Abstract
Cognitive functions are important correlates of health outcomes across the life-course. Individual differences in cognitive functions are partly heritable. Epigenetic modifications, such as DNA methylation, are susceptible to both genetic and environmental factors and may provide insights into individual differences in cognitive functions. Epigenome-wide meta-analyses for blood-based DNA methylation levels at ~420,000 CpG sites were performed for seven measures of cognitive functioning using data from 11 cohorts. CpGs that passed a Bonferroni correction, adjusting for the number of CpGs and cognitive tests, were assessed for: longitudinal change; being under genetic control (methylation QTLs); and associations with brain health (structural MRI), brain methylation and Alzheimer's disease pathology. Across the seven measures of cognitive functioning (meta-analysis n range: 2557–6809), there were epigenome-wide significant (P < 1.7 × 10-8) associations for global cognitive function (cg21450381, P = 1.6 × 10-8), and phonemic verbal fluency (cg12507869, P = 2.5 × 10-9). The CpGs are located in an intergenic region on chromosome 12 and the INPP5A gene on chromosome 10, respectively. Both probes have moderate correlations (~0.4) with brain methylation in Brodmann area 20 (ventral temporal cortex). Neither probe showed evidence of longitudinal change in late-life or associations with white matter brain MRI measures in one cohort with these data. A methylation QTL analysis suggested that rs113565688 was a cis methylation QTL for cg12507869 (P = 5 × 10-5 and 4 × 10-13 in two lookup cohorts). We demonstrate a link between blood-based DNA methylation and measures of phonemic verbal fluency and global cognitive ability. Further research is warranted to understand the mechanisms linking genomic regulatory changes with cognitive function to health and disease.
Similar content being viewed by others
Background
Cognitive function is an important predictor of health outcomes and mortality [1,2,3,4]. Whether this is due to differences in health literacy and lifestyle choices or if there is a biological predisposition is not clear [5]. The complex balance between genetic and environmental contributions to cognitive function is poorly understood [6]. Epigenetic modifications may provide insight into the link between cognitive function, perturbed biological pathways and relevance for lifelong health.
Molecular genetic studies of unrelated individuals show that around 30% of the variance in general cognitive function can be explained by common genetic polymorphisms (single-nucleotide polymorphisms: SNPs) and variants in linkage disequilibrium with them [7,8,9]. However, there are relatively few well-established individual SNP predictors of cognitive function and those that have been identified explain a very small proportion of the variance in cognitive test scores [8].
Epigenetic marks may help us better understand the interaction between genes, the environment, and health-related quantitative traits, such as cognitive function, and common disease outcomes [10, 11]. The epigenome helps to regulate genes via, for example, chemical modifications to DNA. DNA methylation typically refers to the addition of a methyl group to a cytosine nucleotide placed next to a guanine in the DNA sequence. The addition or removal of the methyl group is a dynamic process and can be tissue specific with, for example, different epigenetic signatures in blood and brain. The proportion of cytosines methylated at a specific CpG site can be partly explained by both genetics and lifestyle/environment or a combination of these [12]. Studies have examined the association between DNA methylation with genotype [13, 14], metabolic factors, such as body mass index [15, 16], and environmental factors, such as smoking [17]. However, no large-scale population-based studies have examined the association of cognitive function with DNA methylation in circulating leucocytes.
One aspect of note for epigenetic epidemiology studies of brain-related traits (cognitive functions, schizophrenia, depression, dementia, etc.) is tissue (and cellular) specificity. As brain samples are not likely to be available until post-mortem, a proxy tissue is an attractive possibility to be explored for building relevant epigenetic signatures. In epidemiological studies, the most likely candidate is blood, which, although its methylation patterns are often dissimilar to those in the brain [18, 19], they have still been linked to mental health traits [20,21,22]. Identifying robust methylomic differences in relation to cognitive traits may improve our ability to predict cognitive decline and better understand the mechanistic link between cognitive function and deleterious health outcomes.
Here, we examine, using a meta-analytic approach, the associations between blood-based DNA methylation and several individual tests of cognitive functions in up to 6809 healthy, older-aged adults. First we test which, if any, CpG probes are associated with individual cognitive functions at an epigenome-wide level. Then we investigate these probes to see if they are (1) under genetic control (methQTLs), (2) stable over time, (3) associated with structural brain-imaging measures, (4) associated with Alzheimer's disease case–control status or neuropathology, (5) associated with DNA methylation levels in different brain regions and (6) associated with blood-based gene expression.
Methods
Overview
Epigenome-wide association studies were performed in 11 independent cohorts for seven cognitive function phenotypes. The number of cohorts contributing to each of the seven tests of cognitive function ranged from 3 to 10 (Table S1). A sample-size-based meta-analysis of Z-scores was performed on the overlapping cohort summary output for each cognitive test.
