Abstract
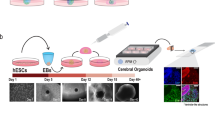
The typical abnormalities observed in the brain of Alzheimer’s disease (AD) patients include synaptic alterations, neuronal death, brain inflammation, and the accumulation of protein aggregates in the form of amyloid plaques and neurofibrillary tangles. Despite the development of many animal and in vitro models for AD, there is a lack of an experimental approach that fully recapitulates essential aspects of the disease in human cells. Here, we report the generation of a new model to study AD, consisting of cerebral organoids (COs) produced from human-induced pluripotent stem cells (iPSCs). Under our experimental conditions, COs grow to form three-dimensional (3D) structures containing neural areas with cortical-like organization. Analysis of COs by histological and biochemical methods revealed that organoids produced from iPSCs derived from patients affected by familial AD or Down syndrome (DS) spontaneously develop over time pathological features of AD, including accumulation of structures highly reminiscent to amyloid plaques and neurofibrillary tangles. These pathological abnormalities were not observed in COs generated from various controls, including human iPSCs from healthy individuals, human iPSCs from patients affected by Creutzfeldt–Jakob disease, mouse embryonic stem cells (ESCs), or mouse iPSCs. These findings enable modeling genetic AD in a human cellular context in a 3D cortical-like tissue developed in vitro from patient-specific stem cells. This system provides a more relevant disease model compared to pre-existing methods and offers a new platform for discovery of novel targets and screening of drugs for therapeutic intervention.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Querfurth HW, LaFerla FM. Alzheimer’s disease. N Engl J Med. 2010;362:329–44.
Laurijssens B, Aujard F, Rahman A. Animal models of Alzheimer’s disease and drug development. Drug Discov Today Technol. 2013;10:e319–27.
Duyckaerts C, Potier MC, Delatour B. Alzheimer disease models and human neuropathology: similarities and differences. Acta Neuropathol. 2008;115:5–38.
Stadtfeld M, Hochedlinger K. Induced pluripotency: history, mechanisms, and applications. Genes Dev. 2010;24:2239–63.
Yagi T, Ito D, Okada Y, Akamatsu W, Nihei Y, Yoshizaki T. et al. Modeling familial Alzheimer’s disease with induced pluripotent stem cells. Hum Mol Genet. 2011;20:4530–9.
Israel MA, Yuan SH, Bardy C, Reyna SM, Mu Y, Herrera C. et al. Probing sporadic and familial Alzheimer’s disease using induced pluripotent stem cells. Nature. 2012;482:216–20.
Kondo T, Asai M, Tsukita K, Kutoku Y, Ohsawa Y, Sunada Y. et al. Modeling Alzheimer’s disease with iPSCs reveals stress phenotypes associated with intracellular Abeta and differential drug responsiveness. Cell Stem Cell. 2013;12:487–96.
Shi Y, Kirwan P, Smith J, MacLean G, Orkin SH, Livesey FJ. A human stem cell model of early Alzheimer’s disease pathology in Down syndrome. Sci Transl Med. 2012;4:124ra29
Muratore CR, Rice HC, Srikanth P, Callahan DG, Shin T, Benjamin LN. et al. The familial Alzheimer’s disease APPV717I mutation alters APP processing and Tau expression in iPSC-derived neurons. Hum Mol Genet. 2014;23:3523–36.
Moore S, Evans LD, Andersson T, Portelius E, Smith J, Dias TB. et al. APP metabolism regulates tau proteostasis in human cerebral cortex neurons. Cell Rep. 2015;11:689–96.
Koch P, Tamboli IY, Mertens J, Wunderlich P, Ladewig J, Stuber K. et al. Presenilin-1 L166P mutant human pluripotent stem cell-derived neurons exhibit partial loss of gamma-secretase activity in endogenous amyloid-beta generation. Am J Pathol. 2012;180:2404–16.
Sproul AA, Jacob S, Pre D, Kim SH, Nestor MW, Navarro-Sobrino M. et al. Characterization and molecular profiling of PSEN1 familial Alzheimer’s disease iPSC-derived neural progenitors. PLoS ONE. 2014;9:e84547
Mahairaki V, Ryu J, Peters A, Chang Q, Li T, Park TS. et al. Induced pluripotent stem cells from familial Alzheimer’s disease patients differentiate into mature neurons with amyloidogenic properties. Stem Cells Dev. 2014;23:2996–3010.
Armijo E, Gonzalez C, Shahnawaz M, Flores A, Davis B, Soto C. Increased susceptibility to Abeta toxicity in neuronal cultures derived from familial Alzheimer’s disease (PSEN1-A246E) induced pluripotent stem cells. Neurosci Lett. 2017;639:74–81.
Lancaster MA, Renner M, Martin CA, Wenzel D, Bicknell LS, Hurles ME. et al. Cerebral organoids model human brain development and microcephaly. Nature. 2013;501:373–9.
Lancaster MA, Knoblich JA. Generation of cerebral organoids from human pluripotent stem cells. Nat Protoc. 2014;9:2329–40.
Sherrington R, Rogaev EI, Liang Y, Rogaeva EA, Levesque G, Ikeda M. et al. Cloning of a gene bearing missense mutations in early-onset familial Alzheimer’s disease. Nature. 1995;375:754–60.
Nethercott HE, Brick DJ, Schwartz PH. Derivation of induced pluripotent stem cells by lentiviral transduction. Methods Mol Biol. 2011;767:67–85.
