Nixdorf Recall Study (HNR) cohort, which was established with the support of the Heinz Nixdorf Foundation. A. Metspalu was supported by EstRC grant IUT20-60 and by the European Union through the European Regional Development Fund (Project No. 2014-2020.4.01.15-0012), Centre of Excellence “GenTransMed”. This work was conducted within the framework of the German multicenter trial “Mechanisms of Action in CBT (MAC)”. The MAC study was funded by the German Federal Ministry of Education and Research (BMBF; project no. 01GV0615), as part of the BMBF Psychotherapy Research Funding Initiative. Principal investigators (PI) with respective areas of responsibility in the MAC study are: VA (Münster: Overall MAC Program Coordination); HUW (Dresden: PI for the Randomized Clinical Trial (RCT) and Manual Development); AH (Greifswald: PI for Psychophysiology); ALG (Münster: PI for Psychophysiology and Panic subtypes); AS (Berlin: PI for Experimental Pharmacology); TK (Marburg: PI for functional neuroimaging); and JD (Würzburg: PI for Genetics). Additional site directors for the RCT component of the program are GWA (Würzburg); TF and LF (Berlin-Adlershof); and TL (Bremen). Acknowledgements and staff members according to site Greifswald (coordinating site for Psychophysiology): Christiane Melzig, Jan Richter, Susan Richter, Matthias von Rad; Berlin-Charité (coordinating Center for Experimental Pharmacology): Harald Bruhn, Anja Siegmund, Meline Stoy, André Wittmann; Berlin-Adlershof: Irene Schulz; Münster (Overall MAC Program Coordination, Genetics and Functional Neuroimaging): Andreas Behnken, Katharina Domschke, Adrianna Ewert, Carsten Konrad, Bettina Pfleiderer, Christina Uhlmann, Peter Zwanzger; Münster (coordinating site for psychophysiology and subtyping): Judith Eidecker, Swantje Koller, Fred Rist, Anna Vossbeck-Elsebusch; Marburg/Aachen (coordinating center for functional neuroimaging): Barbara Drüke, Sonja Eskens, Thomas Forkmann, Siegfried Gauggel, Susan Gruber, Andreas Jansen, Thilo Kellermann, Isabelle Reinhardt, Nina Vercamer-Fabri; Dresden (coordinating site for data collection, analysis, and the RCT): Franziska Einsle, Christine Froehlich, Andrew T. Gloster, Christina Hauke, Simone Heinze, Michael Hoefler, Ulrike Lueken, Peter Neudeck, Stephanie Preiß, Dorte Westphal; Würzburg Psychiatry Department (coordinating center for genetics): AR, Caro Gagel; Würzburg Psychology Department: Julia Duerner, Hedwig Eisenbarth, Antje B. M. Gerdes, Harald Krebs, PP,, Silvia Schad, Nina Steinhäuser; Bremen: Veronika Bamann, Sylvia Helbig-Lang, Anne Kordt, Pia Ley, Franz Petermann, Eva-Maria Schroeder. Additional support was provided by the coordinating center for clinical studies in Dresden (KKS Dresden): Xina Graehlert and Marko Käppler. This work is also part of the German multicenter trial “Mechanisms of CBT-treatment effects in patients with panic disorder and panic disorder with agoraphobia: The role of interoceptive exposure and fear augmentation (MCBT-PDAS)”. The MCBT-PDAS study is funded by the German Federal Ministry of Education and Research (BMBF, 01GV0614), as part of the larger BMBF Psychotherapy Research Funding Initiative “Improving the Treatment of Panic Disorder”. The PIs of the MCBT-PDAS study are: Alfons Hamm (Greifswald: PI Psychophysiology); TL (Bremen: Study Director for the RCT and Manual Development); ALG (Münster: PI Panic subtypes); GWA (Mannheim: PI Ambulatory assessment); CPF (Greifswald: PI Psychophysiology and Panic Disorder); TK (Marburg: PI for functional neuroimaging), and JD (Würzburg: PI for Genetics). Additional site directors for the RCT component of the program are Winfried Rief (Marburg), and PP (Würzburg). Centers of the Research Network: VA (Münster: Overall Network Coordination); HUW (Dresden); Andreas Ströhle (Berlin). Data Access and Responsibility: All PIs take responsibility for the integrity of the respective study data and their components. All authors and co-authors had full access to all study data. Data analysis and manuscript preparation were completed by the authors and co-authors of this article, who take responsibility for its accuracy and content. Acknowledgements and staff members by site: Bremen (coordinating center for the multicenter trial): Veronika Bamann, Sandra Cammin, Sarah Czilwik, Kira Geisler, Sylvia Helbig-Lang, Kirsten Helmes, Anne Kordt, Tanja Leonhard, Mila Plett- Perelshteyn, Christian