Cohorts
Nine of the eleven cohorts that contributed to the analysis included participants of European ancestry: Framingham Heart Study, InCHIANTI, Lothian Birth Cohort 1921, Lothian Birth Cohort 1936, MOBILIZE Boston, Normative Aging Study, Rotterdam Study (Rotterdam Bios and Rotterdam III) and Twins UK. The Atherosclerosis Risk in the Community (ARIC) and Genetic Epidemiology Network of Arteriopathy's (GENOA) cohorts included participants of African American ancestry. Details of each cohort are presented in Appendix 1.
Cognitive measures
Scores from seven different cognitive tests were assessed:
-
1.
Wechsler Logical Memory [23, 24] as a measure of verbal declarative memory. The sum of the immediate and delayed tasks was used.
-
2.
Wechsler Digit Symbol Test [25] or Symbol Digit Modalities Test [26] or Letter Digit Substitution Test [27] as a measure of processing speed, hereafter referred to as Digit Test. The total number of correct answers in the allocated time period was used. The three tests listed above are highly correlated [28].
-
3.
Semantic Verbal Fluency [29] as a measure of an aspect of executive function (animal naming - total score).
-
4.
Phonemic Verbal Fluency [29] as a measure of an aspect of executive function (letter fluency - total score).
-
5.
Trail Making Test Part B [30] as a measure of an aspect of executive function (Natural log (ln) of the time taken in seconds).
-
6.
Boston Naming Test [31] or National Adult Reading Test [32] or any other measure of vocabulary. The total number of correct answers was assessed.
-
7.
Mini-Mental State Examination (MMSE) [33] as a measure of general cognitive function. Individuals with a score of less than 24 out of 30 were excluded from the analysis.
With the exception of the MMSE scores, any cognitive score that fell above or below 3.5 standard deviations from the mean was set to the mean plus or minus 3.5 standard deviations, respectively. These analyses were performed within each cohort independently for each cognitive test. Full details of the tests available within each cohort are provided in Appendix 1.
DNA methylation
Whole-blood DNA methylation was assessed in each cohort using the Illumina HumanMethylation450 BeadChips [34]. Quality control was performed according to cohort-specific thresholds, described in Appendix 1. The blood samples for DNA methylation and cognitive ability were measured concurrently.
Structural brain imaging
1.5 T structural brain imaging was assessed in one of the participating epigenome-wide association study (EWAS) cohorts: The Lothian Birth Cohort 1936. Full details have been reported previously [35]. Here, we considered two measures of white matter connectivity—fractional anisotropy (directional coherence of water diffusion) and mean diffusivity (average magnitude of water diffusion)—that have been previously associated with cognitive function [36, 37].
Gene expression
The association between DNA methylation and gene expression was assessed using the Affymetrix Human Exon 1.0 ST Array in one of the participating cohorts: The Framingham Heart Study. Methodological details are provided in Appendix 1.
Ethics
Ethical permission for each cohort is described in Appendix 1. Written informed consent was obtained from all subjects.
Statistical analysis
Epigenome-wide association testing
For each cognitive test, two linear regression models were considered—a basic-adjustment model and a full-adjustment model. Both models treated methylation at the CpG sites (untransformed methylation beta value) as the dependent variable with the cognitive test score as the independent predictor of interest. In the basic-adjustment model, covariates included age, sex, white-blood cell counts (either measured or imputed [38]), technical covariates such as plate, chip, array and hybridisation date, and, where required, genetic principal components to account for population stratification. In the fully adjusted model, the following additional covariate terms were included: a quadratic term for age, an age x sex interaction; smoking status (current, ever, never) and body mass index. The findings from the fully adjusted model were considered as the primary output. Measurement details for all variables are presented in Appendix 1. Age was standardised within cohort to mean 0, variance 1, to avoid potential model convergence issues. Individuals with prevalent dementia or clinical stroke (including self-reported) were excluded.
Quality control filtering
Prior to the meta analysis, all probes on sex chromosomes were removed along with non-CpG probes, and any cross-reactive probes as reported by Chen et al. [39]. Genomic correction was applied to any cohort-specific results file with an empirical lambda of more than 1. The total number of probes included in the meta-analysis for each cognitive trait ranged between 421,335 and 421,633.
Trait-specific meta-analysis
The primary analyses were conducted in R [40]; sample-size weighted meta-analyses were conducted in METAL [41]. Several significance thresholds were considered. The most liberal threshold was a within meta-analysis Benjamini-Hochberg false discovery rate of Q < 0.05. Next was a within meta-analysis Bonferroni corrected P-value threshold: 0.05/nprobes_max = 0.05/421,633 = 1.2 × 10-7. Finally, the most conservative threshold applied was a Bonferroni corrected P-value that also adjusted for the seven meta analyses: 0.05/(nprobes*nmeta-analyses) = 0.05/(421,633*7) = 1.7 × 10-8.
Summary meta-analysis combining all cognitive traits
Finally, a meta-analysis of the summary output from the seven meta analyses was conducted for the fully adjusted models using the CPASSOC software [42] in R. As the cohorts contributed to multiple EWAS, and as the as cognitive test scores are positively correlated [43], a correlation matrix of the CpG Z-scores for the seven cognitive traits was included to reduce the false-positive rate [42]. A test assuming heterogeneity was assumed and default input arguments were set.