Liu Y, Zheng Y, Li S, Xue H, Schmitt K, Hergenroeder GW. et al. Human neural progenitors derived from integration-free iPSCs for SCI therapy. Stem Cell Res. 2017;19:55–64.
Renner M, Lancaster MA, Bian S, Choi H, Ku T, Peer A. et al. Self-organized developmental patterning and differentiation in cerebral organoids. EMBO J. 2017;36:1316–29.
Klunk WE, Wang Y, Huang GF, Debnath ML, Holt DP, Mathis CA. Uncharged thioflavin-T derivatives bind to amyloid-beta protein with high affinity and readily enter the brain. Life Sci. 2001;69:1471–84.
Kawarabayashi T, Younkin LH, Saido TC, Shoji M, Ashe KH, Younkin SG. Age-dependent changes in brain, CSF, and plasma amyloid (beta) protein in the Tg2576 transgenic mouse model of Alzheimer’s disease. J Neurosci. 2001;21:372–81.
Rostagno A, Ghiso J. Isolation and biochemical characterization of amyloid plaques and paired helical filaments. Curr Protoc Cell Biol. 2009;Chapter 3:Unit 3.33.
Porter AG, Janicke RU. Emerging roles of caspase-3 in apoptosis. Cell Death Differ. 1999;6:99–104.
Zhang D, Pekkanen-Mattila M, Shahsavani M, Falk A, Teixeira AI, Herland A. A 3D Alzheimer’s disease culture model and the induction of P21-activated kinase mediated sensing in iPSC derived neurons. Biomaterials. 2014;35:1420–8.
Choi SH, Kim YH, Hebisch M, Sliwinski C, Lee S, D’Avanzo C. et al. A three-dimensional human neural cell culture model of Alzheimer’s disease. Nature. 2014;515:274–8.
Raja WK, Mungenast AE, Lin YT, Ko T, Abdurrob F, Seo J. et al. Self-organizing 3D human neural tissue derived from induced pluripotent stem cells recapitulate Alzheimer’s disease phenotypes. PLoS ONE. 2016;11:e0161969
Garcez PP, Loiola EC, Madeiro da CR, Higa LM, Trindade P, Delvecchio R. et al. Zika virus impairs growth in human neurospheres and brain organoids. Science. 2016;352:816–8.
Huch M, Knoblich JA, Lutolf MP, Martinez-Arias A. The hope and the hype of organoid research. Development. 2017;144:938–41.
Musiek ES, Holtzman DM. Three dimensions of the amyloid hypothesis: time, space and ‘wingmen’. Nat Neurosci. 2015;18:800–6.
Lancaster MA, Knoblich JA. Organogenesis in a dish: modeling development and disease using organoid technologies. Science. 2014;345:1247125
Qian X, Nguyen HN, Song MM, Hadiono C, Ogden SC, Hammack C. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell. 2016;165:1238–54.
Acknowledgements
We are grateful to Dr. Brian Davis (University of Texas Medical School at Houston) for his valuable help in establishing the iPSC technology in our lab, Dr. Fabrizio Tagliavini (Istituto Carlo Besta, Italy) for kindly providing human fibroblasts from patients affected by inherited Creutzfeldt–Jakob disease. We are also grateful to Dr. Ying Liu (Institute of Molecular Medicine, University of Texas at Houston) for providing an iPSC line obtained from healthy controls. This project was partially funded by a pilot grant from the University of Texas Brain Initiative Program.
Author contributions
CG designed the studies, carried out many of the experiments, analyzed the results, and prepared the first version of the figures. EA generated and characterized induced pluripotent stem cells used in these studies, performed some of the experiments and histological staining, analyzed some of the results, prepared the final version of the figures, and assisted in the preparation and writing of the final version of the manuscript. JB-A performed the biochemical studies to measure the levels of Aβ and tau aggregates. AB-C participated in the studies with mouse cerebral organoids. CM performed the western blot and ELISA studies of p-tau, generated induced pluripotent stem cells, produced the cerebral organoids derived from Creutzfeldt–Jakob disease cells, processed tissue samples, and assisted in the preparation and writing of the final version of the manuscript. CS is the principal investigator on the project and was responsible for coordinating research activity, analyzing the data, funding, writing the manuscript, and producing the final version of the article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Gonzalez, C., Armijo, E., Bravo-Alegria, J. et al. Modeling amyloid beta and tau pathology in human cerebral organoids. Mol Psychiatry 23, 2363–2374 (2018). https://doi.org/10.1038/s41380-018-0229-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-018-0229-8
This article is cited by
-
Neuropathogenesis-on-chips for neurodegenerative diseases
Nature Communications (2024)
-
Genetic forms of tauopathies: inherited causes and implications of Alzheimer’s disease-like TAU pathology in primary and secondary tauopathies
Journal of Neurology (2024)
-
Sporadic Creutzfeldt–Jakob disease infected human cerebral organoids retain the original human brain subtype features following transmission to humanized transgenic mice
Acta Neuropathologica Communications (2023)
-
Amyloid β-based therapy for Alzheimer’s disease: challenges, successes and future
Signal Transduction and Targeted Therapy (2023)
-
Cerebral organoids with chromosome 21 trisomy secrete Alzheimer’s disease-related soluble aggregates detectable by single-molecule-fluorescence and super-resolution microscopy
Molecular Psychiatry (2023)