Soltau, Juliane Sülz, Maxie von Auer; Greifswald (coordinating site for psychophysiology): Anett Hoffmann, Jan Richter; Mannheim (coordinating center for ambulatory assessment): Christoph Biwer, Elisabeth Borgmann, Antje Gerdes, Otto Martin, Kristina Steinbach, Bettina Stemmler, Andrew White; Marburg (coordinating center for functional neuroimaging): Tobias Fehlinger, Andreas Jansen, Nikita Jegan, Carsten Konrad, Marion Mickeler, Silke Rusch, Katrin Schlötterer, Benjamin Straube, Mareike Stumpenhorst, Katrin Wambach, Yunbo Yang; Münster (coordinating site for panic subtypes): Susanne Kettler, Anna Vossbeck-Elsebusch; Würzburg Psychiatry Department (coordinating center for genetics): Carola Gagel, Andreas Reif, Heike Weber; Würzburg Psychology Department: Almut Friedl-Huber, Harald Krebs, Caroline Ott, Nina Steinhäuser. Additional support was provided by the coordinating center for clinical studies in Dresden (KKS Dresden): Marko Käppler. The study was registered with the NCT01323556. Funding for the Netherland Twin Register and Netherlands Study of Depression and Anxiety (NESDA) was obtained from the Netherlands Organization for Scientific Research (NWO) and MagW/ZonMW grants Middelgroot-911-09-032, Spinozapremie 56-464-14192, Geestkracht program of the Netherlands Organization for Health Research and Development (ZonMW 10-000-1002), Center for Medical Systems Biology (CSMB, NOW Genomics), Genetic influences on stability and change in psychopathology from childhood to young adulthood (ZonMW 912-10-020), NBIC/BioAssist/RK(2008.024), Biobanking and Biomolecular Resources Research Infrastructure (BBMRI–NL, 184.021.007), VU University’s Institute for Health and Care Research (EMGO+) and Neuroscience Campus Amsterdam (NCA); the European Science Council (ERC Advanced, 230374). The genotyping and analyses were funded in part by the Genetic Association Information Network (GAIN) of the Foundation for the National Institutes of Health, Rutgers University Cell and DNA Repository (NIMH U24 MH068457-06); the Avera Institute for Human Genetics, Sioux Falls, South Dakota (USA); and the National Institutes of Health (NIH R01 HD042157-01A1, MH081802, Grand Opportunity grants 1RC2 MH089951 and 1RC2 MH089995). The iPSYCH project is funded by the Lundbeck Foundation (grant numbers R102- A9118 and R155-2014-1724) and the universities and university hospitals of Aarhus and Copenhagen. The Danish National Biobank resource was supported by the Novo Nordisk Foundation. Data handling and analysis on the GenomeDK HPC facility was supported by NIMH (1U01MH109514-01 to Michael O’Donovan and ADB). High-performance computer capacity for handling and statistical analysis of iPSYCH data on the GenomeDK HPC facility was provided by the Center for Genomics and Personalized Medicine, Aarhus University and Central Region Denmark, and Centre for Integrative Sequencing, iSEQ, Aarhus University (grant to ADB).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
TET, SS, HS and KS are employed by deCODE Genetics/Amgen. TW has acted as advisor and lecturer to H. Lundbeck A/S. The other authors declare that they have no conflict of interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Forstner, A.J., Awasthi, S., Wolf, C. et al. Genome-wide association study of panic disorder reveals genetic overlap with neuroticism and depression. Mol Psychiatry 26, 4179–4190 (2021). https://doi.org/10.1038/s41380-019-0590-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-019-0590-2
This article is cited by
-
The impact of assortative mating, participation bias and socioeconomic status on the polygenic risk of behavioural and psychiatric traits
Nature Human Behaviour (2024)
-
Enhanced interhemispheric resting-state functional connectivity of the visual network is an early treatment response of paroxetine in patients with panic disorder
European Archives of Psychiatry and Clinical Neuroscience (2024)
-
Multi-trait analysis characterizes the genetics of thyroid function and identifies causal associations with clinical implications
Nature Communications (2024)
-
Causal associations between female reproductive behaviors and psychiatric disorders: a lifecourse Mendelian randomization study
BMC Psychiatry (2023)
-
Causal relationships between blood lipids and major psychiatric disorders: Univariable and multivariable mendelian randomization analysis
BMC Medical Genomics (2023)