Methylation quantitative trait loci
To determine if the significant EWAS findings (at the most conservative threshold of P < 1.7 × 10-8) were partly under genetic control, a methylation QTL analysis lookup was performed using data from the Lothian Birth Cohorts of 1921 and 1936 (combined n = 1366), and the Brisbane Systems Genetics Study (n = 614) [44]. The discovery and replication thresholds set in that study were P < 1 × 10-11 and P < 1 × 10-6, respectively, with the combined LBC cohorts acting as a discovery data set (P < 1 × 10-11) with BSGS as the replication study (P < 1 × 10-6) and vice versa. SNPs within 2 Mbp of a CpG site were labelled cis methylation QTLs, and only the most significant SNP for each CpG were considered.
Longitudinal change in methylation
For the significantly associated CpG probes identified in the meta-analyses, longitudinal data from the Lothian Birth Cohort 1936 were used to chart change in methylation at these CpGs between ages 70 and 76 years. Stability in methylation levels might be indicative of potential genetic control or a long-term fixed effect of differential cognitive function on the probe. Variability in methylation levels may be a by-product or cause of cognitive change over time. Methylation data were available on participants at ages 70 (n = 920), 73 (n = 800) and 76 (n = 618) years. Linear mixed models with random intercept terms, adjusting for sex, imputed white-blood cell counts and technical variables, were used to determine the rate of change over time (the coefficient for the fixed effect age variable in the model) for each probe.
Structural brain-imaging associations with methylation
As cognitive function is a brain-related phenotype, it was of interest to see if blood-based methylation signatures for cognitive function were related to brain-imaging measures. Structural MRI data and covariate information were also available in 552 participants at the second wave of the Lothian Birth Cohort 1936—data from only the first wave of the cohort were included in the EWAS. The top associations from the EWAS meta-analyses were assessed at the second wave of the Lothian Birth Cohort 1936 in relation to age- and sex-adjusted brain structural fractional anisotropy and mean diffusivity using linear regression models, adjusting for age, sex, imputed white cell counts and technical covariates.
Blood–brain methylation correlations
Lookup analyses of significant CpG sites were performed in published data sets for both blood and brain (prefrontal cortex, entorhinal cortex, superior temporal gyrus and cerebellum) based EWAS findings for Braak staging and Alzheimer's disease status [21]. A second lookup was performed using results from blood and Brodmann areas 7, 10 and 20 from post-mortem samples of 16 individuals [45].
Gene expression associations
Transcriptome-wide association studies (TWAS) were conducted in the Framingham Heart Study for any significant probes from the cognitive EWAS. Linear mixed effects models with expression of each gene as the dependent variable, methylation as exposure and identical covariates to the EWAS were considered. A Bonferroni correction was applied (P < 0.05/nprobes = 0.05/17,873 = 2.8 × 10-6) to identify statistically significant associations.
Results
Study sample characteristics
Participants came from 11 cohorts—ranging in size from 219 to 2307 individuals (Q1–Q3: 435–920), with between 0 and 100% female participants (Q1–Q3: 52–65%), mean age ranged from 56 to 79 years (Q1–Q3: 60–73). Two of the cohorts (ARIC and GENOA) included participants of African American ancestry; all other cohorts included participants of European ancestry. The cohort-specific summary details for each cognitive test are presented in Supplementary Table 1. The basic-adjustment meta-analytic sample-size ranged from 2557 individuals for the Trail Making Test to 6809 individuals for the MMSE. Similar sample-sizes were observed for the fully adjusted models with the meta-analytic results presented in Fig. 1 and Table 1.
Epigenome-wide association study model diagnostics
Heterogeneity was observed in the EWAS inflation statistics, both within and across cohorts (Supplementary Table 2). For example, the minimum and maximum lambda values in LBC1936 were 1.05 and 1.25, respectively. Prior to meta-analysis, within-cohort genomic correction was applied where lambda exceeded 1. The meta analysis genomic inflation statistics for the basic and fully adjusted models ranged from 0.93 to 1.30, and 0.92 to 1.26, respectively (Table 1).
Epigenome-wide association study of seven cognitive traits
A list of the within-test epigenome-wide significant associations within a given cognitive test across both models are presented in Supplementary Table 3. Significant associations (P < 1.2 × 10-7) were observed in the basic and full adjustment models for Phonemic Verbal Fluency (n = 4 and n = 2), MMSE (n = 1 for both models), Vocabulary (n = 3 and n = 1), and Digit Test (n = 29 and n = 2). From the basic-adjustment model, significant CpGs were located in genes associated with, for example: alcohol metabolism (ALDH2, Digit Test, cg12142865) [46], smoking (AHRR, Digit Test, cg05575921) [17], inflammation (CCR9 and PRRC2A, cg10475172 and cg14943908, respectively) [47, 48] and neurodegeneration through the beta-amyloid precursor protein interactor GAPDH (Digit Test, cg00252813) [49]. In the fully adjusted model, significant CpGs were located in genes associated with, for example: inflammation (SOCS3, Digit Test, cg18181703) [50], epithelial cell splicing (ESRP2, Vocabulary, cg04513006) [51] and transcription activation of NOTCH proteins (MAML3, Phonemic Verbal Fluency, cg16201957) [52]. No CpGs were significantly associated with the Trail Making, Logical Memory or Semantic Verbal Fluency tests. Methylation at cg21450381 was not associated with any of the six other cognitive traits in the fully adjusted meta-analytic results at a nominal significance threshold of P < 0.05 (Table 2). However, cg12507869 was associated with lower scores for both Logical Memory (P = 0.043) and Vocabulary (P = 9.4 × 10-5).
Variation in results when modifying the significance threshold
Using a less conservative FDR correction for multiple testing identified associations at a q-value threshold of 0.05 in both the basic and fully adjusted models for Phonemic Verbal Fluency (n = 49 and n = 2), MMSE (n = 1 for both models), Vocabulary (n = 7 and n = 3) and Digit Test (n = 309 and n = 14). The FDR-significant probes are presented in Supplementary Table 4.
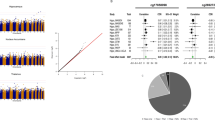
After Bonferroni correction for CpG sites and the seven cognitive traits—P < 0.05/(420,000*7)—two remaining differentially methylated CpGs were cg21450381 (R2 = 0.47%, P = 1.6 × 10-8) with MMSE scores, and cg12507869 (R2 = 0.55%, P = 2.5 × 10-9) with Phonemic Verbal Fluency. In both cases, higher methylation was associated with lower cognitive scores across all of the contributing cohorts. cg21450281 is located in an intergenic region of chromosome 12; cg12507869 is located in the inositol polyphosphate-5-phosphatase, 40 kDa (INPP5A) gene on chromosome 10. Both probes were approximately normally distributed in the Lothian Birth Cohort 1936 (Fig. 2). A forest plot of the Z-scores by cohort sample-size is presented in Fig. 3 and shows no evidence of ethnic outliers or single cohorts driving the associations.
Forest plots of the Z-scores by cohort sample size for the two significant CpGs. ARIC Atherosclerosis Risk in the Community, FHS Framingham Heart Study Offspring Cohort, GENOA Genetic Epidemiology Network of Arteriopathy, InCHIANTI Invecchiare in Chianti, LBC Lothian Birth Cohort, MOBILIZE Maintenance of Balance, Independent Living, Intellect and Zest in the Elderly of Boston, NAS Normative Aging Study, RS Rotterdam Study, RS-Bios Rotterdam Study—Biobank-based Integrative Omics Studies
Combined meta-analysis of all seven cognitive traits
There was no evidence from the combined meta-analytic results of the seven tests for a globally significant CpG across all tests in the fully adjusted model (minimum Benjamini-Hochberg FDR q-value of 0.057 for cg12507869).
Genetic contributions to cognitive-related differential methylation
A methylation QTL lookup [44] analyses identified no SNPs to be associated with cg21450381. The top SNP for cg12507869 (rs113565688 in the INPP5A gene on chromosome 10) explained around 1.2% of the variance in methylation (P-values of 3.6 × 10-13 and 5.4 × 10-5 in the Australian and Scottish cohorts, respectively). There is no overlap of this SNP with cognitive traits based on a recent GWAS conducted in the UK Biobank cohort: rs113565688 association with memory (P = 0.55), reaction time (P = 0.42), verbal-numerical reasoning (P = 0.17) and educational attainment (P = 0.13) [8].
Longitudinal changes in methylation at cognitive-related differential methylation sites
Longitudinal analyses over three waves of data (ages 70, 73 and 76 years) from LBC1936, adjusting for sex, imputed white-blood cell counts and technical variables, found no evidence for a linear change in the methylation of either probe over a relatively narrow period in later-life. The mixed model standardised effect size for change in cg12507869 was 0.02 standard deviations per year, P = 0.13; the standardised effect size for cg21450381 was -0.02, P = 0.40. Without adjustment for covariates, the across wave correlations for cg12507869 were 0.62 (age 70: age 73), 0.63 (age 70: age 76) and 0.68 (age 73: age 76). The corresponding correlations for cg21450381 were 0.04, 0.10 and 0.30, respectively.
Association of brain MRI features with cognitive-related differential methylation
There were no significant associations between the top two CpGs and either of the brain MRI measures of white matter connectivity (mean diffusivity minimum P = 0.56; fractional anisotropy minimum P = 0.28) at age 73 in the LBC1936 (n = 552).
Correlation of blood and brain methylation at the cognitive-related differential methylation
Two blood–brain comparisons were conducted. The first, using a blood–brain DNA methylation comparison tool [18] [http://epigenetics.essex.ac.uk/bloodbrain/], provided no evidence for a significant correlation between blood-methylation at either probe with methylation in four brain regions (prefrontal cortex, entorhinal cortex, superior temporal gyrus and cerebellum, Supplementary Figs. 1 and 2). Whereas the mean of the cg21450381 probe was similar to the means for the four brain regions, the mean of the cg12507869 probe in blood was markedly different (hypomethylated) to the means for the prefrontal cortex, entorhinal cortex, superior temporal gyrus (Supplementary Figs. 1 and 2). It was, however, similar to the mean of the cerebellum. The second comparison, using BECon [45] [https://redgar598.shinyapps.io/BECon/] showed the same mean methylation levels for cg21450381 between blood and Brodmann areas 7, 10 and 20; cg12507869 was again hypomethylated in blood compared to the three brain regions. There were moderate correlations between blood-methylation and Brodmann area 20 for both CpGs (r = 0.43 for cg12507869 and r = -0.46 for cg21450381) and between Brodmann area 7 and cg12507869 (r = 0.31).
Association of cognitive-related differential methylation with Braak staging and Alzheimer’s disease
None of the six CpGs that were epigenome-wide significant in the fully adjusted EWASs at P < 1.2 × 10-7 were associated with Braak staging or Alzheimer's case–control status in either blood or brain-based methylation (minimum FDR q-value 0.51, Supplementary Table 5).
Transcriptome-wide association study
There were no significant TWAS results for cg21450381. The minimum P-value observed was 0.00013 (Q = 0.51). There were nine significant TWAS results for cg12507869 at P < 2.8 × 10-6 and 41 at Q < 0.05. There was a nominal inverse association between the INPP5A transcript and CpG (P = 0.049, Q = 0.65). The full TWAS output for the two CpGs is shown in Supplementary Tables 6 and 7.
Discussion
This study presents a meta-analysis of the relationship between blood-based DNA methylation and cognitive function. We analysed seven different cognitive tests and found two epigenome-wide methylation correlations: cg21450381, located in an intergenic region of chromosome 12, with global cognitive function (as measured by the MMSE); and cg12507869, located in the INPP5A gene on chromosome 10, with phonemic verbal fluency. Methylation at the latter CpG was also associated with two other cognitive tests (logical memory and vocabulary) at a nominal P < 0.05 threshold. Genetic analyses of the top two CpGs showed a modest cis regulation for one of the probes, suggesting that the vast majority of the methylation variation at the cognitive-related differentially methylated sites are due to environmental influences. Blood-based methylation levels at both of the CpGs correlated with methylation levels in Brodmann area 20 (cerebral cortex).
INPP5A is a member of the inositol polyphosphate-5-phosphatase (INPP5) family of genes that encode enzymes that hydrolyse inositol 1,4,5 triphosphate (IP3). It is involved in the mobilisation of intracellular calcium, and has been implicated in cerebellar degeneration in mice [53]. A second INPP5 family member, INPP5D, has been associated with Alzheimer’s Disease and cognitive decline [54, 55], further implicating this gene family in cognitive functions. cg21450381 is located in an intergenic region of chromosome 12, that contains a histone modification mark (H3K27Ac), DNAaseI hypersensitivity clusters and evidence of transcription factor-binding sites, which indicates that the region may be involved in gene regulation [56].
In a TWAS analysis of the top two probes in the Framingham Heart Study (n > 1900), there was no evidence for an association between cg21450381 and blood-based gene expression. Of the nine Bonferroni-significant transcripts in the TWAS of cg12507869, eight were trans associations, with the cis association occurring in ADAM12, which is more than 6 Mb from INPP5A. There was no evidence of a cis effect of the CpG on the INPP5A expression levels.
Disentangling correlation from causation is particularly tricky when studying epigenetic marks in a non-target tissue. By increasing the sample sizes of the meta-analytic EWAS and replicating any findings across different cognitive domains will reduce the chances of false-positive associations. It is, of course, possible that a reliable blood-based epigenetic marker of cognitive function may be several degrees of separation away from the biological processes that drive cognitive skills. For example, the signal could be in response to neurotoxic events, such as inflammation, oxidative stress or small vessel disease. However, the discrimination of cause from consequence is something that affects many epigenetic epidemiology studies. Approaches that may overcome this include Mendelian randomisation studies where a methQTL can be used as an instrument, or the use of mouse models to dissect functional consequences of DNA methylation on gene regulation.
There are additional limitations of this study: a varying number of participants with cognitive data available for each test; heterogeneity in relation to the ethnicity and geographical location of the participants across cohorts; and relating a blood-based methylation signature to a brain-based outcome. We attempted to counter these limitations by: plotting cohort sample-size by Z-score to see if there was bias due to outliers or clustering by ethnicity; adjusting for population stratification in the cohorts with admixture; correlating the blood-based CpG associations with methylation levels in several brain regions; looking at the association between brain region-specific methylation and Alzheimer's disease phenotypes for the blood-based CpG associations. It is possible that bias may have been introduced in the secondary analyses that focussed on the MRI, gene expression and longitudinal methylation data, as both the LBC1936 and Framingham studies contributed to the discovery meta-analyses. Re-running the meta-analyses without these cohorts yielded: P-values of 1.3 × 10-7 and 7.1 × 10-6 for the phonemic verbal fluency finding (cg12507869), excluding Framingham and LBC1936, respectively; and P-values of 1.7 × 10-8 and 3.3 × 10-6 for the MMSE finding (cg21450381), again excluding Framingham and LBC1936, respectively. Whereas the longitudinal methylation and MRI findings were null, the cis and trans expression-methylation associations warrant confirmation in an independent sample. The methQTL findings were based on highly stringent discovery and replication P-value thresholds in both LBC and an independent cohort, BSGS.
Neither of the top two CpGs showed signs of linear change in methylation levels between the ages of 70 and 76 years in one of the participating studies (LBC1936) that had three waves of longitudinal data. It is possible that non-linear changes may be present although additional waves of data collection would be required to test this robustly. In addition, a 6-year window is possibly too narrow to observe substantial changes in the CpG levels.
It is notable that the two significant CpG associations were found for the cognitive tests that were completed by the largest number of participants (n > 6000). The study provided results for a list of cognitive tests that cover several major cognitive domains: memory, processing speed, executive function, vocabulary and global ability. The heterogeneity with respect to ethnicity and geographic location can allows us to generalise our findings to multiple populations.
Blood is the most feasible tissue for epigenetic epidemiology analyses of cognitive function. Brain would be the ideal target tissue although this would make it impossible to have simultaneous cognitive function data. Moreover, epigenome-wide studies of other brain-related outcomes, such as schizophrenia, have identified putative blood-based methylation signatures [22].
In conclusion, we have presented evidence for blood-based epigenetic correlates of cognitive function. Specifically, we identified methylation sites that are linked to an aspect of executive function and global cognitive ability. The latter finding relied on a relatively crude cognitive test (the MMSE), which is commonly used to identify individuals at risk of dementia. One of the two CpG sites identified was under modest genetic control, with a cis SNP explaining over 1% of its variance. Unlike other traits, such as smoking and body mass index [15, 17], there are relatively modest methylation signatures for cognitive function. However, our analyses concur with other recent studies to suggest that blood-based methylation signatures may be useful tools to interrogate differences in brain-related outcomes.
References
Calvin CM, Deary IJ, Fenton C, Roberts BA, Der G, Leckenby N, et al. Intelligence in youth and all-cause-mortality: systematic review with meta-analysis. Int J Epidemiol. 2011;40:626–44.
Deary IJ, Weiss A, Batty GD. Intelligence and personality as predictors of illness and death: How researchers in differential psychology and chronic disease epidemiology are collaborating to understand and address health inequalities. Psychol Sci Public Interest. 2010;11:53–79.
Henderson M, Richards M, Stansfeld S, Hotopf M. The association between childhood cognitive ability and adult long-term sickness absence in three British birth cohorts: a cohort study. BMJ Open. 2012;2:e000777.
Sörberg A, Lundin A, Allebeck P, Melin B, Falkstedt D, Hemmingsson T. Cognitive ability in late adolescence and disability pension in middle age: follow-up of a national cohort of Swedish males. PLoS ONE. 2013;8:e78268.
Hagenaars SP, Harris SE, Davies G, Hill WD, Liewald DC, Ritchie SJ, et al. Shared genetic aetiology between cognitive functions and physical and mental health in UK Biobank (N = 112 151) and 24 GWAS consortia. Mol Psychiatry. 2016;21:1624–32.
Deary IJ, Johnson W, Houlihan LM. Genetic foundations of human intelligence. Hum Genet. 2009;126:215–32.
Davies G, Armstrong N, Bis JC, Bressler J, Chouraki V, Giddaluru S, et al. Genetic contributions to variation in general cognitive function: a meta-analysis of genome-wide association studies in the CHARGE consortium (N = 53949). Mol Psychiatry. 2015;20:183–92.
Davies G, Marioni RE, Liewald DC, Hill WD, Hagenaars SP, Harris SE, et al. Genome-wide association study of cognitive functions and educational attainment in UK Biobank (N = 112 151). Mol Psychiatry. 2016;21:758–67.
Marioni RE, Davies G, Hayward C, Liewald D, Kerr SM, Campbell A, et al. Molecular genetic contributions to socioeconomic status and intelligence. Intelligence. 2014;44:26–32.
Bakulski KM, Fallin MD. Epigenetic epidemiology: promises for public health research. Environ Mol Mutagen. 2014;55:171–83.
Rakyan VK, Down TA, Balding DJ, Beck S. Epigenome-wide association studies for common human diseases. Nat Rev Genet. 2011;12:529–41.
Shah S, McRae AF, Marioni RE, Harris SE, Gibson J, Henders AK, et al. Genetic and environmental exposures constrain epigenetic drift over the human life course. Genome Res. 2014;24:1725–33.
Lemire M, Zaidi SH, Ban M, Ge B, Aïssi D, Germain M, et al. Long-range epigenetic regulation is conferred by genetic variation located at thousands of independent loci. Nat Commun. 2015;6:6326.
Gaunt TR, Shihab HA, Hemani G, Min JL, Woodward G, Lyttleton O, et al. Systematic identification of genetic influences on methylation across the human life course. Genome Biol. 2016;17:61.
Dick KJ, Nelson CP, Tsaprouni L, Sandling JK, Aïssi D, Wahl S, et al. DNA methylation and body-mass index: a genome-wide analysis. Lancet. 2014;383:1990–8.
Mendelson MM, Marioni RE, Joehanes R, Liu C, Hedman ÅK, Aslibekyan S, et al. Association of body mass index with DNA methylation and gene expression in blood cells and relations to cardiometabolic disease: A Mendelian randomization approach. PLoS Med. 2017;14:e1002215.
Joehanes R, Just AC, Marioni RE, Pilling LC, Reynolds LM, Mandaviya PR, et al. Epigenetic signatures of cigarette smoking. Circ Cardiovasc Genet. 2016;9:436–7.
Hannon E, Lunnon K, Schalkwyk L, Mill J. Interindividual methylomic variation across blood, cortex, and cerebellum: implications for epigenetic studies of neurological and neuropsychiatric phenotypes. Epigenetics. 2015;10:1024–32.
Walton E, Hass J, Liu J, Roffman JL, Bernardoni F, Roessner V, et al. Correspondence of DNA methylation between blood and brain tissue and its application to schizophrenia research. Schizophr Bull. 2016;42:406–14.
Hannon E, Dempster E, Viana J, Burrage J, Smith AR, Macdonald R, et al. An integrated genetic-epigenetic analysis of schizophrenia: evidence for co-localization of genetic associations and differential DNA methylation. Genome Biol. 2016;17:176.
Lunnon K, Smith R, Hannon E, De Jager PL, Srivastava G, Volta M, et al. Methylomic profiling implicates cortical deregulation of ANK1 in Alzheimer's disease. Nat Neurosci. 2014;17:1164–70.
Montano C, Taub MA, Jaffe A, Briem E, Feinberg JI, Trygvadottir R, et al. Association of DNA methylation differences with Schizophrenia in an Epigenome-Wide Association Study JAMA Psychiatry. 2016;73:506–14.
Wechsler D. WMS-IIIUK administration and scoring manual. London, UK: Psychological Corporation; 1998.
Wechsler D. Wechsler Memory Scale - Revised. New York: Psychological Corporation, New York, NY; 1987
Wechsler D. WAIS-IIIUK administration and scoring manual. London, UK: Psychological Corporation; 1998.
Smith A. Symbol Digit Modalities Test manual - revised. Los Angeles, CA, USA: Western Psychological Services; 1992.
van der Elst W, van Boxtel MP, van Breukelen GJ, Jolles J. The Letter Digit Substitution Test: normative data for 1,858 healthy participants aged 24-81 from the Maastricht Aging Study (MAAS): influence of age, education, and sex. J Clin Exp Neuropsychol. 2006;28:998–1009.
Ibrahim-Verbaas CA, Bressler J, Debette S, Schuur M, Smith AV, Bis JC, et al. GWAS for executive function and processing speed suggests involvement of the CADM2 gene. Mol Psychiatry. 2016;21:189–97.
Lezak M. Neuropsychological testing. Oxford, UK: Oxford University Press; 2004.
Bowie CR, Harvey PD. Administration and interpretation of the Trail Making Test. Nat Protoc. 2006;1:2277–81.
Kaplan E, Goodglass H, Weintraub S. Boston Naming Test. Philadelphia: Lea & Febiger; 1983.
Nelson HE, Willison JR. National Adult Reading Test (NART) Test Manual (Part II). Windsor, UK: NFER-Nelson; 1991.
Folstein MF, Folstein SE, McHugh PR. "Mini-mental state". A practical method for grading the cognitive state of patients for the clinician. J Psychiatr Res. 1975;12:189–98.
Bibikova M, Barnes B, Tsan C, Ho V, Klotzle B, Le JM, et al. High density DNA methylation array with single CpG site resolution. Genomics. 2011;98:288–95.
Wardlaw JM, Bastin ME, Valdes Hernandez MC, Munoz Maniega S, Royle NA, Morris Z, et al. Brain ageing, cognition in youth and old age, and vascular disease in the Lothian Birth Cohort 1936: rationale, design and methodology of the imaging protocol. Int J Stroke. 2011;6:547–59.
Booth T, Bastin ME, Penke L, Maniega SM, Murray C, Royle NA, et al. Brain white matter integrity and cognitive abilities in community-dwelling older people: the Lothian Birth Cohort 1936. Neuropsychology. 2013;27:595–607.
Penke L, Maniega SM, Bastin ME, Hernandez MCV, Murray C, Royle NA, et al. Brain white matter integrity as a neural foundation for general intelligence. Mol Psychiatry. 2012;17:1026–30.
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinforma. 2012;13:86.
Chen YA, Lemire M, Choufani S, Butcher DT, Grafodatskaya D, Zanke BW, et al. Discovery of cross-reactive probes and polymorphic CpGs in the Illumina Infinium HumanMethylation450 microarray. Epigenetics. 2013;8:203–9.
R Core Team R: A language and environment for statistical computing. 2017. R Foundation for Statistical Computing, Vienna, Austria.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
Zhu X, Feng T, Tayo BO, Liang J, Young JH, Franceschini N, et al. Meta-analysis of correlated traits via summary statistics from GWASs with an application in hypertension. Am J Hum Genet. 2015;96:21–36.
Spearman C. "General Intelligence," objectively determined and measured. Am J Psychol. 1904;15:201–92.
McRae AF, Marioni RE, Shah S, Yang J, Powell JE, Harris SE, Gibson J, Henders AK, Bowdler L, Painter JN, Murphy L, Martin NG, Starr JM, Wray NR, Deary IJ, Visscher PM, Montgomery GW. Identification of 55,000 Replicated DNA Methylation QTL. bioRxiv 166710; https://doi.org/10.1101/166710
Edgar RD, Jones MJ, Meaney MJ, Turecki G, Kobor MS. BECon: A tool for interpreting DNA methylation findings from blood in the context of brain. BioRxiv https://doi.org/10.1101/111609
Edenberg HJ. The genetics of alcohol metabolism: role of alcohol dehydrogenase and aldehyde dehydrogenase variants. Alcohol Res Health. 2007;30:5–13.
Hashimoto M, Nakamura N, Obayashi H, Kimura F, Moriwaki A, Hasegawa G, et al. Genetic contribution of the BAT2 gene microsatellite polymorphism to the age-at-onset of insulin-dependent diabetes mellitus. Hum Genet. 1999;105:197–9.
Zabel BA, Agace WW, Campbell JJ, Heath HM, Parent D, Roberts AI, et al. Human G protein-coupled receptor GPR-9-6/CC chemokine receptor 9 is selectively expressed on intestinal homing T lymphocytes, mucosal lymphocytes, and thymocytes and is required for thymus-expressed chemokine-mediated chemotaxis. J Exp Med. 1999;190:1241–56.
Butterfield DA, Hardas SS, Lange ML. Oxidatively modified glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and Alzheimer's disease: many pathways to neurodegeneration. J Alzheimers Dis. 2010;20:369–93.
Carow B, Rottenberg ME. SOCS3, a major regulator of infection and inflammation. Front Immunol. 2014;5:58.
Warzecha CC, Sato TK, Nabet B, Hogenesch JB, Carstens RP. ESRP1 and ESRP2 are epithelial cell-type-specific regulators of FGFR2 splicing. Mol Cell. 2009;33:591–601.
Wu L, Sun T, Kobayashi K, Gao P, Griffin JD. Identification of a family of mastermind-like transcriptional coactivators for mammalian notch receptors. Mol Cell Biol. 2002;22:7688–700.
Yang AW, Sachs AJ, Nystuen AM. Deletion of Inpp5a causes ataxia and cerebellar degeneration in mice. Neurogenetics. 2015;16:277–85.
Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer's disease. Nat Genet. 2013;45:1452–8.
Andrews SJ, Das D, Anstey KJ, Easteal S. Late onset Alzheimer's disease risk variants in cognitive decline: The PATH through life study. J Alzheimers Dis. 2017;57:423–36.
Rosenbloom KR, Sloan CA, Malladi VS, Dreszer TR, Learned K, Kirkup VM, et al. ENCODE data in the UCSC Genome Browser: year 5 update. Nucl. Acids Res. 2013;41:D56–63. (Database issue)
Acknowledgements
This project was carried out through the Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) consortium: Epigenetics Working Group. Individual cohort acknowledgements are presented in Appendix 1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marioni, R.E., McRae, A.F., Bressler, J. et al. Meta-analysis of epigenome-wide association studies of cognitive abilities. Mol Psychiatry 23, 2133–2144 (2018). https://doi.org/10.1038/s41380-017-0008-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-017-0008-y
This article is cited by
-
DNA methylation as a potential mediator of the association between indoor air pollution and neurodevelopmental delay in a South African birth cohort
Clinical Epigenetics (2023)
-
A correlation map of genome-wide DNA methylation patterns between paired human brain and buccal samples
Clinical Epigenetics (2022)
-
Genetic variation, brain, and intelligence differences
Molecular Psychiatry (2022)
-
A comparison of the genes and genesets identified by GWAS and EWAS of fifteen complex traits
Nature Communications (2022)
-
Novel data archival system for multi-omics data of human exposure to harmful substances
Molecular & Cellular Toxicology (2022)